Figures & data
Figure 2. Histochemical gus assay showing gene expression (A) in transgenic plantlets while no expression can be detected in non-transgenic plant (C). (B) In transgenic T 1 seeds and spikes (B and C) while no expression can be detected in non-transgenic seeds and spikes (A).
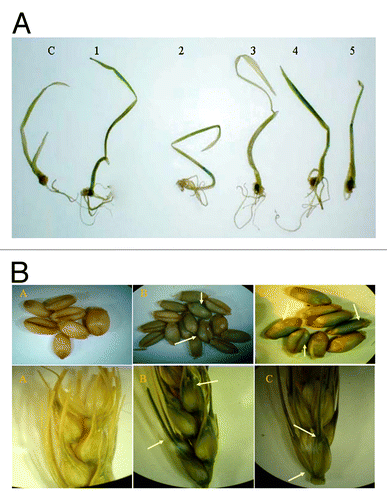
Figure 1.Effect of co-cultivation time on transient GUS gene expression in wheat cultivar Gemmeiza 9.
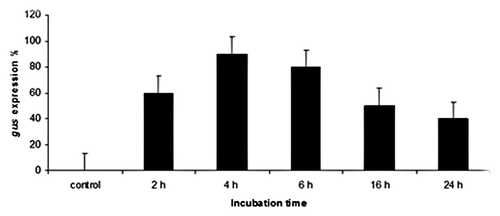
Table 1. Segregation analysis of npt-II gene among T2 progeny of transgenic events as revealed by PCR analysis.
Figure 3. Recovery of fertile transgenic wheat plants expressing the AtNHX1 gene. (A) and (B) mature seeds, (C) embryonic excise, (D) inoculation step, (E-F) germination of transgenic plant, (G-I) acclimatization and maturation.

Figure 4. PCR analysis indicating the transformation status of wheat genome with the AtNHX1 gene. A and B PCR with AtNHX1 gene (500 bp) and npt-II (250bp), respectively. M, 100 bp DNA ladder; P, plasmid; C, control.
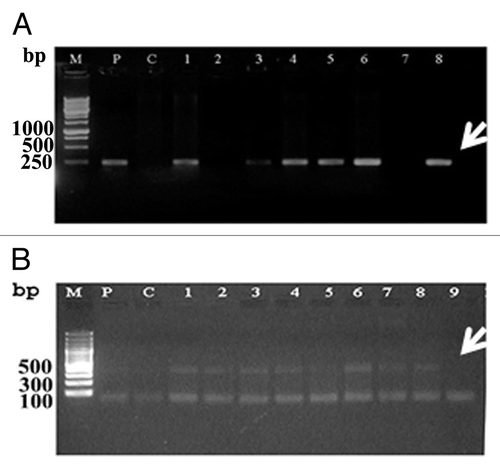
Table 2. Transformation percentages of two hexaploid wheat cultivars with ATNHX1.
Figure 5. Dot blot analysis with AtNHX1 gene specific probe (A). Northern blot analysis confirming AtNHX1 gene expression in three transgenic lines of wheat. WT, non transgenic and T1-T3: transgenic lines (B).
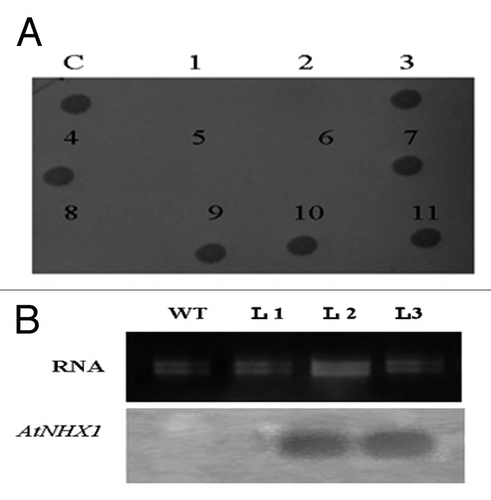
Figure 6. The effect of salt stress on growth rate of the wild type (WT) (upper panel) and AtNHX1 expressing wheat lines (lower panel).

Figure 7. Plant dry weight of two wheat transgenic lines compared with the control (non-transgenic) under different NaCl concentrations. Data are averages of three replication ± SE
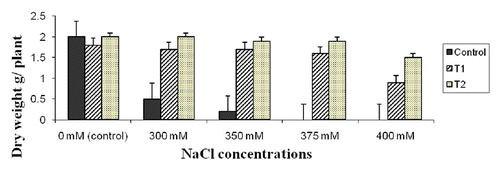
Table 3. Sodium, potassium and chloride concentrations in leaves of the transgenic wheat lines.
Table 4. The pedigrees of the two wheat cultivars used.
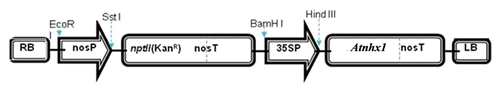
Figure 8. Schematic representation of the transformation vector pBI-121 containing the ATNHX1 under the genetic control of 35S promoter.