ABSTRACT
Marine benthic proliferations are increasing in occurrence, range and duration by way of anthropogenic nutrient loading and climate change. The spread of these cyanobacteria within a variety of regions has made it fundamental to recognize the diversity and the species involved in these blooms. To expand knowledge on marine cyanobacterial diversity and reveal their phylogenetic relationships, sampling and isolation of benthic proliferative events from underexplored regions were conducted in the USA (Florida) and France (Loire-Atlantique). Cyanobacteria were described with the polyphasic approach using morphology, 16S rRNA gene phylogeny, 16S–23S rRNA ITS secondary structures and pairwise distances. Sampling of marine cyanobacteria from seagrasses, lagoons and marine coastal waters from nine Florida localities, in addition to Florida freshwater canals and French salt flats, revealed floating, epipsammic and epiphytic mats. A total of 30 cyanobacteria were isolated of which 21 represented two novel genera, Sirenicapillaria and Tigrinifilum. Sirenicapillaria is a genus that is found in massive benthic blooms throughout the western and southern Florida Coast.
INTRODUCTION
Coastal marine areas are notably changing through anthropogenic nutrient loading, temperature fluctuations due to climate change, and acidification (Pearce & Feng Citation2013; Di Lorenzo & Mantua Citation2016; Oliver et al. Citation2017). With warming weather and nutrients in favour, benthic cyanobacterial mats (BCMs) can form dense proliferations over shallow and deeper regions that were once dominated by seagrass beds or hard-bodied corals, respectively (Paerl & Paul Citation2012; Gurgel et al. Citation2020; Urrutia-Cordero et al. Citation2020; Benny et al. Citation2021). These dense BCMs are increasing and becoming more prevalent (Coleman et al. Citation2020; Wood et al. Citation2020; Ford et al. Citation2021) where they can dominate reefs and coastal and littoral regions (Paul et al. Citation2005; Sneed et al. Citation2017; Ford et al. Citation2018; Lydon et al. Citation2020; Berthold et al. Citation2021; FDEP Citation2019–2021). BCMs can also be a nuisance due to producing toxins or unknown secondary compounds, emitting noxious odours, decreasing dissolved oxygen when decomposing leading to hypoxia and fish kills, or harbouring microorganisms that can cause skin irritation (Ford et al. Citation2018; Reilly Citation2020; Wood et al. Citation2020). With the expansion of BCMs causing negative ecological effects, it is necessary to recognize and understand their biological and chemical diversity (Sharp et al. Citation2009; Engene et al. Citation2011; Sendall & McGreggor Citation2018).
Although benthic marine cyanobacterial proliferations (i.e. benthic blooms) are frequently observed, the extent of their spatial and temporal occurrence as well as the species involved remain largely unexplored. To further compound the issues, BCM proliferations covering seagrasses, corals and the benthos are often misidentified and reported as ‘Lyngbya’, ‘Lyngbya majuscula’, ‘Microseira wollei’, ‘Moorena producens’, or ‘Dapis’, an indication for the need of more taxonomic studies (Engene et al. Citation2012, Citation2013, Citation2018; FDEP 2021). With proper cyanobacterial descriptions and taxonomic designations, our understanding of cyanobacteria that bloom on the coast is strengthened, which is also important for public outreach (Collins Citation2021).
Accurate identification of cyanobacteria within mats can be challenging as BCMs are usually entangled masses of several cyanobacterial species (Biessy et al. Citation2021; Berthold et al. Citation2021). Since marine BCMs are highly diverse, integrative molecular techniques are often necessary for proper specific and generic delimitations (Caires et al. Citation2018a; Lefler et al. Citation2021). The combination of molecular and ultrastructural approaches together with ecological and morphological analyses is necessary to unravel cyanobacterial diversity (Komárek Citation2016). This polyphasic method is especially important in delimiting simple homocytous trichal cyanobacteria in the order Oscillatoriales, such as Lyngbya, Oscillatoria and Phormidium, which have few morphological features and still include polyphyletic assemblages of species (Komárek Citation2016; Mühlsteinová et al. Citation2018). In addition to describing cyanobacteria on a generic and specific level, family level taxonomy is equally necessary. Recent genetic delimitation of Oscillatoria princeps Vaucher ex Gomont by Mühlsteinová et al. (Citation2018) provided a reliable taxonomic reference point for the genus Oscillatoria and thus the family Oscillatoriaceae, and revealed a need for the revision of the genera that have been traditionally classified as Oscillatoriaceae, because these are polyphyletic as currently circumscribed (Suda et al. Citation2013; Berthold et al. Citation2021; Zimba et al. Citation2021).
To investigate cyanobacteria that form BCMs, benthic proliferations from coastal western Florida, South Florida and western France were studied. From the sampled cyanobacterial mats, we propose two novel genera: Sirenicapillaria gen. nov., represented by three new species (S. glauca sp. nov., S. rigida sp. nov. and S. stauglerae sp. nov.), and Tigrinifilum gen. nov., represented by two species (T. floridanum sp. nov. and T. guerandense sp. nov.). Two new species of Capilliphycus, C. guerandensis sp. nov. and C. flaviceps sp. nov. are also described.
MATERIAL AND METHODS
Cyanobacteria sampling, isolation and cultivation
Cyanobacterial benthic mats which possessed filamentous ‘Lyngbya-like’ taxa were sampled from various marine locations around Florida (USA), including the west (Gulf) coast near Lemon Bay, the east (Atlantic) coast in Biscayne Bay, and the Florida Keys. A total of 10 sampling events were conducted in Florida coastal waters between August 2017 and July 2020 (Table S1). Samples were collected mostly from epilithic or epipsammic mats, dislodged floating mats and among seagrass beds. Cyanobacterial mats and dried mat material found in hypersaline flats from western France (Guérande) and freshwater canals in Florida were also included in this study for robust taxon sampling of BCMs.
Cyanobacteria were isolated with traditional microbiological methods using solidified (15 g l–1 agar) sterile source water and micromanipulation of single filaments onto solid BG-11 media with added aquarium salts (35 g l–1) (SWBG-11). Cyanobacterial isolates were treated with cycloheximide to remove contaminating eukaryotes (355 mM; SIGMA) and cultured in liquid SWBG-11 (35 ppt) at 25°C under 12:12 h light cycles. Live cultures are maintained in the BLCC (University of Florida, Fort Lauderdale Research and Education Center, Davie, Florida, USA) and the BCCM/ULC (University of Liège, Liège, Belgium); both dried and preserved (4% formaldehyde) type material for each novel taxon is deposited in the US National Herbarium (National Museum of Natural History, Smithsonian Institution, Washington, DC, USA).
Morphological and phylogenetic analyses
Images of cyanobacterial thalli were taken using a DSLR camera. Morphometric data were derived from images captured on a DIC (differential interference contrast) compound microscope (Leica DM5500 B) and corresponding LAS X software (Life Sciences; Leica Microsystems, Wetzlar, Germany).
Prior to DNA extraction, fresh biomass (100 mg) of each unialgal culture was washed twice with sterile deionized water to remove salts. Total genomic DNA was extracted with a DNeasy UltraClean Microbial Kit (Qiagen, Hilden, Germany) and then quantified using NanoDrop Lite (Thermo Scientific). The 16S rRNA gene sequence and 16S–23S rRNA internal transcribed spacer (ITS) region were amplified by polymerase chain reaction (PCR) on a thermocycler (ProFlex PCR system; Applied Biosystems; Life Technologies) using the primers 359F and 1487R, and 1337F and 23S30R, respectively (Wilmotte et al. Citation1993; Nübel et al. Citation1997) following protocols outlined in Berthold et al. (Citation2021).
PCR products were purified with a QIAquick PCR Purification Kit (Qiagen) and were subsequently visualized on a 0.8% agarose gel. The amplified products of the 16S rRNA gene were directly sequenced; however, the 16S–23S rRNA ITS products were cloned using a TOPO TA cloning kit with TOP10 chemically competent cells (Invitrogen; Life Technologies) according to manufacturer protocols. Recombinant plasmids from several positive transformants for each sample were extracted from clonal libraries and purified (PureLink® Quick Plasmid Miniprep Kit). Sanger sequencing of both 16S rRNA PCR products and plasmid DNA was carried out by Eurofins Genomics (Kentucky, USA) using BigDye Terminator v3.1 (Applied Biosystems). Sequences are deposited in GenBank (National Center for Biotechnology Information, NCBI) under the accession numbers listed in Table S1.
The Basic Local Alignment Search Tool (BLASTN), NCBI, was used for identifying sequences of representative strains similar to the 16S rRNA gene of the cyanobacterial isolates. A total of 141 sequences were used to align with the 30 isolates from our study using MUSCLE (SeaView v4.4; Edgar Citation2004), and then annotated manually based on conserved regions, with Gloeobacter violaceus (AY485484) as an outgroup. A total of 1013 sites of the 16S rRNA gene were used in the alignment. The best fit model was assessed using jModelTest through MEGA v10.1.7 (Stecher et al. Citation2020). Phylogenetic trees were constructed using Bayesian inferencing (BI) with MrBayes v3.2.7a and maximum likelihood (ML) using RAxML v8.2.12, through the CIPRES network v3.3 and MEGA, respectively (Miller et al. Citation2010). The ML analysis was carried out using the K2 + G + I model with 1,000 bootstrap resampling replicates. The BI analysis was conducted with MrBayes v3.2.7a (Ronquist & Huelsenbeck Citation2003) using 1.5 × 106 generations, a 0.25 burnin rate and resampling every 100 generations.
Predicted secondary structures including the D1-D1’ and Box-B helices of the 16S–23S rRNA ITS region were generated using Mfold (Zuker Citation2003) with default settings and the structures were drawn using untangle with loop fix setting. Pairwise distances for the 16S rRNA gene and 16S–23S rRNA ITS sequences between isolates and representative related clades were calculated using MEGA. Mean p-distance between and within cyanobacterial families based on 16S rRNA gene sequences were also calculated using MEGA, and taxonomic family classification was based on Guiry & Guiry (Citation2021) and Hauer & Komárek (Citation2021).
RESULTS
Taxonomic treatments
Subclass Oscillatoriophycidae L. Hoffmann, Komárek & Kaštovský Order Oscillatoriales J.H. Schaffner
Sirenicapillariaceae D.E. Berthold, Lefler & Laughinghouse fam. nov.
Description
Homocytous, uniseriate, isopolar, straight to slightly curved to coiled, filamentous. Thalli planktonic and/or attached to substrata and vegetation, prostrate, erect, fascicular or not. Euryhaline. Filaments with or without sheaths. Trichomes with or without constriction at cross wall. Cell content granulated typically at cross-walls. Cells discoid, apical cells range from bluntly rounded, to conical rounded, to rounded, calyptra present or not. Reproduction by single cell release, hormogonia, and/or diagonal fragmentation. False branching is rare. Currently, this family includes the genera Affixifilum, Capilliphycus, Limnoraphis, Limnospira, Neolyngbya, Sirenicapillaria and Tigrinifilum, which differ by 16S rRNA phylogeny.
Diagnosis
Monophyletic group of marine/brackish/freshwater genera based on the 16S rRNA gene phylogeny, sister to the Laspinemataceae clade.
Comment
Genera are commonly found in marine coastal areas, with some found in inland waters. Some genera display desiccation tolerance and growth in hypersalinity.
Etymology
Name based on the genus Sirenicapillaria, a rigid marine filamentous cyanobacterium that forms nuisance benthic blooms across the Florida western and southern coasts.
Type Genus
Sirenicapillaria gen. nov.
Subclass Oscillatoriophycidae L. Hoffmann, Komárek &KaštovskýOrder Oscillatoriales J.H. Schaffner Family Sirenicapillariaceae D.E. Berthold, Lefler & Laughinghouse fam. nov.
Sirenicapillaria D.E. Berthold, Lefler & Laughinghouse gen. nov.
Description
Homocytous filamentous cyanobacteria, forming dark brown, blue green erect thallus on benthos. Long coarse filaments (c. 10 cm) with thick, sometimes facultative and lamellate hyaline sheath. Trichomes cylindrical, thicker in the centre and slowly attenuated towards apices, uniseriate, constricted at cross-walls. Cells discoid, shorter than wide. Cell content light to dark brown, greyish green and blue green. Apical cell rounded, narrowed at times, with occasional calyptra. Reproduction by release of individual cells, straight or diagonal trichome fragmentation, necridia, hormogonia. Rarely false branches.
Type Species
Sirenicapillaria stauglerae D.E. Berthold, Lefler & Laughinghouse sp. nov.
Etymology
From Latin Siren (Greek Σειρήν), a Siren, and Latin capillus, hair, in reference to the long thalli and ‘mermaid’s hair’ appearance of these marine cyanobacteria.
Sirenicapillaria glauca D.E. Berthold, Lefler & Laughinghouse sp. nov.
Figs 1–6. Images of Sirenicapillaria glauca (reference strain BLCC-M125).

Description
Thallus erect, light to dark blue green. Filaments straight and somewhat flexuous, longer than 10 cm, 18.4–25.2(–28) µm wide with thick hyaline sheath. Trichomes cylindrical, (14–)15–20(–21) µm wide, thicker in the centre and gradually attenuated towards the ends. Cells slightly constricted at cross walls. Cells discoid, wider than long, 1.3–2.8 µm long. Cell content light to dark green, blue green to greyish, with lightly pigmented cells towards apices. Apical cell rounded, at times conical, with occasional calyptra. Reproduction by straight or diagonal trichome fragmentation, hormogonia, and necridia.
Type Locality
USA. Florida: Englewood, seagrass beds within Lemon Bay Aquatic Preserve (26°54.503ʹN, 82°20.365ʹW).
Habitat
Epipsammic, epiphytic and floating on marine water surface.
Etymology
Latin adjective glaucus, -a, -um, in reference to the ‘sea green’ cell and thalli colour of this cyanobacterium.
Holotype
US 227772 (dried material in a metabolically inactive state of reference strain BLCC-M125), deposited in US National Herbarium, National Museum of Natural History, Smithsonian Institution, Washington, DC, USA.
Reference Strain
BLCC-M125.
materials analysed
BLCC-M125.
Sirenicapillaria rigida D.E. Berthold, Lefler & Laughinghouse sp. nov.
Figs 7–15. Microscope images of Sirenicapillaria rigida (reference strain BLCC-M116).
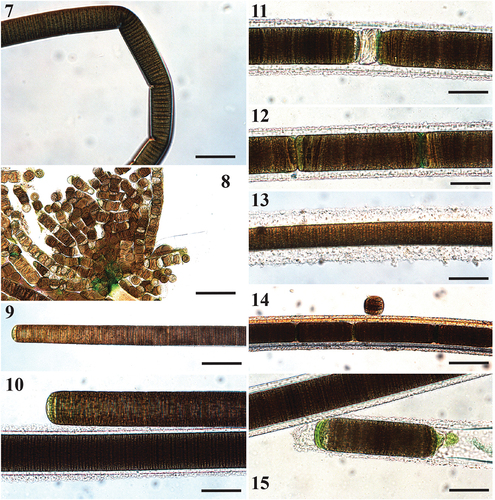

Description
Thallus dark brown, tangled, web-like. Filaments straight and rigid, 5 cm long, 28–83(–94) µm wide with thick, lamellate hyaline sheath. Trichomes cylindrical, 22.8–63.3 µm wide, thicker in the middle and gradually attenuated towards apices. Cells slightly constricted at cross walls. Cells discoid, wider than long, 1.8–3.9 µm long. Cell content light to dark brown, with pigmentless cells towards apices. Apical cell rounded, at times conical, with occasional calyptra. Reproduction by release of individual cells, straight or diagonal trichome fragmentation, necridia, hormogonia.
Comment
Individual filaments observed with naked eye and reminiscent of the freshwater Microseira wollei. Hormogonia form attached clumps on individual filaments.
Type Locality
USA. Florida: Duck Key (24°46.635ʹN, 80°54.527ʹW).
Habitat
Benthic, wrapped around seagrasses in marine coasts.
Etymology
Latin adjective rigidus, -a, -um, in reference to the physical rigidity of the filaments.
Holotype
US 227691 (dried material in a metabolically inactive state of reference strain BLCC-M116), deposited in US National Herbarium, National Museum of Natural History, Smithsonian Institution, Washington, DC, USA.
Reference Strain
BLCC-M116.
materials analysed
BLCC-M116 (US 227691), BLCC-M134 (US 227777).
Sirenicapillaria stauglerae D.E. Berthold, Lefler & Laughinghouse sp. nov.
Figs 16–26. Images of Sirenicapillaria stauglerae (reference strain BLCC-M121).
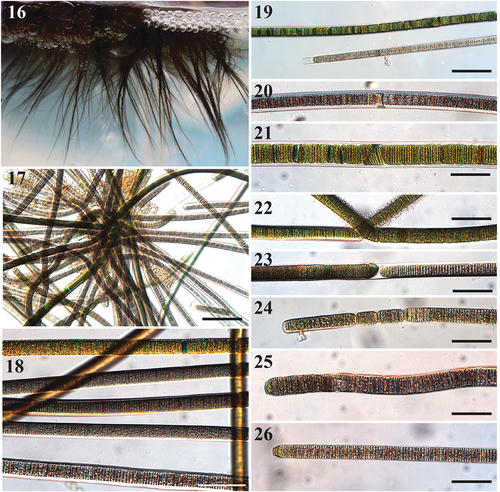

Description
Thallus erect, light to dark brown, almost black. Filaments straight and flexuous, 5 cm long, 12.5–33(–35) µm wide with thick hyaline sheath. Trichomes cylindrical, 10–26.8 µm wide, thicker in the centre and gradually attenuated towards apices. Cells slightly constricted at cross walls. Cells discoid, wider than long, 1.0–2.4 µm long. Cell content light to dark brown, grey to red, with pigmentless cells towards apices. Apical cell rounded, at times conical, with occasional calyptra. Reproduction by straight or diagonal trichome fragmentation, hormogonia and necridia. Rarely false branches.
Comment
Filament ends usually do not have thick sheaths, while mid-filaments usually have thick lamellate hyaline sheaths.
Type Locality
USA. Florida: Englewood, seagrass beds within Lemon Bay Aquatic Preserve (26°54.503ʹN, 82°20.365ʹW).
Habitat
Benthic on marine coasts.
Etymology
The epithet stauglerae is in honour of Elizabeth ‘Betty’ Staugler for her committed work in studying and surveying coastal marine habitats and the ‘Eyes on Seagrass’ Florida Sea Grant Extension programme, which first noted the emergence of this cyanobacterium blanketing areas that had previously been inhabited by seagrasses.
Holotype
US 227770 (dried material in a metabolically inactive state of reference strain BLCC-M121), deposited in US National Herbarium, National Museum of Natural History, Smithsonian Institution, Washington, DC, USA.
Reference Strain
BLCC-M121.
materials analysed
BLCC-M121, BLCC-M122, BLCC-M123, BLCC-M138.
Subclass Oscillatoriophycidae L. Hoffmann, Komárek & KaštovskýOrder Oscillatoriales J.H. SchaffnerFamily Sirenicapillariaceae D.E. Berthold, Lefler & Laughinghouse fam. nov.
Tigrinifilum D.E. Berthold, Lefler & Laughinghouse gen nov.
Description
Homocytous filamentous cyanobacteria forming solitary and free-floating, straight, wavy to slightly flexuous filaments, with or without thin hyaline sheath. Trichomes cylindrical. Cells slightly constricted between meristematic zones, highly constricted between dividing cells in older filaments. Cross-walls densely granulated and darkened except at the filament ends. Cells discoid, wider than long. Cell content light to dark green, blue green, brown, and grey. Apical cell rounded, with occasional clear calyptra. Reproduction by fragmentation or hormogonia.
Type Species
Tigrinifilum floridanum D.E. Berthold, Lefler & Laughinghouse sp. nov.
Etymology
From Latin tigrinus, striped like a tiger, and Latin filum, a thread, in reference to the resemblance of a ‘banded tiger’ characteristic of the filaments.
Comment
Species of this genus are halotolerant-halophilic and resistant to desiccation.
Tigrinifilum floridanum D.E. Berthold, Lefler & Laughinghouse sp. nov.
Figs 27–38. Microscope images of species of Tigrinifilum.
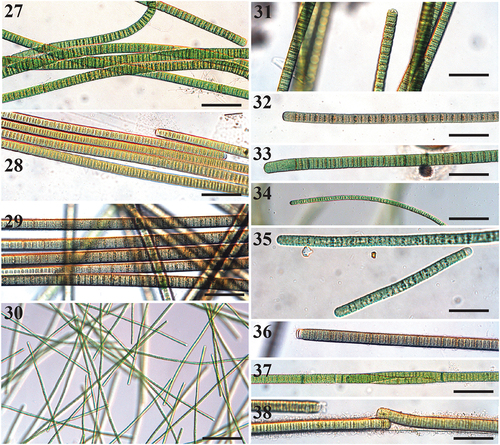

Description
Filaments solitary and free-floating, or attached to substrate. Filaments straight, wavy, to flexuous, (9–)10.2–19 µm wide with thick hyaline sheath. Trichomes cylindrical, (7–)8.0–19.2 µm wide. Cells slightly constricted between meristematic zones, highly constricted between dividing cells in older filaments. Cross-walls densely granulated and darkened. Cells discoid, wider than long, rarely isodiametric, 1.0–2.7 µm long. Cell content light to dark green, blue green, brown, and grey. Apical cell rounded, with occasional clear calyptra. Reproduction by fragmentation or hormogonia.
Type Locality
USA. Florida: Elliot Key (Biscayne Bay) (25°23.810ʹN, 80°14.043ʹW).
Habitat
Entangled within larger benthic cyanobacteria or dislodged floating mats.
Etymology
Latin floridanum, in reference to the widespread occurrence of this cyanobacterium throughout the southern Florida coast.
Holotype
US 223427 (dried material in a metabolically inactive state of reference strain BLCC-M48), deposited in US National Herbarium, National Museum of Natural History, Smithsonian Institution, Washington, DC, USA.
Reference Strain
BLCC-M48.
materials analysed
BLCC-M48, BLCC-M50, BLCC-M51, BLCC-M57, BLCC-M66, BLCC-M73, BLCC-M77, BLCC-M85, BLCC-M87, BLCC-M102, BLCC-M107, BLCC-M118, BLCC-M119, BLCC-M120.
Tigrinifilum guerandense D.E. Berthold, Lefler & Laughinghouse sp. nov.
Description
Thallus solitary, floccose, free-floating, light to dark blue green. Filaments straight, wavy to flexuous, without sheath. Trichomes cylindrical, 5.3–6.2 µm wide. Cells slightly constricted between meristematic zones, highly constricted between dividing cells in older filaments. Cross-walls densely granulated and darkened. Cells discoid, wider than long, rarely isodiametric, 1.5–2.9 µm long. Cell content light to dark green, blue green, brown and grey. Apical cell rounded. Reproduction by filament disintegration or fragmentation.
Type Locality
FRANCE. Loire-Atlantique: Guérande, Terre de Sel (47°18.938ʹN, 2°27.197ʹW).
Habitat
Entangled within cyanobacterial mats of inland salt flats.
Etymology
The epithet guerandense, is a reference to the salt flats in Guérande where the species originates.
Holotype
US 227675 (dried material in a metabolically inactive state of reference strain BLCC-M99), deposited in US National Herbarium, National Museum of Natural History, Smithsonian Institution, Washington, DC, USA.
Reference Strain
BLCC-M99.
materials analysed
BLCC-M99.
Subclass Oscillatoriophycidae L. Hoffmann, Komárek & KaštovskýOrder Oscillatoriales J.H. SchaffnerFamily Sirenicapillariaceae D.E. Berthold, Lefler & Laughinghouse fam. nov.
Capilliphycus flaviceps Lefler, D.E. Berthold & Laughinghouse sp. nov.
Figs 39–48. Microscope images of Capilliphycus. Scale bars = 50 µm.
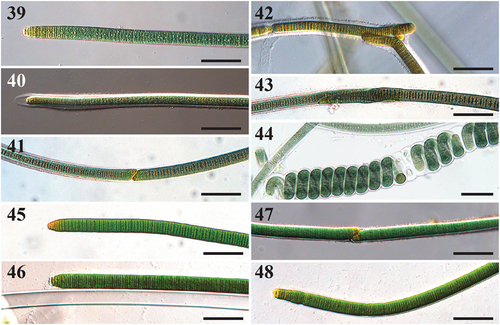
Description
Thallus amorphous and entangled, mat-like. Filaments straight, 14–18.9 µm in diameter, rarely coiled. Sheath thick, facultative, hyaline, not lamellate. Cells slightly constricted. Cross walls densely granulated. Cells discoid, wider than long, 10.7–14 µm wide, 1.4–2.4 µm long. Cell content green, green-yellow, and yellow at apices. Apices conical to rounded, without calyptra. Hormogonia formed by straight fragmentation, with or without necridic cells. Diagonal fragmentation occurs with or without necridic cells.
Type Locality
USA. Florida: Key Biscayne (25°43.588ʹN, 80°09.513ʹW).
Habitat
Found within floating marine mats of larger filamentous cyanobacteria.
Etymology
Latin adjective flavus, -a, -um, yellow, and Latin ending -ceps (caput), head, in reference to the tendency for yellowing of the apices of the trichome.
Holotype
US 227779 (dried material in a metabolically inactive state of reference strain BLCC-M137), deposited in US National Herbarium, National Museum of Natural History, Smithsonian Institution, Washington, DC, USA.
Reference Strain
BLCC-M137.
materials analysed
BLCC-M137, BLCC-M53 (US 227634).
Capilliphycus guerandensis Lefler, D.E. Berthold & Laughinghouse sp. nov.
Description
Thallus light to dark forest green, and yellow to orange when exposed to air (drying). Filaments straight to flexuous, 18.5–29.2 µm in diameter, rarely coiled. Sheath thin, facultative, hyaline, not lamellate. Cells not constricted. Cells discoid, wider than long, 15.8–24.9 µm wide, 1–2.2 µm long. Cell content green. Apices conical to rounded, attenuating, without calyptra. Apical cell yellow. Hormogonia formed by straight fragmentation, with or without necridic cells. Diagonal fragmentation occurs with or without necridic cells.
Comment
This species grows partially submerged.
Type Locality
FRANCE. Loire-Atlantique: Guérande, Terre de Sel (47°18.938ʹN, 2°27ʹ11.850”W)
habitat
Hypersaline pools within inland salt flats.
Etymology
The epithet guerandensis is a reference to the salt flats in Guérande where the species originates.
Holotype
US 227655 (dried material in a metabolically inactive state of reference strain BLCC-M76), deposited in US National Herbarium, National Museum of Natural History, Smithsonian Institution, Washington, DC, USA.
Reference Strain
BLCC-M76.
Materials Analysed
BLCC-M76, BLCC-M92 (US 227668).
Capilliphycus tropicalis T.A. Caires, Sant’Anna & J.M. Nunes
Comment
Our material fits the original description of this species in Caires et al. (Citation2018b) and clusters together with Capilliphycus tropicalis in the 16S rRNA gene sequence phylogenetic tree.
Species Features
Thallus green and entangled, mat-like. Filaments straight, 11.8–16.8 µm in diameter. Sheath hyaline. Cells light to dark green. Cells discoid, wider than long, 8.4–12 µm wide, 1–2.1 µm long. This is the first report of this species outside of Brazil.
Materials Analysed
BLCC-M106 (US 227681).
Cyanobacterial isolation and morphological characteristics
A total of 30 unicyanobacterial isolates were used in this study (Table S1). All 30 cyanobacterial strains contained discoid cells with or without a sheath. Sampling of marine cyanobacteria from seagrasses, lagoons and marine coastal waters from nine Florida localities revealed floating, epipsammic and epiphytic mats. From Florida BCMs, a total of 24 unicyanobacterial strains were isolated, of which 21 isolates represent two novel cyanobacterial genera, Sirenicapillaria and Tigrinifilum. In addition, three novel cyanobacterial isolates of Capilliphycus and Tigrinifilum from France (Guérande) are described, as well as the phylogenetic information (16S rRNA gene and 16S–23S rRNA ITS) of three isolates of Limnoraphis from freshwater canals in Florida.
Morphological analyses of the 24 Floridian marine cyanobacterial strains demonstrated cyanobacteria with cell dimensions ranging between 10.2 and 83 µm for filament width and discoid cells ranging between 5.3–63.3 and 1.0–3.9 µm for both trichome cell width and length (). Sirenicapillaria has cyanobacteria with the largest cell size and sheath, especially S. rigida.
Table 1. Morphological characteristics of known species of Sirenicapillaria gen. nov., Tigrinifilum gen. nov. and Capilliphycus. Measurements (μm) are represented as min–max and mean ± st. dev
Morphological analyses demonstrated that Sirenicapillaria is composed of taxa with very long thalli (c. 20 cm), filaments with facultative sheaths, and three species: S. glauca (), S. rigida (), and S. stauglerae (). In a healthy lab environment, this genus has concentrated pigments and appears dark (), whereas in the field, in high light, or with unfavourable conditions, it turns light brown to gold colours (Fig. S2). Sirenicapillaria filaments are very long (up to 10 cm), with a thick sheath at the middle of the filament () and a gradual attenuation of the filament towards the ends with a thin sheath or none at all (). The largest species in terms of filament width is S. rigida, often observed with a lamellate or thickened gelatinous sheath (). Compared to its sister species, S. rigida has extremely tough filaments (), forming an intertwined, rigid and net-like mat; this is potentially the justification for the misidentification of Sirenicapillaria as Microseira [Lyngbya] in local reports.
Sirenicapillaria stauglerae and S. glauca overlap in both cell width and length. Although S. stauglerae and S. glauca have similar dimensions, these species are easily differentiated based on thallus colour and cell colour; S. stauglerae is generally brown and golden brown while S. glauca maintains a greyish, blue-green pigmentation (). The thallus of Sirenicapillaria is reminiscent of that of Lyngbya majuscula Harvey ex Gomont and Dapis, where a very long thalli resembles fine hair underwater (). Unlike L. majuscula that was originally described as black hair from the coast of England (Gomont Citation1892), Sirenicapillaria is more gold to brown and was discovered from the Florida Gulf coast through the Florida Keys. While the morphology of S. rigida overlaps with L. majuscula, the latter reaches larger widths and, together with their different type regions, we do not suggest this to be L. majuscula.
The genus Tigrinifilum is somewhat comparable to previously described cyanobacterial taxa in Lyngbya, including L. aestuarii F. Liebman ex Gomont and L. salina Kützing ex Starmach (Komárek & Anagnostidis Citation2005). Shared morphological features between T. floridanum and L. aestuarii include rare false branching, pronounced cross wall granulation and a marine habitat. However, T. floridanum and L. aestuarii differ considerably in climatic conditions of the type locality, which for L. aestuarii is the North Sea off the coast of present day Friesland in northwestern Germany. Furthermore, there is a large difference in filament and cell sizes between T. floridanum and L. aestuarii, with the latter being up to 10 μm wider. Tigrinifilum floridanum also resembles L. salina in having similar filament width and cell length, as well as in the marine habitat. However, the type locality for L. salina is a thermal spa in Kissingen, Germany, and morphological differences including the presence of a layered and stratified sheath and a lack of constriction between cells within L. salina suggest that T. floridanum is novel. Additionally, T. floridanum demonstrates a calyptra while L. salina does not.
16S rRNA phylogeny, intergeneric and family p-distance analyses
BLASTN analysis of the 16S rRNA gene sequence of the 30 marine cyanobacterial isolates shows them as closely related to the genera Affixifilum, Capilliphycus, Limnoraphis, Limnospira and Neolyngbya with 97% or lower identity. BLASTN analysis of isolates from Tigrinifilum indicated 99.4% or lower identity to uncultured cyanobacteria found in coastal sediments of the Mediterranean Sea (Païssé et al. Citation2010) and in sulfidic springs of western USA (Headd & Engel Citation2014). Analysis of the 16S rRNA gene sequence of Sirenicapillaria isolates indicated high similarity to sequences of ‘cf. Lyngbya sp. BAN TS02ʹ (99.2%; HQ419195) from Australia and 97.1% or lower similarity to Limnoraphis hieronymusii (JN854140), Capilliphycus tropicalis (MF190468), cyanobacteria found in the guts of marine fish (Jones et al. Citation2018), and uncultured cyanobacteria ‘cf. Lyngbya’ or ‘cf. Limnoraphis’ from the freshwater Clear Lake in California (Kurobe et al. Citation2013) and from Lake Atitlán, Guatemala (Rejmánková et al. Citation2011; Komárek et al. Citation2013), respectively.
BI and ML phylogenetic analyses of the 16S rRNA gene sequence of the 30 isolates demonstrated a large clade containing other taxa including Affixifilum, Capilliphycus, Limnoraphis, Limnospira and Neolyngbya with high support (BS: 89%; PP: 1.0) (triangle in ). The large Sirenicapillariaceae clade is sister to the Laspinemataceae clade (BS: –; PP: 0.99) and closely related to the Lyngbya confervoides clade (BS: –; PP: 1.0). Within Sirenicapillariaceae, Sirenicapillaria (BS: 88%; PP: 1.0) and Tigrinifilum (BS: 76%; PP: 1.0) formed distinct clades (). The BI and ML analyses also showed clades of described genera including Capilliphycus (BS: –; PP: 1.0) and Limnoraphis (BS: 100%; PP: 1.0). Within Capilliphycus, C. flaviceps (BS: 97%; PP: 1.0) as well as C. guerandensis (BS: 99%; PP: 1.0) formed separate subclades. Within Sirenicapillaria, three separate clades formed including S. glauca (BS: 88%; PP: 1.0), S. rigida (BS: 77%; PP: 0.93) and S. stauglerae (BS: 91%; PP: 1.0). The clade that included Tigrinifilum had two subclades of T. floridanum (BS: 100%; PP: 0.66) and T. guerandense (BS: 74%; PP: 1.0).
Fig. 49. Maximum likelihood and Bayesian inference, showing Bayesian topology of the phylogenetic relationship of the 16S rRNA gene sequence of the 30 cyanobacterial strains presented in this work and 141 cyanobacterial taxa including Gloeobacter violaceus PCC 7421 (NR074282) as outgroup. Bootstrap support and posterior probabilities ≥50 and 0.5 are shown at consensus nodes of both inferential analyses, respectively. (*) represents the reference strains for the species; triangle indicates the branch where the family Sirenicapillariaceae is delimited; stars denote the type species of the genus.
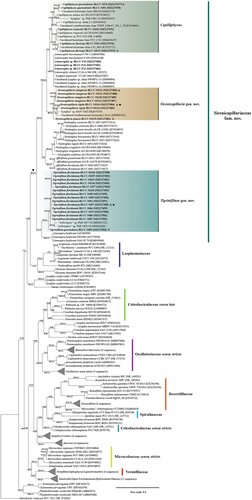
Intergeneric pairwise distances of the 16S rRNA gene sequence of the genera within the family Sirenicapillariaceae revealed genetic distances with 93.62%–97.97% among Affixifilum, Capilliphycus, Limnospira, Limnoraphis, Neolyngbya, Sirenicapillaria and Tigrinifilum (Table S2). Sirenicapillaria and Tigrinifilum showed ≤97.03% and ≤96.08% genetic identity to all other genera within Sirenicapillariaceae, respectively (Table S2).
The genetic similarity analyses of the 16S rRNA gene sequence across analysed established homocytous cyanobacterial families demonstrated ≤94.5% similarity for the novel family Sirenicapillariaceae (). Sirenicapillariaceae is closely related to Coleofasciculaceae sensu stricto (94.5%), Microcoleaceae (94.5%) and Laspinemataceae (94.4%), but comparatively distant to Oscillatoriaceae sensu stricto (92.6%). The within family genetic similarity of Sirenicapillariaceae was 97.2% while the within similarities of the established families were comparatively lower for Spirulinaceae (94.7%) and Desertifilaceae (95.3%) and ≥98.0% for the remaining families including Microcoleaceae, Oscillatoriaceae, Laspinemataceae, Vermifilaceae and Coleofasciculaceae. Morphological descriptions of the genera within Sirenicapillariaceae are summarized in .
Table 2. Mean p-distance (percentage of genetic similarity) between and within (last column) established cyanobacterial families, based on 16S rRNA gene sequences. Values are based on an alignment of 91 taxa and 7 established cyanobacterial families including the novel family Sirenicapillariaceae with eight genera: Affixifilum, Capilliphycus, Lyngbya, Limnospira, Limnoraphis, Neolyngbya, Sirenicapillaria and Tigrinifilum. Taxonomic family classifications were based on Guiry & Guiry (Citation2021) and Hauer & Komárek (Citation2021)
Table 3. Morphological characteristics of the genera Affixifilum, Neolyngbya, Limnoraphis, Capilliphycus, Sirenicapillaria gen. nov. and Tigrinifilum gen. nov. within the family Sirenicapillariaceae fam. nov.
16S–23S rRNA internal transcribed spacer (ITS) secondary structure and p-distance analyses
Lengths of the identifiable domains in the 16S–23S rRNA ITS sequence of all genera can be found in . Within Capilliphycus, the 16S–23S rRNA ITS secondary structure of both the D1-D1ʹ () and the Box-B () showed contrasting lengths and structures among both proposed species. Two D1-D1ʹ helices of C. guerandensis were recovered, tRNA dependent. Both helices were comparable in length (60 nucleotides) and were characterized by a six-nucleotide clamp (5ʹ- GACCUA—UAGGUC -3’) followed by a unilateral bulge on the 3ʹ side. A unilateral bulge before the terminal loop caused by a single nucleotide on the 5ʹ side was seen in the helix from the operon lacking tRNA. In the helix for the operon with tRNA, this unilateral bulge is present as a bilateral bulge (5ʹ- GA—A -3’). Capilliphycus flaviceps BLCC-M53 had a five-nucleotide clamp (5ʹ- GACCU—AGGUC -3’) followed by a unilateral bulge on the 3ʹ side. A bilateral bulge (5ʹ- AG—A -3’) occurs in the main stem before the large terminal loop. Capilliphycus flaviceps BLCC-M137 possessed two operons, with and without tRNAs, but only one D1-D1ʹ helix, which had a six-nucleotide clamp (5ʹ- GACCUA—UAGGUC -3’) followed by a unilateral bulge on the 3ʹ side. A bulge of unmatched base pairs (5ʹ- A—G -3’) occurs before a bilateral bulge (5ʹ- AA—A -3’), which is followed by the large terminal loop.
Table 4. Lengths of identifiable domains in the 16S–23S rRNA Internal Transcribed Spacer (ITS) regions of strains of Capilliphycus, Limnoraphis, Sirenicapillaria gen. nov. and Tigrinifilum gen. nov. Duplicated strain numbers indicate different ITS patterns (with or without tRNAs). Multiple values indicate variable operon length within species and/or strains. A dash (-) denotes not observed
Fig. 50. The 16S–23S rRNA ITS sequence secondary structures of the D1-D1ʹ of four cyanobacterial genera (Capilliphycus, Limnoraphis, Sirenicapillaria, Tigrinifilum) presented in this work. Nonbolded taxa represent previously published data. A: C. guerandensis BLCC-M76 & BLCC-M92. B: C. guerandensis BLCC-M79 & BLCC-M92 with tRNA. C: C. flaviceps BLCC-M53. D: C. flaviceps BLCC-M137 with and without tRNA. E: C. salinus. F: C. tropicalis. G: Limnoraphis sp. BLCC-F19 & BLCC-F23 with and without tRNA. H: S. glauca BLCC-M125. I: S. rigida BLCC-M116 & BLCC-M134 with and without tRNA. J: S. stauglerae BLCC-M121, BLCC-M122, BLCC-M123, & BLCC-M138. K: S. stauglerae BLCC-M121 with tRNA. L: T. floridanum BLCC-M48, BLCC-M50 clone A, BLCC-M57 clone C, BLCC-M66 clone B, BLCC-M77 clone C, BLCC-M87 clone C, BLCC-M107, BLCC-M118 clone B, BLCC-M119 clone B, & BLCC-M120 clones A & B, with tRNA. M: T floridanum BLCC-M50, BLCC-M57 clones A & B, BLCC-M66 clones A & C, BLCC-M73, BLCC-M87 clones A & B, BLCC-M118 clones A & C, BLCC-M119 clone C, & BLCC-M120 clone C, with tRNA. N: T. guerandensis BLCC-M99, with tRNA.
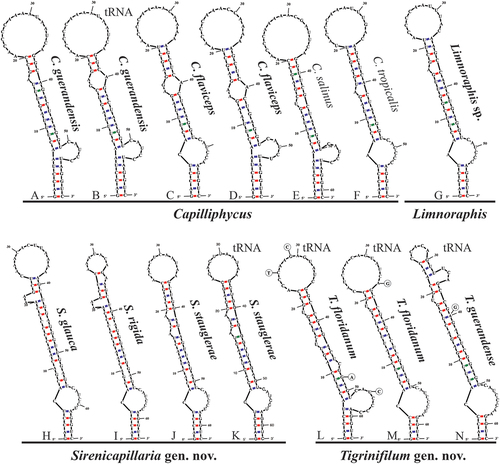
Fig. 51. The 16S–23S rRNA ITS sequence secondary structures of the Box-B of the four cyanobacterial genera (Capilliphycus, Limnoraphis, Sirenicapillaria, Tigrinifilum) presented in this work. Nonbolded taxa represent previously published data. A: C. flaviceps BLCC-M53. B: C. flaviceps BLCC-M137. C: C. flaviceps BLCC-M137 with tRNA. D: C. guerandensis BLCC-M76 & BLCC-M92 with and without tRNA. E: C. salinus. F: C. tropicalis. G: Limnoraphis sp. BLCC-F19 & BLCC-F23 with and without tRNA. H: S. glauca BLCC-M125. I: S. rigida BLCC-M116 & BLCC-M134 with and without tRNA. J: S. stauglerae BLCC-M121, BLCC-M122, BLCC-M123, & BLCC-M138 with and without tRNA. K: T. floridanum BLCC-M48, BLCC-M50, BLCC-M57, BLCC-M66, BLCC-M73, BLCC-M77, BLCC-M87, BLCC-M107, BLCC-M118, BLCC-M119, & BLCC-M120, with tRNA. L: T. guerandensis BLCC-M99, with tRNA.
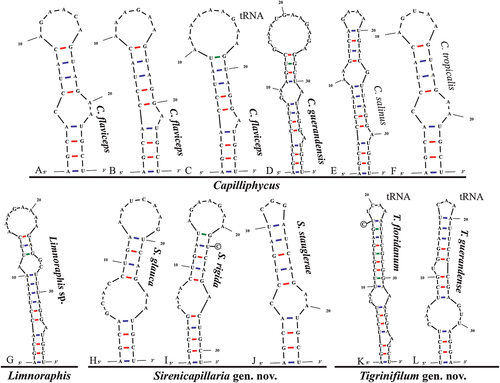
The Box B helices of all Capilliphycus species had a four-nucleotide clamp (5ʹ- AGCA—UGCU -3ʹ), although the helices varied in length and structure. The Box B of C. guerandensis is 40 nucleotides with a unilateral bulge caused by a single nucleotide on the 3ʹ side as well as a bilateral bulge (5ʹ- GA—A -3ʹ) before the terminal loop. The Box B of C. flaviceps BLCC-M53 is shorter (25 nucleotides) with a unilateral bulge (5ʹ- C—AA -3ʹ) in the main stem. Capilliphycus flaviceps BLCC-M137 contained two, tRNA dependent, Box B helices characterized by a unilateral bulge caused by a single nucleotide on the 3ʹ side and a terminal loop. The Box B helix of the tRNA operon was longer, 28 nucleotides compared to 25 nucleotides. No V3 helices could be identified.
Within Sirenicapillaria, the 16S–23S rRNA ITS secondary structure of both the D1-D1ʹ and the Box-B showed different lengths and structures among all proposed species. Sirenicapillaria glauca had a five-nucleotide clamp (5ʹ- GACCU—AGGUC -3ʹ) followed by a unilateral bulge on the 3ʹ side. A bulge of unmatched base pairs occurs in the stem after the first bulge (5ʹ- GA—GA -3ʹ) and again (5ʹ- C—A -3ʹ) before a unilateral bulge on the 5ʹ side before the terminal loop. Although operons with and without tRNAs were recovered, Sirenicapillaria rigida only possessed one D1-D1ʹ helix with a five-nucleotide clamp (5ʹ- GACCU—AGGUC -3ʹ) followed by a unilateral bulge on the 3ʹ side. A bulge of unmatched base pairs occurs in the stem after the first bulge (5ʹ- GA—GA -3ʹ) and again (5ʹ- C—A -3ʹ) before a unilateral bulge on the 5ʹ side. This is followed by a unilateral bulge on the 3ʹ side before a small terminal loop. Sirenicapillaria stauglerae possessed two operons, with and without tRNAs. The operon without tRNAs had a five-nucleotide clamp (5ʹ- GACCU—AGGUC -3ʹ) followed by a unilateral bulge on the 3ʹ side. A bulge of unmatched base pairs occurs after the first bulge (5ʹ- A—A -3ʹ). Two unilateral bulges on the 5ʹ side before the terminal loop. The D1-D1ʹ helix of the operons with tRNAs was similar to that of the helix without differing only at nucleotide 44, although this did not affect the structure. The helix is characterized by a five-nucleotide clamp (5ʹ- GACCU—AGGUC -3ʹ) followed by a unilateral bulge on the 3ʹ side. A bulge of unmatched base pairs occurs after the first bulge (5ʹ- A—A -3ʹ). Two unilateral bulges on the 5ʹ side before the terminal loop.
The Box B helices of all Sirenicapillaria species contained a four-nucleotide clamp (5ʹ- AGCA—UGCU -3ʹ), although the helices varied in length and structure. The Box-B of Sirenicapillaria glauca BLCC-M125 was 29 nucleotides long with a bilateral bulge after the clamp (5ʹ- GCAC—AA -3ʹ), causing a rightward lean. Only one BoxB helix of Sirenicapillaria rigida was recovered, tRNA independent, it contained a five base pair clamp (5ʹ- AGCAG—CUGCU -3ʹ) followed by a unilateral bulge after the clamp (5ʹ- CAA—A -3ʹ), causing a rightward lean, and a large terminal loop. Only one Box B helix from Sirenicapillaria stauglerae was recovered, tRNA independent. This BoxB helix was the shortest at 24 nucleotides and a unilateral bulge after the clamp (5ʹ- C—AA -3ʹ), causing a leftward lean, and a small (3 nucleotide) loop. V3 helices were recovered for all Sirenicapillaria species (data not shown).
Within Tigrinifilum, the 16S–23S rRNA ITS secondary structure of both the D1-D1ʹ and the Box-B showed dissimilar lengths and structures between both proposed species. Tigrinifilum floridanum had two D1-D1ʹ helices, which were 62 nucleotides and 63 nucleotides. The longest D1-D1ʹ helix is characterized by a five-nucleotide clamp (5ʹ- GACCU—AGGUC -3ʹ) followed by a large unilateral bulge on the 3ʹ side. A unilateral bulge caused by a single nucleotide on the 3ʹ occurs followed by two unilateral bulges by a single nucleotide on the 5ʹ side before the terminal loop, there were four variable bases throughout the helix, although they had no effect on the structure. The shorter D1-D1ʹ helix possessed a shorter, four base pair, clamp (5ʹ- GACC—GGUC -3ʹ) followed by a unilateral bulge on the 3ʹ side. A pair of unmatched nucleotides occurs after the first bulge (5ʹ- U—C -3ʹ). Both helices were recovered from different clones of the same strains in several isolates. Tigrinifilum guerandense contained one D1-D1ʹ helix characterized by a four-nucleotide clamp (5ʹ- GACC—GGUC -3ʹ) followed by a unilateral bulge on the 3ʹ side. A pair of unmatched nucleotides (5ʹ- A—A -3ʹ) occurs before a unilateral bulge by a single nucleotide on the 5ʹ side. A unilateral bulge of three nucleotides on the 3ʹ side occurs before a small terminal loop.
Each species of Tigrinifilum possessed one Box B helix. Tigrinifilum floridanum had a short three nucleotide clamp (5ʹ- AGC—GGU -3ʹ) followed by a unilateral bulge on the 3ʹ side caused by a single nucleotide. A bilateral bulge (5ʹ- GA—A -3ʹ) occurs within the helix before the small terminal loop. The Box B helix of T. guerandense was shorter than that of T. floridanum (34 vs 40 nucleotides). Tigrinifilum guerandense contained a four-nucleotide clamp (5ʹ- UCCU—AGGA -3ʹ) with a bilateral bulge (5ʹ- CAA—GTT -3ʹ). A unilateral bulge on the 5ʹ side caused by a single nucleotide occurs before the short terminal loop. The V3 helix of T. guerandense was recovered (data not shown) but no V3 helix was recovered from T. floridanum.
We also present the first full 16S–23S rRNA ITS sequence of Limnoraphis sp. from Florida, USA. Operons were recovered with and without tRNA but did not affect the D1-D1ʹ nor BoxB helices. The D1-D1ʹ helix is 58 nucleotides long and possessed a five-nucleotide clamp (5ʹ- GACCU—AGGUC -3ʹ) with a unilateral bulge caused by a single nucleotide on the 3ʹ before the terminal loop. The Box B helix is 38 nucleotides long and had a four-nucleotide clamp (5ʹ- AGCA—UGGU -3ʹ) followed by a unilateral bulge on the 3ʹ side caused by a single nucleotide. There is a bilateral bulge before the terminal bulb (5ʹ- C—AA -3ʹ); a V3 helix was not recovered.
The 16S–23S rRNA ITS sequence dissimilarity between species of Capilliphycus (without tRNA) including C. tropicalis, C. salinus T.A. Caires, Sant’Anna & J.M. Nunes, C. flaviceps and C. guerandensis was ≥14.8% (Table S3). The sequence dissimilarity between species of Sirenicapillaria including S. glauca, S. rigida and S. stauglerae was ≥11.6% (Table S4). The 16S–23S rRNA ITS region (with tRNAs) dissimilarity between the two species of Tigrinifilum, T. floridanum and T. guerandense, was ≥5.8% (Table S5).
DISCUSSION
Sampling of marine cyanobacteria in this work from seagrasses, lagoons, salt flats, and marine coastal areas, in addition to freshwater canals, yielded a total of 30 unicyanobacterial isolates (Table S1). The isolates in this work were used to demonstrate the taxonomic relationship among filamentous homocytous cyanobacteria using the 16S rRNA gene sequence phylogeny as well as the 16S–23S rRNA ITS region genetic distances and secondary structure of the Box-B and D1-D1ʹ helices. The 16S rRNA gene sequence phylogeny, reinforced by both BI and ML analyses and morphological analyses, supported the delineation of two novel cyanobacterial genera, Sirenicapillaria and Tigrinifilum. Pairwise distance analyses of the 16S rRNA gene sequence demonstrated similarities of ≤97.01% among the two novel genera Sirenicapillaria and Tigrinifilum and other closely related cyanobacterial genera, namely Affixifilum, Capilliphycus, Limnospira, Limnoraphis and Neolyngbya. Although the genetic distance is above the suggested generic cut-off rate of 94.5% (Yarza et al. Citation2014), additional genetic regions, especially the 16S–23S rRNA ITS region, are often helpful for delimiting genera of cyanobacteria along with morphological and ecological data.
The percentage of dissimilarity of the 16S–23S rRNA ITS region is another standard, in addition to gene phylogenies, that is used to delimitate cyanobacterial species (Pietrasiak et al. Citation2014; Vázquez-Martínez et al. Citation2018; González-Resendiz et al. Citation2019; Berthold et al. Citation2021; Lefler et al. Citation2021). The intrageneric percent dissimilarity within Capilliphycus and the two novel genera Sirenicapillaria and Tigrinifilum demonstrated high support for the delimitation of the seven novel species presented herein (Tables S3–S5). Based on the value of 7% dissimilarity of the 16S–23S rRNA ITS region for strong support in separating species, the novel species within the genera Capilliphycus (C. flaviceps and C. guerandensis) and Sirenicapillaria (S. glauca, S. rigida and S. stauglerae) demonstrate high support for their delimitations in addition to the robust 16S rDNA phylogenetic analyses, morphological differences and 16S–23S rRNA ITS region secondary structures. Although the intrageneric p-distance within Tigrinifilum was below the cut-off (5.8%), where 3%–7% dissimilarity requires additional lines of evidence for species delimitation (González-Resendiz et al. Citation2019), the strong support from the 16S rRNA gene phylogenetic analyses in addition to the morphological disparities support two novel species, T. floridanum and T. guerandense.
Sirenicapillaria is responsible for many coastal BCMs, especially on the western Florida coast in Lemon Bay where the reference strain was originally sampled (FDEP 2019–2021; Fig. S2). The cyanobacterium forms huge floating brownish mats, especially in spring and summer, which are reported to produce noxious odours when decomposing, with consequences for human health and coastal economies (Reilly Citation2019; Lane Citation2021). Although Sirenicapillaria is a prevalent bloomer, it is often reported as ‘Lyngbya’, ‘Lyngbya-like’, ‘Lyngbya majuscula’ or even the freshwater ‘Microseira wollei’. Sirenicapillaria is often found proliferating along with Vaucheria and other green algae (FDEP 2021), wrapped around turtlegrass (Thalassia testudinum Banks ex König) and manatee grass (Syringodium filiforme Kützing) or floating and decomposing around red mangrove (Rhizophora mangle Linnaeus) prop roots or marinas (Fig. S2). Lemon Bay is an aquatic preserve supporting mangrove, seagrass and oyster communities, and Sirenicapillaria can grow across several miles of benthos. Once Sirenicapillaria reaches large densities, through high productivity and oxygen bubble production, the mats can raise the seagrasses out of the sediments. The benthic blooms can inhibit coral larval settlement, produce secondary compounds that deter fish herbivory and indirectly affect aquatic mammal health by displacing seagrasses (Lydon et al. Citation2020; Ritson-Williams et al. Citation2020). Though some physical effects of Sirenicapillaria proliferations have been observed on seagrasses, the synergistic physical or chemical effects on the marine flora and fauna of these regions remain largely unknown. Further research is warranted to uncover potential toxicity and novel secondary compounds within Sirenicapillaria.
We found two species of Tigrinifilum, T. floridanum from the Florida coast and T. guerandense from salt flats in France (Fig. S2). Tigrinifilum is a genus that is distinctive by the highly granulated crosswalls that give the cyanobacterium a striped tiger appearance (). Another unique morphological feature of Tigrinifilum is the lack of granulations at the filament ends (). The species within Tigrinifilum are morphologically distinct, where T. guerandense is much smaller than T. floridanum, with no overlap in cell widths. Both species are slightly motile and tolerate high levels of salinity, especially T. guerandense, which was isolated from salt flats containing pools of briny seawater (>60 psu). Tigrinifilum is especially desiccation tolerant; mats of T. floridanum withstand drying for extensive periods (up to two months in laboratory cultures) and T. guerandense was also isolated from completely dried cyanobacterial mat material (Fig. S2). Tigrinifilum floridanum is a species that varies widely in pigmentation () and trichome width (8–19 µm; , 32–34), and is also widespread across the southeastern Florida coast, from Hollywood down to the Florida Keys.
Two species of Capilliphycus were previously known, including the type, C. salinus, and C. tropicalis. The novel species C. guerandensis from French salt flats differs by being wider than the other described species in the genus. Capilliphycus flaviceps overlaps in filament width with previously described Capilliphycus species, though C. flaviceps has a yellowing pigmentation at the apices and an extension of apical sheaths (). This yellowing feature can also be seen in C. guerandensis, although confined to the apical cells (), and not extending into a region as in C. flaviceps (). A feature shared between species in Capilliphycus is the spiralling or contortion of trichomes or the rearrangement of cells within a filament, as seen in both C. tropicalis (Caires et al. Citation2018b, p. 297, fig. 3E) and C. flaviceps ().
In agreement with previous observations, Capilliphycus contains cyanobacteria with tolerance for high salt concentrations. Caires et al. (Citation2018b) indicated that C. salinus was isolated from tidal pools of 50 psu. Comparatively, C. guerandensis was isolated from hypersaline ponds and tidal flats (>60 psu) and grew in media with a salinity of up to 100 psu. Herein we also present a strain of C. tropicalis (BLCC-M106) with a full 16S–23S rRNA ITS region isolated from Florida marine waters, extending the known range of this species from Brazil to Florida. The occurrence of Capilliphycus in both countries is another indication of the similarities between (sub)tropical Florida marine coastal cyanobacterial diversity and that of tropical northern Brazil (Caires & Affe Citation2021; Lefler et al. Citation2021). In our study, the Capilliphycus species that were isolated from Florida were components of benthic proliferations mainly formed by other larger homocytous filamentous cyanobacteria, such as Sirenicapillaria and Okeania.
In addition to the novel genera and species presented herein, we also recognize a new family, Sirenicapillariaceae. The new family is supported by its monophyly () as well as genetic p-distance similarities using the 16S rRNA gene sequences of taxa from established families including Microcoleaceae, Coleofasciculaceae sensu stricto, Vermifilaceae and Oscillatoriaceae sensu stricto (). Sirenicapillariaceae is currently composed of seven genera of marine and freshwater cyanobacteria: Affixifilum, Capilliphycus, Limnoraphis, Limnospira, Neolyngbya, Sirenicapillaria and Tigrinifilum (; ).
CONCLUSION
Cyanobacterial mats from the coast of Florida were sampled and isolated along with cyanobacteria from a French salt flat. A total of 30 cyanobacterial isolates are presented herein, of which 21 represent two novel genera isolated from Florida coasts and seagrasses: Sirenicapillaria and Tigrinifilum. Three isolates from the French salt flat led to the erection of two novel species: Capilliphycus guerandensis and Tigrinifilum guerandense. Additionally, three freshwater isolates of Limnoraphis were included for a more robust taxon sampling of the novel family Sirenicapillariaceae.
To describe the novel cyanobacterial diversity presented in this work, a combination of techniques was applied including morphology, 16S rRNA gene phylogeny and genetic similarities, along with 16S–23S rRNA ITS secondary structure and genetic p-distances. The polyphasic approach demonstrated congruency among analyses and strong support for the delimitation of the two novel genera Sirenicapillaria and Tigrinifilum, and the new species of Capilliphycus. Family level phylogenetic analysis of the 16S rRNA gene sequence supported the erection of Sirenicapillariaceae which includes Capilliphycus, Limnoraphis, Limnospira, Neolyngbya, Sirenicapillaria and Tigrinifilum.
A most crucial taxonomic establishment within this research was the identification of the frequent proliferating benthic cyanobacterial genus Sirenicapillaria, which has been commonly misidentified as the freshwater Microseira (‘Lyngbya’) wollei or the marine L. majuscula, or species of Dapis or Okeania. Sirenicapillaria proliferates across the western Florida Coast and is also found in floating mats in the Florida Keys. Large swaths of seagrasses succumb annually to blooms of these cyanobacteria, especially in spring and summer, imposing large constraints on local economies and negative impacts on marine biota. Recognizing Sirenicapillaria, a large player in benthic cyanobacterial bloom events, is a fundamental step in deciphering the diversity and putative emerging threats within marine environments.
Supplemental Material
Download Zip (59.6 MB)ACKNOWLEDGEMENTS
We thank Dr. Michael Wynne and Dr. Craig Schneider for nomenclatural enlightenments and Betty Staugler for sampling some of the benthic proliferations from which strains were isolated for this work.
DISCLOSURE STATEMENT
No potential conflict of interest was reported by the authors.
SUPPLEMENTARY MATERIAL
Supplemental data for this article can be accessed on the publisher’s website.
Additional information
Funding
REFERENCES
- Benny N., Thomas L.C. & Padmakumar K.B. 2021. Environmental influences on the cyanobacterial mat formation in the mangrove ecosystems along the southwest coast of India. Marine Ecology [Online]: Article e12685. DOI:https://doi.org/10.1111/maec.12685.
- Berthold D.E., Lefler F.W. & Laughinghouse H.D., IV. 2021. Untangling filamentous marine cyanobacterial diversity from the coast of South Florida with the description of Vermifilaceae fam. nov. and three new genera: Leptochromothrix gen. nov., Ophiophycus gen. nov., and Vermifilum gen. nov. Molecular Phylogenetics and Evolution 160: Article 107010. DOI: https://doi.org/10.1016/j.ympev.2020.107010.
- Biessy L., Wood S.A., Chinain M., Roué M. & Smith K.F. 2021. Exploring benthic cyanobacterial diversity and co-occurring potentially harmful dinoflagellates in six Islands of the South Pacific. Hydrobiologia 848: 2815–2829. DOI: https://doi.org/10.1007/s10750-021-04599-6.
- Caires T.A. & Affe H.M.J. 2021. Brazilian coast: a significant gap in the knowledge of Cyanobacteria and their applications. In: Cyanobacteria – recent advances in taxonomy, ecology and applications (Ed. by W.N. Hozzein). IntechOpen, Rijeka, Croatia. 16 pp. DOI: https://doi.org/10.5772/intechopen.97151.
- Caires T.A., Lyra G.M., Hentschke G.S., Pedrini A.G., Sant’Anna C.L. & Nunes J.M.C. 2018a. Neolyngbya gen. nov. (Cyanobacteria, Oscillatoriaceae): a new filamentous benthic marine taxon widely distributed along the Brazilian coast. Molecular Phylogenetics and Evolution 120: 196–211. DOI: https://doi.org/10.1016/j.ympev.2017.12.009.
- Caires T.A., Lyra G.M., Hentschke G.S., Silva A.M.S., Araújo V.L., Sant’Anna C.L. & Nunes J.M.C. 2018b. Polyphasic delimitation of a filamentous marine genus, Capillus gen. nov. (Cyanobacteria, Oscillatoriaceae) with the description of two Brazilian species. Algae 33: 291–304. DOI: https://doi.org/10.4490/algae.2018.33.11.25.
- Coleman M.A., Minne A.J.P., Vranken S. & Wernberg T. 2020. Genetic tropicalisation following a marine heatwave. Scientific Reports 10: Article 12726. DOI: https://doi.org/10.1038/s41598-020-69665-w.
- Collins A. 2021. Resources for understanding blue-green algae (cyanobacteria) blooms. UF/IFAS, University of Florida BLOGS. http://blogs.ifas.ufl.edu/manateeco/2020/07/14/resources-for-understanding-blue-green-algae-cyanobacteria-blooms/http://blogs.ifas.ufl.edu/manateeco/2020/07/14/resources-for-understanding-blue-green-algae-cyanobacteria-blooms/; searched on 19 April 2021.
- Di Lorenzo E. & Mantua N. 2016. Multi-year persistence of the 2014/15 North Pacific marine heatwave. Nature Climate Change 6: 1042–1047. DOI: https://doi.org/10.1038/nclimate3082.
- Edgar R.C. 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research 32: 1792–1797. DOI: https://doi.org/10.1093/nar/gkh340.
- Engene N., Choi H., Esquenazi E., Rottacker E.C., Ellisman M.H., Dorrestein P.C. & Gerwick W.H. 2011. Underestimated biodiversity as a major explanation for the perceived rich secondary metabolite capacity of the cyanobacterial genus Lyngbya: secondary metabolite diversity of Lyngbya. Environmental Microbiology 13: 1601–1610. DOI: https://doi.org/10.1111/j.1462-2920.2011.02472.x.
- Engene N., Rottacker E.C., Kaštovsky J., Byrum T., Choi H., Ellisman M.H., Komárek J. & Gerwick W.H. 2012. Moorea producens gen. nov., sp. nov. and Moorea bouillonii comb. nov., tropical marine cyanobacteria rich in bioactive secondary metabolites. International Journal of Systematic and Evolutionary Microbiology 62: 1171–1178. DOI: https://doi.org/10.1099/ijs.0.033761-0.
- Engene N., Paul V.J., Byrum T., Gerwick W.H., Thor A. & Ellisman M.H. 2013. Five chemically rich species of tropical marine cyanobacteria of the genus Okeania gen. nov. (Oscillatoriales, Cyanoprokaryota). Journal of Phycology 49: 1095–1106. DOI: https://doi.org/10.1111/jpy.12115.
- Engene N., Tronholm A. & Paul V.J. 2018. Uncovering cryptic diversity of Lyngbya: the new tropical marine cyanobacterial genus Dapis (Oscillatoriales). Journal of Phycology 54: 435–446. DOI: https://doi.org/10.1111/jpy.12752.
- FDEP (Florida Department of Environmental Protection). 2019–2021. Algal bloom sampling sites. https://floridadep.gov/dear/algal-bloom/content/algal-bloom-sampling-results ; searched on 01 April 2021.
- Ford A.K., Bejarano S., Nugues M.M., Visser P.M., Albert S. & Ferse S.C.A. 2018. Reefs under siege – the rise, putative drivers, and consequences of benthic cyanobacterial mats. Frontiers in Marine Science 5: Article 18. DOI: https://doi.org/10.3389/fmars.2018.
- Ford A.K., Visser P.M., van Herk M.J., Jongepier E. & Bonito V. 2021. First insights into the impacts of benthic cyanobacterial mats on fish herbivory functions on a nearshore coral reef. Scientific Reports 11: 7147. DOI: https://doi.org/10.1038/s41598-021-84016-z.
- Gomont M. 1892 ‘1893ʹ. Monographie des Oscillariées (Nostocacées Homocystées). Deuxième partie. – Lyngbyées. Annales des Sciences Naturelles, Botanique, Série 7 16: 91–264.
- González-Resendiz L., Johansen J.R., León-Tejera H., Sanchéz L., Segal-Kischinevzky C., Escobar-Sánchez V. & Morales M. 2019. A bridge too far in naming species: a total evidence approach does not support recognition of four species in Desertifilum (Cyanobacteria). Journal of Phycology 55: 898–911. DOI: https://doi.org/10.1111/jpy.12867.
- Guiry M.D. & Guiry G.M. 2021. AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. http://www.algaebase.org; searched on 31 May 2021.
- Gurgel C.F.D., Camacho O., Minne A.J.P., Wernberg T. & Coleman M.A. 2020. Marine heatwave drives cryptic loss of genetic diversity in underwater forests. Current Biology 30: 1199–1206. DOI: https://doi.org/10.1016/j.cub.2020.01.051.
- Hauer T. & Komárek J. 2021. CyanoDB 2.0 – on-line database of cyanobacterial genera. World-wide electronic publication, Univ. of South Bohemia & Inst. of Botany AS CR. http://www.cyanodb.cz; searched on 31 May 2021.
- Headd B. & Engel A.S. 2014. Biogeographic congruency among bacterial communities from terrestrial sulfidic springs. Frontiers in Microbiology 5: Article 473. DOI: https://doi.org/10.3389/fmicb.2014.00473.
- Jones J., DiBattista J.D., Stat M., Bunce M., Boyce M.C., Fairclough D.V., Travers M.J. & Huggett M.J. 2018. The microbiome of the gastrointestinal tract of a range-shifting marine herbivorous fish. Frontiers in Microbiology 9: Article 2000. DOI: https://doi.org/10.3389/fmicb.2018.02000.
- Komárek J. 2016. A polyphasic approach for the taxonomy of cyanobacteria: principles and applications. European Journal of Phycology 51: 346–353. DOI: https://doi.org/10.1080/09670262.2016.1163738.
- Komárek J. & Anagnostidis K. 2005. Cyanoprokaryota. II. Oscillatoriales. In: Süßwasserflora von Mitteleuropa, vol. 19 (2) (Ed. by B. Büdel, L. Krienitz, G. Gärtner & M. Schagerl). Spektrum Akademischer Verlag, Heidelberg, Germany. 759 pp.
- Komárek J., Zapomelová E., Smarda J., Kopecký J., Rejmanková E., Woodhouse J., Neilan B.A. & Komárková J. 2013. Polyphasic evaluation of Limnoraphis robusta a water-bloom forming cyanobacterium from Lake Atitlán, Guatemala, with a description of Limnoraphis gen. nov. Fottea 13: 39–52. DOI: https://doi.org/10.5507/fot.2013.004.
- Kurobe T., Baxa D.V., Mioni C.E., Kudela R.M., Smythe T.R., Waller S., Chapman A.D. & Teh S.J. 2013. Identification of harmful cyanobacteria in the Sacramento-San Joaquin Delta and Clear Lake, California by DNA barcoding. SpringerPlus 2: Article 491. DOI: https://doi.org/10.1186/2193-1801-2-491.
- Lane C. 2021. Mats of algae clog marina, affect business. https://www.amisun.com/2021/05/24/mats-of-algae-clog-marina-affect-business/https://www.amisun.com/2021/05/24/mats-of-algae-clog-marina-affect-business/; searched on 24 May 2021.
- Lefler F.W., Berthold D.E. & Laughinghouse H.D., IV. 2021. The occurrence of Affixifilum gen. nov. and Neolyngbya (Oscillatoriaceae) in South Florida (USA), with the description of A. floridanum sp. nov. and N. biscaynensis sp. nov. Journal of Phycology 57: 92–110. DOI: https://doi.org/10.1111/jpy.13065.
- Lydon C.A., Mathivathanan L., Sanchez J., Dos Santos L.A.H., Sauvage T., Gunasekera S.P., Paul V.J. & Berry J.P. 2020. Eudesmacarbonate, a eudesmane-type sesquiterpene from a marine filamentous cyanobacterial mat (Oscillatoriales) in the Florida Keys. Journal of Natural Products 83: 2030–2035. DOI: https://doi.org/10.1021/acs.jnatprod.0c00203.
- Miller M.A., Pfeiffer W. & Schwartz T. 2010. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Proceedings of the Gateway Computing Environments Workshop (GCE), pp 1–8. New Orleans, Louisiana, USA.
- Mühlsteinová R., Hauer T., De Ley P. & Pietrasiak N. 2018. Seeking the true Oscillatoria: a quest for a reliable phylogenetic and taxonomic reference point. Preslia 90: 151–169. DOI: https://doi.org/10.23855/preslia.2018.151.
- Nübel U., Garcia-Pichel F. & Muyzer G. 1997. PCR primers to amplify 16S rRNA genes from cyanobacteria. Applied and Environmental Microbiology 63: 3327–3332. DOI: https://doi.org/10.1128/aem.63.8.3327-3332.1997.
- Oliver E.C.J., Benthuysen J.A., Bindoff N.L., Hobday A.J., Holbrook N.J., Mundy C.N. & Perkins-Kirkpatrick S.E. 2017. The unprecedented 2015/16 Tasman Sea marine heatwave. Nature Communications 8: Article 16101. DOI: https://doi.org/10.1038/ncomms16101.
- Paerl H.W. & Paul V.J. 2012. Climate change: links to global expansion of harmful cyanobacteria. Water Research 46: 1349–1363. DOI: https://doi.org/10.1016/j.watres.2011.08.002.
- Païssé S., Goñi-Urriza M., Coulon F. & Duran R. 2010. How a bacterial community originating from a contaminated coastal sediment responds to an oil input. Microbial Ecology 60: 394–405. DOI: https://doi.org/10.1007/s00248-010-9721-7.
- Paul V.J., Thacker R.W., Banks K. & Golubić S. 2005. Benthic cyanobacterial bloom impacts the reefs of South Florida (Broward County, USA). Coral Reefs 24: 693–697. DOI: https://doi.org/10.1007/s00338-005-0061-x.
- Pearce A.F. & Feng M. 2013. The rise and fall of the “marine heat wave” off Western Australia during the summer of 2010/2011. Journal of Marine Systems 111-112: 139–156. DOI: https://doi.org/10.1016/j.jmarsys.2012.10.009.
- Pietrasiak N., Mühlsteinová R., Siegesmund M.A. & Johansen J.R. 2014. Phylogenetic placement of Symplocastrum (Phormidiaceae, Cyanophyceae) with a new combination S. californicum and two new species: S. flechtnerae and S. torsivum. Phycologia 53: 529–541. DOI: https://doi.org/10.2216/14-029.1.
- Reilly S. 2019. Algae still mucking, stinking up Lemon Bay. Englewood Sun, Englewood, Florida, USA. https://www.yoursun.com/englewood/news/algae-still-mucking-stinking-up-lemon-bay/article_07446afc-7029-11e9-892f-5f57c138c062.htmlhttps://www.yoursun.com/englewood/news/algae-still-mucking-stinking-up-lemon-bay/article_07446afc-7029-11e9-892f-5f57c138c062.html; searched on 19 April 2021.
- Reilly S. 2020. Algae mats lingering in Lemon Bay. Englewood Sun, Englewood, Florida, USA. https://www.yoursun.com/englewood/news/algae-mats-lingering-in-lemon-bay/article_62ace6ca-71c4-11e9-99ba-6398317cc9f6.htmlhttps://www.yoursun.com/englewood/news/algae-mats-lingering-in-lemon-bay/article_62ace6ca-71c4-11e9-99ba-6398317cc9f6.html; searched on 19 April 2021.
- Rejmánková E., Komárek J., Dix M., Komárková J. & Girón N. 2011. Cyanobacterial blooms in Lake Atitlán, Guatemala. Limnologica 41: 296–302. DOI: https://doi.org/10.1016/j.limno.2010.12.003.
- Ritson-Williams R., Arnold S.N. & Paul V.J. 2020. The impact of macroalgae and cyanobacteria on larval survival and settlement of the scleractinian corals Acropora palmata, A. cervicornis and Pseudodiploria strigosa. Marine Biology 167: Article 31. DOI: https://doi.org/10.1007/s00227-019-3639-5.
- Ronquist F. & Huelsenbeck J.P. 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: 1572–1574. DOI: https://doi.org/10.1093/bioinformatics/btg180.
- Sendall B.C. & McGreggor G.B. 2018. Cryptic diversity within the Scytonema complex: characterization of the paralytic shellfish toxin producer Heterosyctonema crispum, and the establishment of the Family Heteroscytonemataceae (Cyanobacteria/Nostocales). Harmful Algae 80: 158–170. DOI: https://doi.org/10.1016/j.hal.2018.11.002.
- Sharp K., Arthur K.E., Gu L., Ross C., Harrison G., Gunasekera S.P., Meickle T., Matthew S., Luesch H., Thacker R.W. et al. 2009. Phylogenetic and chemical diversity of three chemotypes of bloom-forming Lyngbya species (Cyanobacteria: Oscillatoriales) from reefs of Southeastern Florida. Applied and Environmental Microbiology 75: 2879–2888. DOI: https://doi.org/10.1128/AEM.02656-08.
- Sneed J.M., Meickle T., Engene N., Reed S., Gunasekera S. & Paul V.J. 2017. Bloom dynamics and chemical defenses of benthic cyanobacteria in the Indian River Lagoon, Florida. Harmful Algae 69: 75–82. DOI: https://doi.org/10.1016/j.hal.2017.10.002.
- Stecher G., Tamura K. & Kumar S. 2020. MEGA: Molecular Evolutionary Genetics Analysis (MEGA) for macOS. Molecular Biology and Evolution 37: 1237–1239. DOI: https://doi.org/10.1093/molbev/msz312.
- Suda S., Moriya R., Sumimoto S., Ohno O. & Suenaga K. 2013. Genetic diversity of filamentous cyanobacteria from shore regions of Okinawa. Journal of Marine Science and Technology 21: 175–180. DOI: https://doi.org/10.6119/JMST-013-1220-7.
- Urrutia-Cordero P., Zhang H., Chaguaceda F., Geng H. & Hansson L.-A. 2020. Climate warming and heat waves alter harmful cyanobacterial blooms along the benthic–pelagic interface. Ecology 101: Article e03025. DOI: https://doi.org/10.1002/ecy.3025.
- Vázquez-Martínez J., Gutierrez-Villagomez J.M., Fonseca-García C., Ramírez-Chávez E., Mondragón-Sánchez M.L., Partida-Martínez L., Johansen J.R. & Molina-Torres J. 2018. Nodosilinea chupicuarensis sp. nov. (Leptolyngbyaceae, Synechococcales) a subaerial cyanobacterium isolated from a stone monument in central Mexico. Phytotaxa 334: 167–182. DOI: https://doi.org/10.11646/phytotaxa.334.2.6.
- Wilmotte A., Auwera G.V. & Wachter R.D. 1993. Structure of the 16S ribosomal RNA of the thermophilic cyanobacterium Chlorogloeopsis HTF (‘Mastigocladus laminosus HTF’) strain PCC7518, and phylogenetic analysis. FEBS Letters 317: 96–100. DOI: https://doi.org/10.1016/0014-5793(93)81499-P.
- Wood S.A., Kelly L.T., Bouma-Gregson K., Humbert J.-F., Laughinghouse IV H.D., Lazorchak J., McAllister T.G., McQueen A., Pokrzywinski K., Puddick J. et al. 2020. Toxic benthic freshwater cyanobacterial proliferations: challenges and solutions for enhancing knowledge and improving monitoring and mitigation. Freshwater Biology 65: 1824–1842. DOI: https://doi.org/10.1111/fwb.13532.
- Yarza P., Yilmaz P., Pruesse E., Glöckner F.O., Ludwig W., Schleifer K., Whitman W.B., Euzéby J., Amann R. & Rosselló-Mora R. 2014. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nature Reviews Microbiology 12: 635–645. DOI: https://doi.org/10.1038/nrmicro3330.
- Zimba P.V., Shalygin S., Huang I.-S., Momčilović M. & Abdulla H. 2021. A new boring toxin producer – perforafilum tunnelli gen. & sp. nov. (Oscillatoriales, Cyanobacteria) isolated from Laguna Madre, Texas, USA. Phycologia 60: 10–24. DOI: https://doi.org/10.1080/00318884.2020.1808389.
- Zuker M. 2003. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Research 31: 3406–3415. DOI: https://doi.org/10.1093/nar/gkg595.
