Figures & data
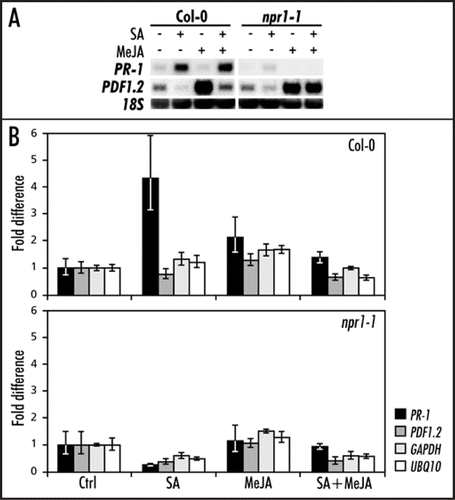
Figure 1 Chromatin immunoprecipitation (ChIP) analysis of histone H3 acetylation at the PR-1 and PDF1.2 promoters in Col-0 and npr1-1. (A) Northern blot analysis of PDF1.2 and PR-1 expression in Col-0 and npr1-1 leaves treated with 1 mM SA, 0.1 mM MeJA, or a combination of both chemicals. Leaf tissue was harvested 24 h after chemical treatment. Equal loading of RNA samples was checked using a probe for 18S rRNA. (B) ChIP analysis of Col-0 and npr1-1 treated with 1 mM SA, 0.1 mM MeJA, or a combination of both chemicals. Chromatin samples were subjected to immunoprecipitation using an AcH3 antibody. The immunoprecipitated DNA was analyzed for the enrichment of PR-1, PDF1.2, UBQ10 and GAPDH promoter sequences by Q-RT-PCR. The fold increase of immunoprecipitated DNA was calculated relative to the input DNA.