Figures & data
Figure 1. IgG and IgG-RNase constructs. (A) Schematic illustrations of IgG and IgG-RNase, as well as (B) the corresponding gene expression cassettes. In addition to IgG-RNase containing wild type human pancreatic RNase (RNase), a control construct is shown containing enzymatically inactive RNase variant RNase(H12A, H119A). The illustrations are not drawn to scale. attB1–2, BP recombination cloning sites; BGH Poly A, bovine growth hormone poly-adenylation signal; CMV, cytomegalovirus; RNase, human pancreatic ribonuclease; V, C, variable and constant regions of light (L) and heavy (H) IgG chain)
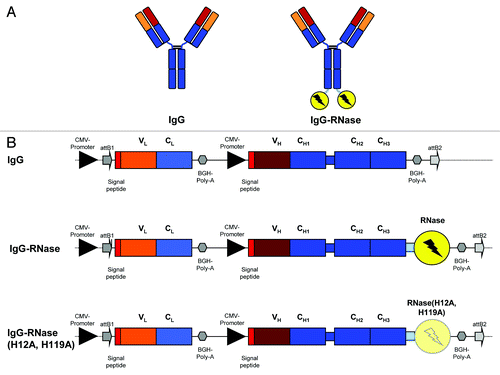
Table 1. Overview of the human IgGs and RNases used in this study
Figure 2. SDS-PAGE of purified MN-IgGs and IgG-RNase constructs. A total of 1 µg purified protein per lane was tested by SDS-PAGE under non-reducing (lane 1–4) and reducing (lane 5–8) conditions followed by Coomassie staining. Lane 1 and 5: MN-IgG; lane 2 and 6: MN-IgG-RNase; lane 3 and 7: MN-IgG-RNase(H12A, H119A); lane 4 and 8: CTX-IgG-RNase; M: protein standard All Blue (BioRad)
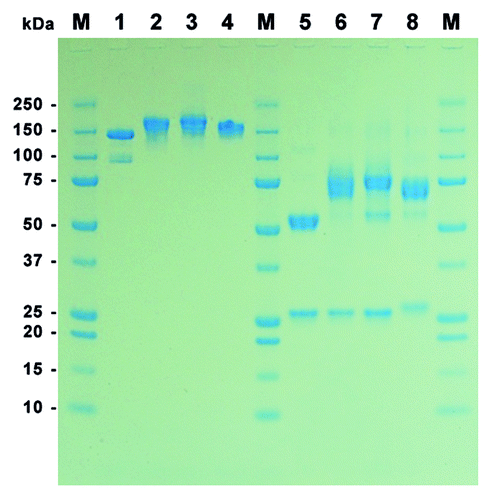
Figure 3. Analytical SEC of IgG and IgG-RNases. (A) MN-IgG, (B) MN-IgG-RNase, (C) MN-IgG-RNase(H12A, H119A) and (D) CTX-IgG-RNase. Base line is shown as gray dotted line.
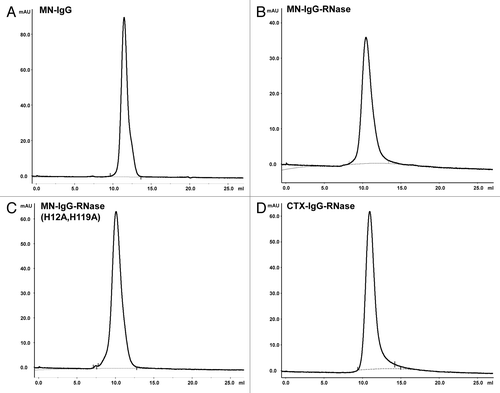
Table 2. Binding kinetic parameters from SPR calculated by 1:1 Langmuir binding model
Table 3. Catalytic efficiency of purified immunoRNases and control constructs
Figure 4. RNase activity in the presence of human placental RNase inhibitor. RNase activity of (A) MN-IgG-RNase and (B) CTX-IgG-RNase was measured in the presence of RNase inhibitor. 10−10 M RNase incubated with up to 50-fold molar excess of RI (RNasin). (C) Bovine RNase was tested as control.
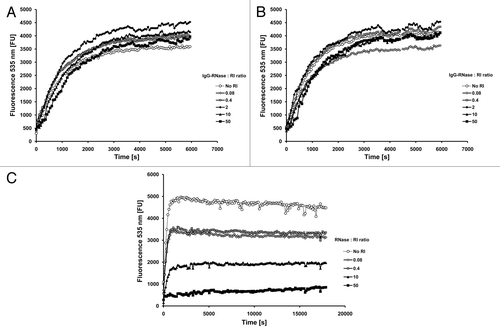
Figure 5. Stability tests. (A) IgG and IgG-RNase were incubated in 50% mouse serum at 37 °C for up to 24 h followed by testing of binding to the corresponding antigen by ELISA. Detection was done with an anti-human IgG-Fc specific secondary antibody HRP conjugate. (B) MN-IgG-RNase and (C) CTX-IgG-RNase were also tested by incubation in 50% human or mouse serum at 37 °C for up to 7 d followed by a novel immunoassay combining ELISA with an RNase activity assay. BSA was used as negative control antigen. All binding and activity data were normalized to values measured with samples freshly thawed (time 0 = 100%).
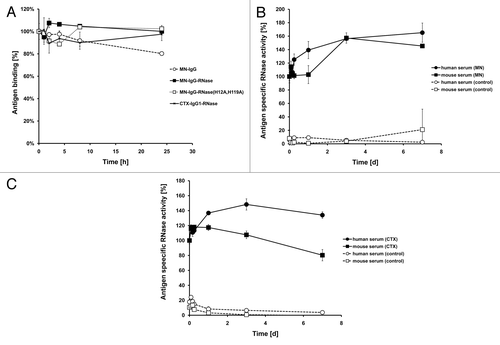
Table 4. Thermostability of the IgG-RNases
Figure 6. Internalization of fluorescently labeled MN-specific IgG-RNases. IgGs, IgG-RNases and control constructs were chemically conjugated with CypHer 5E and incubated for up to 24 h on MIAPaCa-MN+ cell overexpressing MN antigen. CTX-IgG-RNase was used as control. (A) Fluorescence microscopy was performed after different time points, images after 3 and 24 h are shown as examples. Hoechst 33342 was used to counter stain for nuclei. (B) Internalization was quantified by counting of red fluorescent granules per cell for up to 24 h.
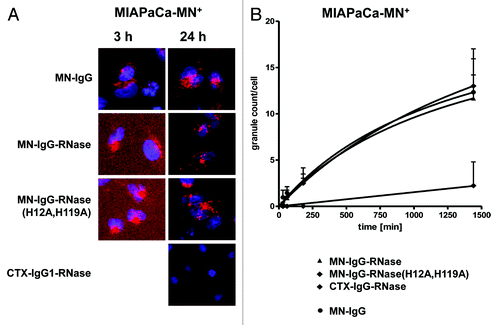
Figure 7. Growth inhibition of MN+ overexpressing tumor cell lines. MN+ overexpressing MIAPaCa 2 cells were incubated (A) with MN-IgG-RNase containing catalytic active human pancreatic RNase. MN-IgG-RNase(H12A, H119A) with a catalytic inactive RNase, and CTX-IgG-RNase, were used as negative controls, whereas Onconase was used as non-targeted positive control. B) Additionally, MN+ overexpressing MIAPaCa 2 were also incubated with (B) MN-IgG based immunoRNase constructs fused with RI evasive human pancreatic RNase variants (Jo2007, QBI-119) as well as MN-IgG-RNase and MN-IgG-QBI119 constructs containing other linker sequences (GGFKGG, GGGGGG, GGFLGG, GGAANG, and GALALAG), which are putatively cleavable in endosomes. Free QBI119 and Onconase were also tested.
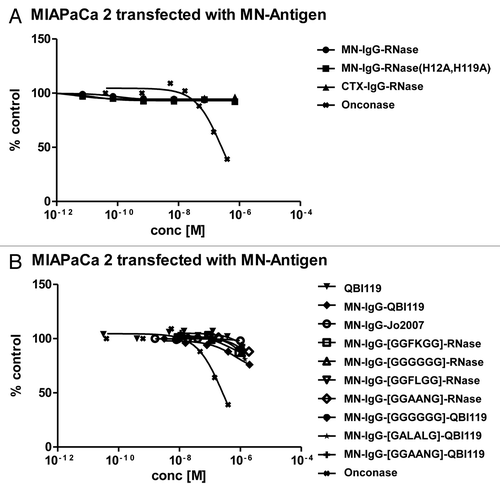
Figure 8. Internalization of fluorescently labeled X-antigen specific IgG-RNases. IgGs, IgG-RNases and control constructs were chemically conjugated with CypHer 5E and incubated for up to 24 h on A549-X+ cell or MCF7 cells which either overexpress or endogenously express X-antigen, respectively. CTX-IgG-RNase was used as control. (A) Fluorescence microscopy was performed at different time points, images after 3 and 24 h are shown. Hoechst 33342 was used as counter stain for nuclei. (B) Internalization was quantified by counting of red fluorescent granules per cell for up to 24 h.
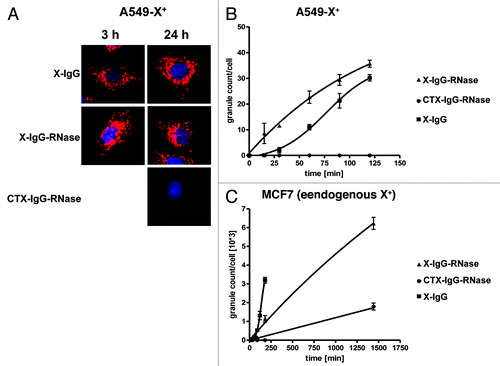
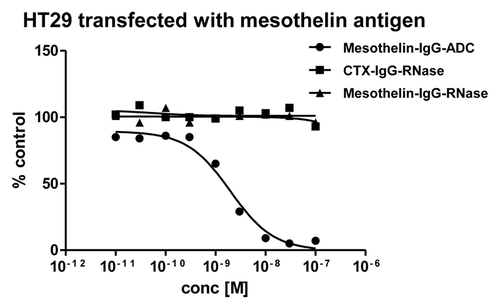
Figure 9. Growth inhibition of mesothelin-antigen expressing tumor cell lines. Mesothelin-antigen stably overexpressing HT29 cells were incubated with mesothelin-IgG based immunoRNase fusion protein. CTX-IgG-RNase was used as negative control. Mesothelin-IgG conjugated to a maytansinoid-based toxophore was used as positive control.