Figures & data
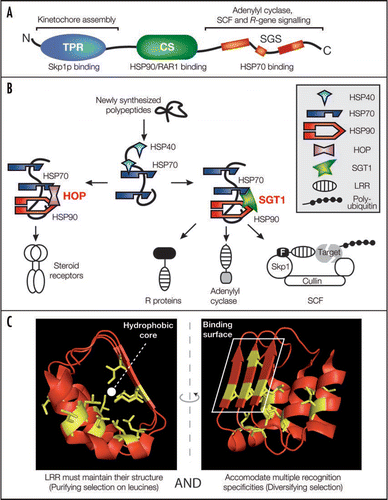
Figure 1 Model for SGT1/chaperone complex functions in the folding of LRR-containing proteins. (A) The structural domains of SGT1, their sites of action (above) and respective binding partners (below) are shown. N- and C-termini are indicated. TPR, tetratricopeptide repeat; CS, CHORD/SGT1; SGS, SGT1-specific. (B) Conceptual analogy between steroid receptor folding by the HOP/chaperone machinery and LRR protein folding by the SGT1/chaperone machinery. LRR motifs are overrepresented in processes requiring SGT1 such as plant immune receptor signalling, yeast adenylyl cyclase activity and plant or yeast SCF (Skp1/Cullin/F-box) E3 ubiquitin ligase activities. (C) Opposite forces drive LRR evolution. Structure of LRRs 16 to 18 of the F-box auxin receptor TIR1 is displayed as an illustration of the LRR folds.Citation30 Leucine/isoleucine residues (side chain displayed in yellow) are under strong purifying selection and build the hydrophobic LRR backbone (Left). By contrast, solvent-exposed residues of the β-strands define a polymorphic and hydrophilic binding surface conferring substrate specificity to the LRR (Right) and are often under diversifying selection.