Figures & data
Figure 1. (A) Coomassie-stained SDS-PAGE of the SHaPrP27–30 sample subjected to SAXS analysis. (B) After SDS-PAGE, a sample was electroblotted onto a PVDF membrane and blotted with antibody 3F4. (C) Negative stain TEM of the same sample.
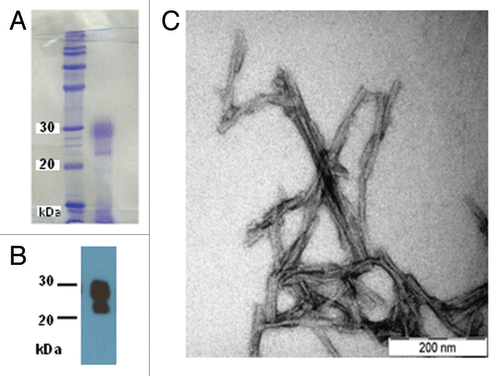
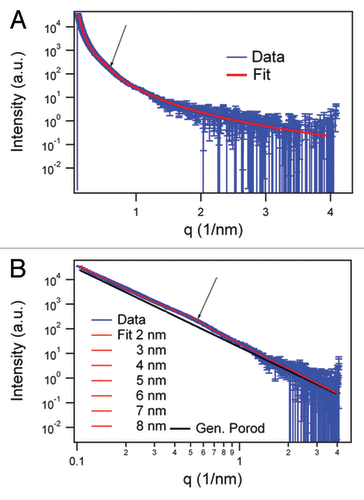
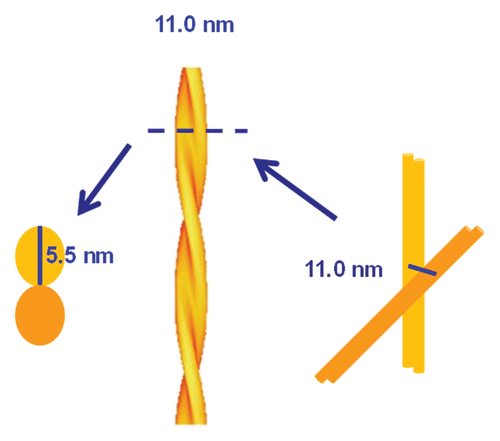
Figure 2. SAXS data in log-linear (A) representations and fit results of the infinite cylinder model (R = 5 nm) with hard-sphere structure factor and generalized Porod slope of SHaPrP27–30 (1 mg/mL). In the log-log representation of the SAXS data. (B) fit results varying the radius of the cylinders between 2 and 8 nm are shown together with the generalized Porod contribution (black). The arrow in both graphs indicates the hump of the structure factor corresponding to 11.0 ± 0.2 nm.