Figures & data
Figure 1 Relaxed syntenies shared between the R. oryzae and E. cuniculi genomes. The numbers on each genome indicate the common shared gene clusters. Pink boxes indicate the sex and sex-related locus. Note the size of the scale bar differs between the species.
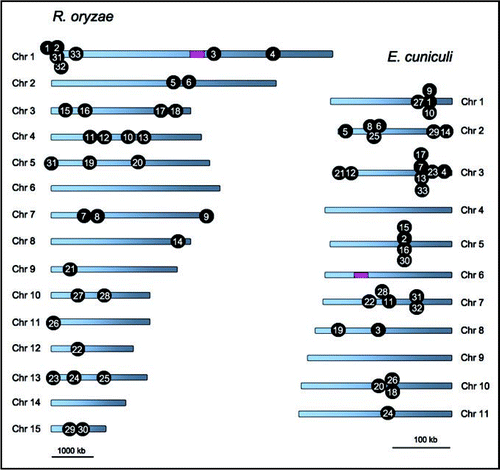
Figure 2 The RPL21 and RPS9 gene cluster is fungal-specific. The synteny is conserved in euascomycetes (i.e., Neurospora crassa, Paracoccidioides brasiliensis, Botrytis cinerea, Chaetomium globosum) and hemi-ascomycete (i.e., Saccharomyces cerevisiae), homobasidiomycete (i.e., Laccaria bicolor) and heterobasidiomycetes (i.e., Puccinia graminis f. sp. tritici and Cryptococcus neoformans) as well as in Mucor circinelloides and the three microsporidia. However, the genes are not syntenic and are completely unlinked in three protozoan, Entamoeba histolytica, Entamoeba dispar and Giardia lamblia genomes. The genome of Schizosaccharomyces pombe encodes RPS9 unlinked to RPL21, and instead RPS4 forms a gene cluster with RPL21. Gene sizes are not to scale.
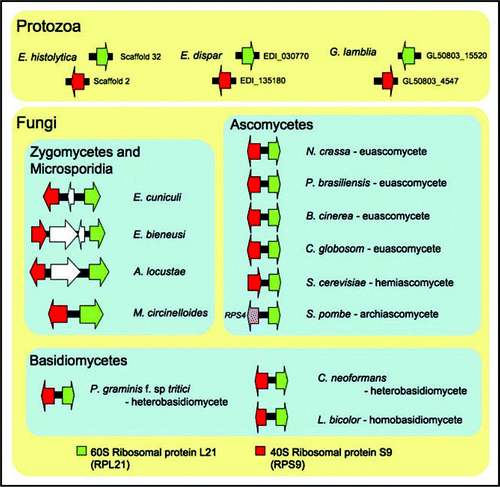
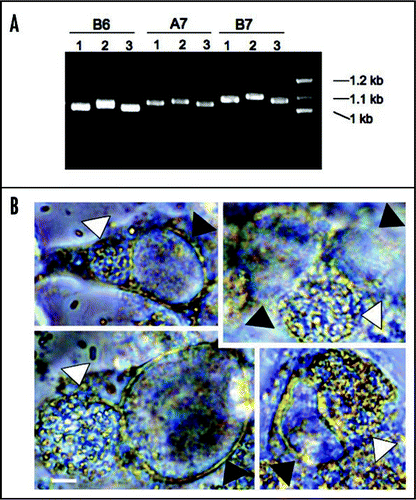
Figure 3 Genetic variation of tandem repeat DNAs in different E. cuniculi isolates and co-infection of E. cuniculi and C. trachomatis. (A) Examples of TR DNAs. Three E. cuniculi isolates (1 is for EC1, 2 for EC2 and 3 for EC3) have different sized banding patterns. B6, A7 and B7 are TR areas chosen with the Tandem Repeat Finder (see the text). (B) E. cuniculi and C. trachomatis infect the same host cell forming two independent parasitophorous vacuoles. White arrowheads indicate E. cuniculi vacuoles and black arrowheads indicate C. trachomatis vacuoles. Scale = 5 µm.