Figures & data
Figure 1 ER-mitochondria juxtaposition and high Ca2+ microdomains on the cytosolic surface of OMM in SH-SY5Y cells. Cells were transfected with cDNAs coding for a FAD-PS2 mutant (PS2-T122R, B) or void vector (Control, A), together with those coding for a red mitochondrial and a green ER markers (mit-RFP and ER-GFP, respectively; upper cell in each part), in order to visualize ER-mitochondria juxtaposition sites (yellow pixels); increased number of interactions is observed in FAD-PS2 expressing cells. Alternatively, cells were transfected with cDNAs coding for a FAD-PS2 mutant (PS2-T122R, B) or void vector (Control, A) and the cameleon Ca2+ probe N33-D1cpv, targeted to the OMM,Citation15 and then analyzed to detect the generation of high Ca2+ microdomains on the cytosolic surface of OMM (yellow-to-red spikes) upon ER Ca2+ release induced by bradykinin (lower cell in each part). At a similar average cytosolic Ca2+ rise, higher and more abundant Ca2+ microdomains are generated close to the OMM in the FAD-PS2 expressing cell compared to the control one. See reference Citation13 for details.
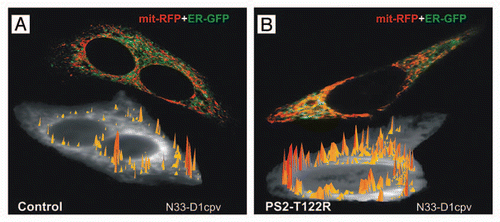
Figure 2 Model depicting the possible effects of FAD-PS2 mutations on ER-mitochondria interactions and Ca2+ cross-talk. In physiological condition (A), mitochondria receive constant Ca2+ signals from ER that ensure their proper activity.Citation30 Expression of FAD-PS2 mutants (B and C) decreases [Ca2+]ERCitation27,Citation28 and recruits mitochondria closer to ER Ca2+ releasing sites.Citation13 As a consequence, closer ER-mitochondria juxtaposition may compensate [Ca2+]ER reduction allowing mitochondria to receive appropriate Ca2+ signals that drive AT P production (B); alternatively, closer ER-mitochondria juxtaposition may expose mitochondria to excessive Ca2+ stimulation, triggering the apoptotic cascadeCitation29 by mitochondria Ca2+ overload, outer membrane permeabilization and release of pro-apoptotic factors (C).
![Figure 2 Model depicting the possible effects of FAD-PS2 mutations on ER-mitochondria interactions and Ca2+ cross-talk. In physiological condition (A), mitochondria receive constant Ca2+ signals from ER that ensure their proper activity.Citation30 Expression of FAD-PS2 mutants (B and C) decreases [Ca2+]ERCitation27,Citation28 and recruits mitochondria closer to ER Ca2+ releasing sites.Citation13 As a consequence, closer ER-mitochondria juxtaposition may compensate [Ca2+]ER reduction allowing mitochondria to receive appropriate Ca2+ signals that drive AT P production (B); alternatively, closer ER-mitochondria juxtaposition may expose mitochondria to excessive Ca2+ stimulation, triggering the apoptotic cascadeCitation29 by mitochondria Ca2+ overload, outer membrane permeabilization and release of pro-apoptotic factors (C).](/cms/asset/976cce8d-e08c-445e-97b2-896765ab419f/kcib_a_10915160_f0002.gif)