Figures & data
Figure 1 (A) The enrichment of GO annotation terms (biological process level 3) in the phenotypic classes “centrosome duplication/separation.” “Centrosome structure” and “no phenotype” were analyzed using DAVID Functional Annotation Tool with the 251 MS-identified centrosomal candidate proteins as background. Bars indicate the fold enrichment of the respective GO terms in the phenotypic classes. (B) Semantic similarity scores (i) within and (ii) between the three protein sets. Scores were calculated with the Bioconductor package ‘GOSemSim’ (Version 1.8.2). N gives the total number of proteins, N effective gives the number of proteins with GO annotation sufficient for score calculation. p values for score differences were calculated using the Mann-Whitney U test.
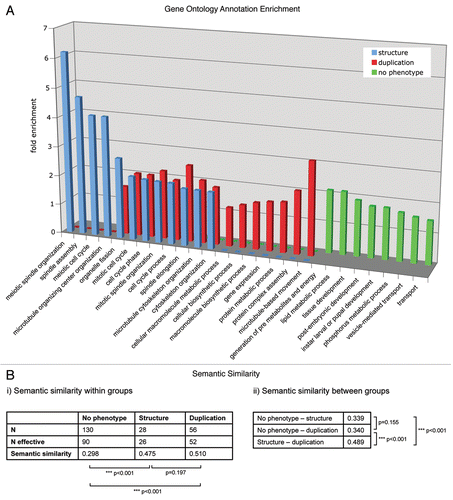
Figure 2 Comparative GO term enrichment analysis reveals functional homology between the Drosophila and human centrosome proteome. GO term enrichment in the category biological process level 3 was analyzed for all proteins classified as human centrosomal, centrosome candidate and centrosome novelCitation5 using the DAVID bioinformatics tool and the whole human genome as the background. The Drosophila centrosome proteome was analyzed accordingly and statistically significantly enriched terms (p value < 0.01) were compared between the two datasets. Less informative terms not directly related to centrosome structure or function were merged and labeled with the term of the next higher level of the GO hierarchy (italic).
