Figures & data
Figure 1 Key structures of the mAb probed by fluorescent dyes. N and U are native and unfolded monomers, respectively. “n” reactive monomers form aggregates.
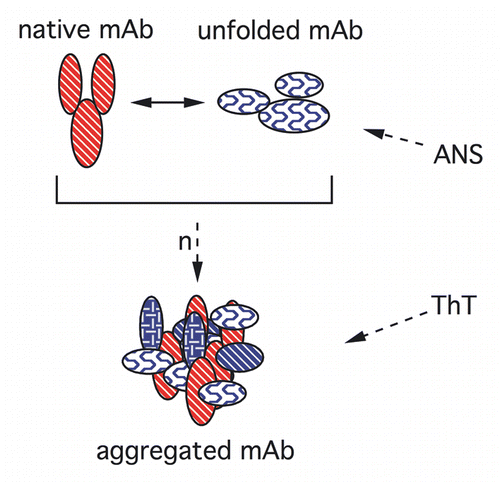
Figure 2 Binding of ANS to the mAb. (A) ANS binding equilibrium to native mAb with increasing dye concentration. Inset: Double reciprocal representation.Citation20 (B) Temperature-dependence of mAb unfolding studied with ANS binding. Dye binding rates were determined from these data as described in the text and shown in .
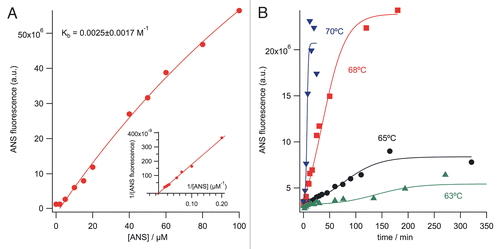
Figure 3 Arrhenius plots for unfolding and aggregate formation of the mAb as a function of temperature (63–70°C). Rate constants were calculated with the first-order reaction for both of the proteins' unfolding and aggregation with the empirical sigmoid fit function. SEC-HPLC results (55–68°C) were calculated using the second-order reaction rate. Data fit was performed using Arrhenius equation (ln kobs = ln A − Ea/RT). The activation energies for the mAb unfolding and aggregation from the ANS and ThT binding and SEC-HPLC experiments are 610, 544 and 549 kJ/mol, respectively.
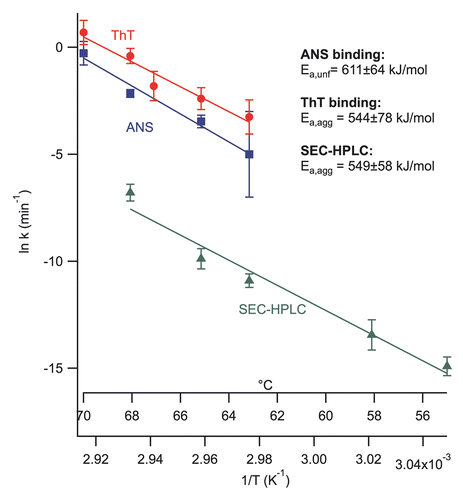
Table 1 Observed unfolding and aggregation rate constants of the mAb by the dye-binding studies