Figures & data
Figure 1 (See previous page). A model for focus formation at a double-strand break. After a DSB is induced (A), the MRN complex quickly relocalizes to the break site, bringing ATM along through an interaction with a C-terminal motif in NBS1 (B). ATM phosphorylates adjacent histone H2AX molecules, tagging nearby chromatin with γ-H2AX (C). MDC1 binds γ-H2AX through its tBRCT domain and recruits additional ATM molecules, both directly via its FHA domain, as well as indirectly via an interaction with NBS1 (D). This allows ATM to phosphorylate more distant histone H2AX molecules, thus spreading the initial signal and amplifying it. MDC1 also recruits the E3 Ub ligase RNF8, which attaches Ub molecules onto histones H2A and H2AX (D). RNF168 binds ubiquitinated histones H2A/X and catalyzes the formation of poly-Ub chains on these histones (E). These events are required for the subsequent arrival of important DDR proteins such as 53BP1 and the BRCA1-Abraxas-RAP80 complex. The events described in (D and E) occur on both sides of the break but only one side of the break is depicted for the sake of simplicity.
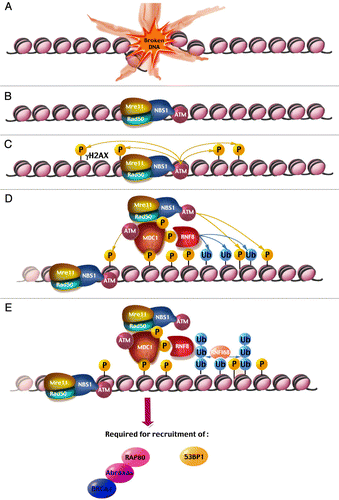
Figure 2 A schematic representation of human MDC1. All features are drawn to scale. Above each feature appears its name with relevant post-translational modifications and below are proteins that interact with it. The fourth TQXF repeat (depicted darker) is not as conserved as the first three. Also, the first three and last three PST repeats (light orange) show weak homology to the central 13 repeats (dark orange). See main text for more details.
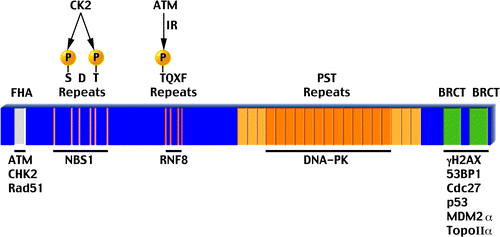
Table 1 A summary of all currently known MDC1 interactions