Figures & data
Figure 1 Domain structures of the Type I Hsp40 Ydj1 and Type II Hsp40 Sis1 (A). Both Hsp40s possess N-terminal J-domains and adjacent G/F-rich regions. Ydj1 also contains a zinc finger-like region with two zinc-binding domains (denoted as I and II). Sis1 contains a G/M-rich region in addition to the G/F-rich region. C-terminal domains (CTD) in these Hsp40s share limited homology although both possess hydrophobic polypeptide-binding pockets (PBP) and dimerization domains (DD). Furthermore, Ydj1 is farnesylated at a C-terminal CaaX motif. (B) Quaternary structures of Ydj1 and Sis1 homodimers (for details see ref. Citation42). The polypeptide-binding pockets of each Hsp40 are noted by a small grove in the C-terminus. While the J-domains are highly conserved between Ydj1 and Sis1, the orientations of these domains with respect to the polypeptide-binding pocket are very different.
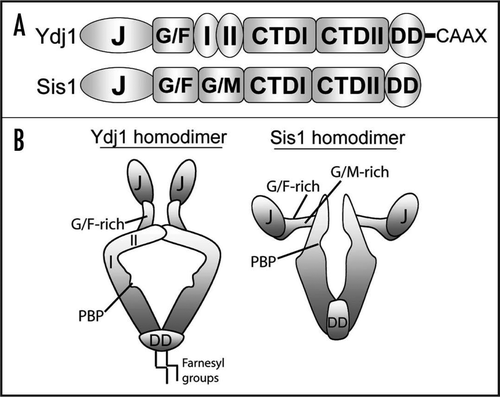
Figure 2 Domain structure of Rnq1 from S. cerevisiae (A). Rnq1 contains a N-terminal non-prion domain (aa1–152) and a C-terminal, Gln/Asn-rich prion domain (aa153–405). Sis1 binds a short, hydrophobic motif in the non-prion domain while Ydj1 binds numerous motifs in prion domain. (B) Model for Hsp40 action in [RNQ+] prion assembly pathway. Native Rnq1 is converted to the [RNQ+] prion state and assembles into large, [RNQ+] prion aggregates that are sheared to generate heritable [RNQ+] prion seeds. Sis1 might facilitate shearing to maintain a pool of [RNQ+] prion seeds thereby propagating the [RNQ+] prion state. In addition Sis1 might promote the elongation of [RNQ+] prion particles. In contrast, Ydj1 might antagonize [RNQ+] prion assembly through binding an early [RNQ+] prion conformer thus hindering [RNQ+] assembly or by accelerating the disassembly of large, [RNQ+] prions such that Rnq1 is eventually converted to its native state.
![Figure 2 Domain structure of Rnq1 from S. cerevisiae (A). Rnq1 contains a N-terminal non-prion domain (aa1–152) and a C-terminal, Gln/Asn-rich prion domain (aa153–405). Sis1 binds a short, hydrophobic motif in the non-prion domain while Ydj1 binds numerous motifs in prion domain. (B) Model for Hsp40 action in [RNQ+] prion assembly pathway. Native Rnq1 is converted to the [RNQ+] prion state and assembles into large, [RNQ+] prion aggregates that are sheared to generate heritable [RNQ+] prion seeds. Sis1 might facilitate shearing to maintain a pool of [RNQ+] prion seeds thereby propagating the [RNQ+] prion state. In addition Sis1 might promote the elongation of [RNQ+] prion particles. In contrast, Ydj1 might antagonize [RNQ+] prion assembly through binding an early [RNQ+] prion conformer thus hindering [RNQ+] assembly or by accelerating the disassembly of large, [RNQ+] prions such that Rnq1 is eventually converted to its native state.](/cms/asset/8b8e8f2c-2e47-4653-9e06-7b91374fc3f4/kprn_a_10909062_f0002.gif)