Figures & data
Figure 1. High throughput compound screening protocol for E. histolytica and G. lamblia trophozoites in 96-well microtiter plate.
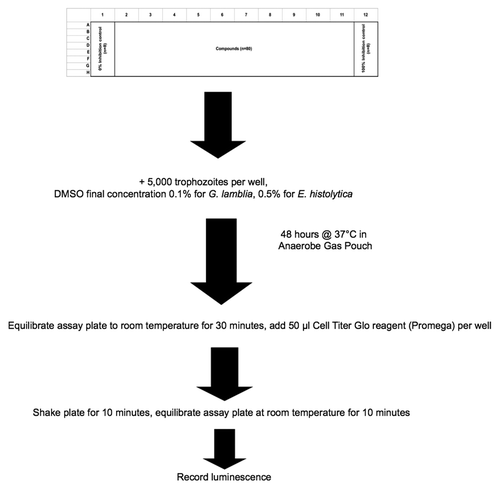
Figure 2. In vitro efficacy of different concentrations of aurnofin on the growth of C. parvum, as determined by real-time PCR. Bars represent standard errors of the means derived from three replicates.
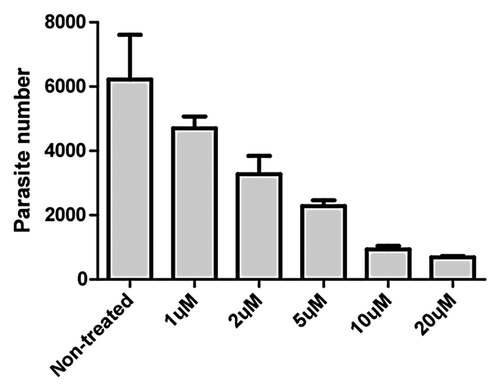
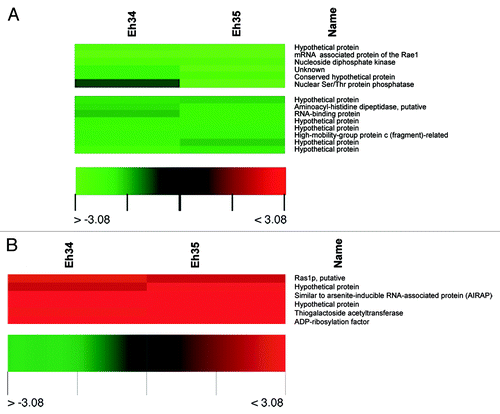
Figure 3. Clustered display of data from a portion of E. histolytica microarray, indicating downregulated (A) and upregulated transcripts (B) due to auranofin treatment. Microarray hybridizations (A and B) were performed using RNA from DMSO-treated E. histolytica and 1 µM auranofin-treated E. histolytica. Data from related genes in two separate hybridizations (Eh34 and Eh35) were analyzed by use of Acuity software.Citation22