Figures & data
Figure 1. Phylogenetic analysis and consensus engineering of CMV immunogens. Phylogenetic trees of selected CMV immunogens are shown. The significance of the unrooted phylogenetic trees was verified by bootstrap analysis and significant support values (≥ 80%; 1,000 bootstrap replicates) are indicated by asterisks at major nodes. Previously reported genotypes are illustrated (white) and reference strains are indicated; AD - AD169, DV - Davis, JH - JHC, JP - JP, ML - Merlin, TO - Toledo, TN - Towne, VR - VR1814. CMV vaccine immunogens as derived by alignment of clinical strain sequence data are displayed (Vac) and scale bars signify distance of amino acids per site.
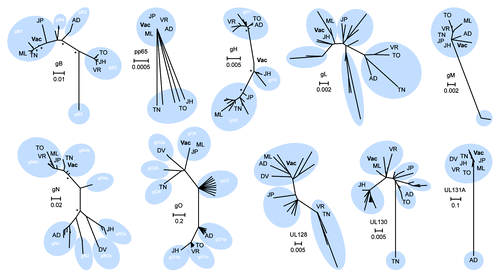
Figure 2. Construction of consensus CMV immunogens and structurally-relevant poly-proteins. Cartoon of consensus CMV immunogens. Poly-proteins are displayed and express multiple structurally-relevant proteins (labeled in white) that are separated by a furin cleavage site (*) for host cell post-translational modification. Amino acid length is defined. All proteins were subsequently genetically-optimized for expression in humans, commercially synthesized, and then subcloned into a modified pVAX mammalian expression vector.
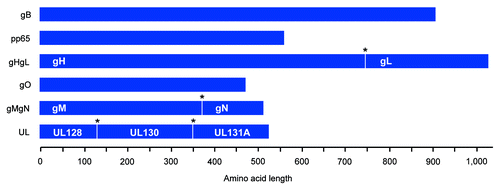
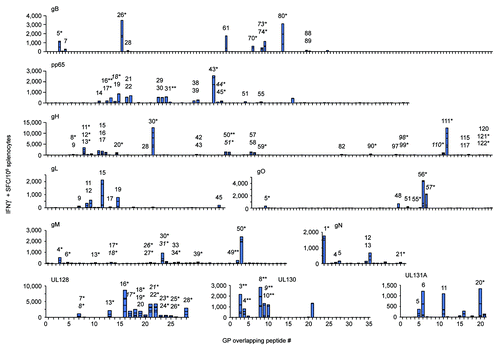
Figure 3. E-DNA vaccination was highly T cell immunogenic. Mice (n = 5/group) were immunized twice with EP delivery of the indicated CMV vaccine plasmid and IFNγ responses were measured by modified IFNγ ELISPOT. Splenocytes were incubated in the presence of individual peptides spanning each consensus CMV immunogen and results are shown in stacked bar graphs (summarized in ). Peptides eliciting CD4-restriced IFNγ responses, CD8-restricted (*), and dual CD4/CD8-restricted responses (**), as determined by FACS analysis, are individually numbered and displayed. Putative shared epitopes for contiguous positive peptide responses are italicized.