Figures & data
Figure 1. The three host plant B. tabaci Q populations (cotton, cucumber, and cabbage) used in this study.
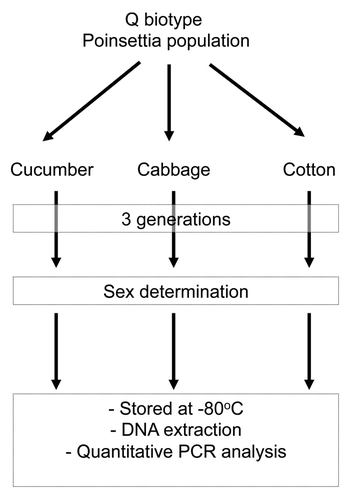
Table 1. Oligonucleotide primers used in quantitative PCR
Figure 2. Relative amount of Portiera and Hamiltonella in three B. tabaci Q populations (cucumber, cabbage, and cotton) as determined by quantitative PCR (normalized according to the amount of actin gene). Values for relative amount of symbionts are means ± SEM of three replicates for each kind of plant. The data were analyzed with one-way analysis of variance. For each kind of symbiont, different letters above the bars indicate significant differences among the three populations (Tukey test, P < 0.05).
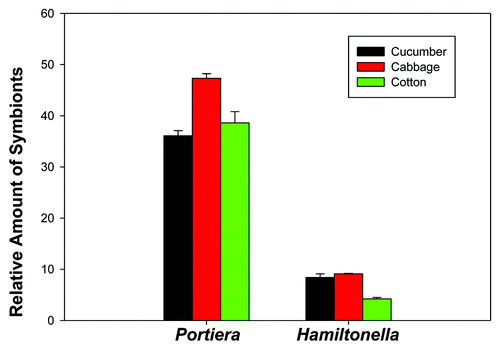
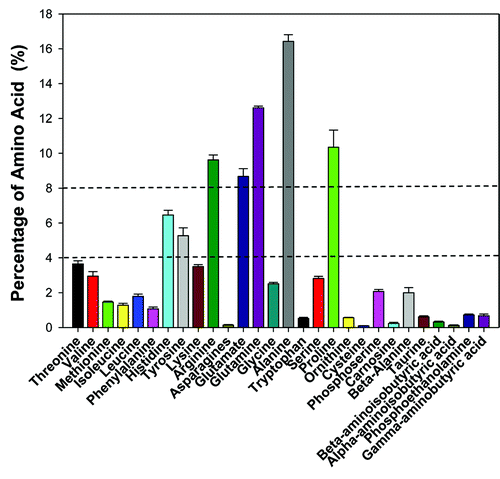
Figure 3. Free amino acids composition in the B. tabaci Q poinsettia population. Free amino acid in whiteflies was analyzed by amino acids analyzer (S433, sykam, Germany). A total of 28 different peaks were detected, the peak area represents the respective amino acid content. The percentage of the various amino acids was calculated by the formula: the respective amino acid content / the total amino acids content × 100%.