Abstract
Correction to:Emerging Microbes & Infections (2018) 7,43 10.1038/s41426-018-0045-x; published online 29 March 2018
The original article can be found online at 10.1038/s41426-018-0045-x.
Correction to:Emerging Microbes & Infections (2018) 7,43 10.1038/s41426-018-0045-x; published online 29 March 2018
Upon publication of the original articleCitation1 the authors were contacted by Dr. Trevor Bedford who has provided invaluable methods to improve the temporal analysis work presented in this paper. Due to conversations with Dr. Bedford the authors improved the calculation of the time to most recent common ancestor (TMRCA) and the clock rate of evolution within Candida auris, to take into account the whole genome, not just the variable sites.
This author correction provides an updated Fig. and an updated paragraph related to this new information. The updated information has been indicated in bold in the below paragraphs.
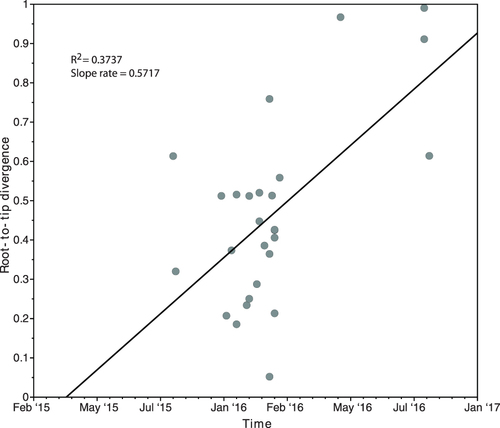
Fitting root-to-tip regression showed there was a linear relationship between sampling time (measured in days) and the expected number of nucleotide substitutions along the tree, demonstrating clock-like evolution across the timescale of the outbreak (Fig. ). There was a good association between genetic distances and sampling dates (R2 = 0.3737), showing that the dataset has a reasonable temporal signal for further molecular clock analysis. Given 0.5757 substitutions per site per year in variable sites, which represent 0.01% of total sites across the genomes, the evolutionary rate of C. auris nuclear DNA equates to 5.7e-5substitutions per site per year, which is comparable to nuclear DNA of other fungal species, such as Schizosaccharomyces pombe beer strains (3.0e-3 28) and Saccharomyces cerevisiae (5.7e-3 29). The time to the most recent common ancestor (TMRCA) was estimated to be late March 2015, weeks prior to the first patient identified with a C. auris infection.
Future alignments will require clade-specific references due to the large evolutionary distances between the South American/African and Indian/Pakistani clades. Although the mode of introduction into the UK is unknown, temporal analysis of the outbreak isolates placed the most recent common ancestor as late March 2015, which correlates closely to the first confirmed infection within the hospital weeks later, suggesting a recent introduction into the UK.