ABSTRACT
In most organisms ranging from cyanobacteria to humans, the endogenous timekeeping system temporally coordinates the behavioral, physiological, and metabolic processes with a periodicity close to 24 h. The timing of these daily rhythms is orchestrated by the synchronized oscillations of both the central pacemaker in the brain and the peripheral clocks located across multiple organs and tissues. A growing body of evidence suggests that the central circadian clock and peripheral clocks residing in the metabolically active tissues are incredibly well coordinated to confer coherent metabolic homeostasis. The interplay between nutrient metabolism and circadian rhythms can occur at various levels supported by the molecular clock network, multiple systemic mechanisms, and the neuroendocrine signaling pathways. While studies suggest the reciprocal regulation between circadian clock and metabolism, it is important to understand the precise mechanisms and the underlying pathways involved in the cross-talk among circadian oscillators and diverse metabolic networks. In addition to the internal synchronization of the metabolic rhythms, feeding time is considered as a potential external synchronization cue that fine tunes the timing of the circadian rhythms in metabolic peripheral clocks. A deeper understanding of how the timing of food intake and the diet composition drive the tissue-specific metabolic rhythms across the body is concomitantly important to develop novel therapeutic strategies for the metabolic disorders arising from circadian misalignment. This review summarizes the recent advancements in the circadian clock regulation of nutrient metabolism and discusses the current understanding of the metabolic feedback signals that link energy metabolism with the circadian clock.
Introduction
“It is not the most intellectual of the species that survives; it is not the strongest that survives, but the species that survives is the one that is able best to adapt and adjust to the changing environment in which it finds itself.” – Charles Darwin (Origin of species)
To survive and adapt to the changing environmental day-night cycles caused due to our Earth’s rotation, there evolved 24 h rhythms in organisms, the circadian rhythms. These endogenous, biological clocks help the organisms to adapt to the daily as well as seasonal cycles and are thus indispensable (Sharma and Chandrashekaran Citation2005).
It is believed that circadian clocks evolved as an adaptation to anticipate the daily changes in the external cyclic environmental factors and to align the phase of behavioral, physiological, and metabolic rhythmicity of an organism with the most suitable time of the day. These rhythms are generated by the core clock machinery consisting of interlocked transcriptional-translational feedback loops (TTFLs) with several clock genes and their product proteins (Hardin Citation2005; Hurley et al. Citation2016). Based on the expression of clock proteins and circadian neuropeptides, circadian pacemaker neurons that drive behavioral rhythms were identified in both mammals and Drosophila (Hastings et al. Citation2019; Helfrich-Förster et al. Citation2007; Kaneko and Hall Citation2000; Kaneko et al. Citation1997; Shafer et al. Citation2006). The circadian clock gene expression is not exclusive to the brain, it is widespread across multiple organs and tissues throughout the body and the circadian pacemaker in the brain synchronizes these peripheral clocks through neuronal and endocrine signaling (Albrecht Citation2012; Glossop and Hardin Citation2002). The relative contributions of the individual clock components vary in different peripheral clocks to function in diverse cellular contexts (Mohawk et al. Citation2012).
An interesting aspect of circadian physiology is to understand how clock genes function in different metabolic tissues and the specific roles of both the central clock in the brain and the peripheral clocks in the metabolically active tissues for the maintenance of metabolic homeostasis. Emerging evidence indicates that food, energy status, and small biomolecules send feedback signals to the clock to sense the changes in the metabolism (Reinke and Asher Citation2019). Comprehensive understanding of how the metabolic pathways are regulated by the circadian clock and how the energy state and small metabolites feedback into clock mechanisms provide new insights into circadian physiology and help to develop novel approaches to treat metabolic disorders. In this review, we summarize the current understanding of the crosstalk between circadian clock and metabolism by including the recent advancements based on the studies carried out so far in Drosophila and mammals. We further discuss the multiple systemic, endocrine signals and the molecular pathways that couple the circadian clock with metabolism.
Circadian regulation of metabolism
Molecular machinery of the circadian clock
Our current understanding of the molecular machinery of the circadian clock is profoundly influenced by the molecular genetics analyses carried out in the genetically amenable model system Drosophila melanogaster. Genetic analysis conducted by Seymour Benzer and Ronald Konopka revealed the first clock gene period (per) and subsequent studies identified its heterodimer partner timeless (tim) in Drosophila (Konopka and Benzer Citation1971; Sehgal et al. Citation1994). The evolutionarily conserved transcriptional activator Clock was initially identified in mice and later in Drosophila (Clk) as a heterodimeric partner for cycle (cyc) (Allada et al. Citation1998; Rutila et al. Citation1998; Vitaterna et al. Citation1994). Collectively, studies in Drosophila unveiled the core clock machinery as an interlocked TTFL with CLOCK (CLK) and CYCLE (CYC) forming the positive limb while PERIOD (PER) and TIMELESS (TIM) making up the negative limb (Benito et al. Citation2007; Darlington et al. Citation1998; Hurley et al. Citation2016) (). This TTFL is remarkably conserved across numerous species (Brown et al. Citation2012; Dunlap and Loros Citation2017; Hurley et al. Citation2016). In mammals, the transcriptional activator CLOCK heterodimerizes with CYCLE homolog BMAL1 and activates the transcription of Period (Per1, Per 2, Per3) and Cryptochrome (Cry1, Cry2) genes (Kondratov et al. Citation2003; Kwon et al. Citation2006) (). The PERIOD (PER) and CRYPTOCHROME (CRY) protein heterodimers translocate back to the nucleus to inhibit their own transcription by interacting with CLOCK and BMAL1 (Griffin et al. Citation1999; Kume et al. Citation1999). Although CRY is a circadian photoreceptor in Drosophila, it functions as a transcriptional repressor along with PER to drive the molecular oscillations in mammals (Kume et al. Citation1999; Stanewsky et al. Citation1998).
Figure 1. Circadian clock in Drosophila and mammals. (a) Primary and secondary feedback loops in the Drosophila circadian clock. The core clock machinery in Drosophila is composed of an interlocked TTFL with the positive limb CLK/CYC and a negative limb PER/TIM. Circadian photoreceptor CRY mediates the light entrainment of circadian clocks in Drosophila. The secondary feedback loop consists of VRI and PDP1 that inhibit and activate Clk transcription, respectively. (b) Primary and secondary feedback loops in the mammalian circadian clock. In mammals, the TTFL consists of the positive arm CLOCK/BMAL1 and a negative arm PER/CRY. REV-ERB and ROR, inhibit and activate Bmal1 transcription, respectively.
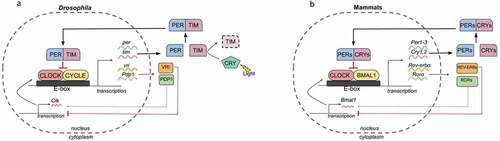
In Drosophila, the mRNA and protein levels of Clk oscillate in antiphase with respect to the mRNA and protein levels of per/tim, indicating a differential transcriptional regulation of Clk. Subsequent studies showed that a secondary feedback loop interlocked with the per/tim primary feedback loop indeed regulates the Clk transcription to confer robustness to the oscillators (Cyran et al. Citation2003). CLOCK/CYCLE activates the transcription of vrille (vri) and Par Domain Protein 1 (Pdp1). The transcription factors VRI and PDP1 in turn feedback to rhythmically inhibit and activate the Clk transcription, respectively. Thus, vri, Pdp1, and Clk form a secondary feedback loop to generate accurate circadian rhythms in Drosophila (Cyran et al. Citation2003; Glossop et al. Citation2003) (). The secondary feedback loop in mammals consists of the receptors REV-ERB and retinoic acid-related orphan receptor (ROR), which inhibit and activate Bmal1 transcription, respectively, and fine-tune the circadian rhythms (Guillaumond et al. Citation2005; Preitner et al. Citation2002) ().
In addition to the transcriptional-translational feedback loop regulation, post-translational regulation plays a crucial role in maintaining the circadian oscillations. Among the post-translational modifications, kinase-mediated phosphorylation of clock proteins plays an important role in controlling transcriptional repression potency, nucleocytoplasmic distribution, and stability of clock proteins (Reischl and Kramer Citation2011). Casein kinase I (CKI) isoforms and Casein kinase 2 (CK2) are the most important kinases in mammals that phosphorylate PER, whereas in Drosophila, PER is phosphorylated by DOUBLE TIME (DBT), an ortholog of CKIε and by CK2 (Akten et al. Citation2003; Cyran et al. Citation2005; Kloss et al. Citation1998; Knippschild et al. Citation2005; Lin et al. Citation2005; Maier et al. Citation2009; Price et al. Citation1998; Reischl and Kramer Citation2011; Tsuchiya et al. Citation2009). In addition to this, glycogen synthase kinase 3 (GSK3) is another important kinase involved in circadian regulation. Its homolog SHAGGY (SGG) phosphorylates TIM in Drosophila and influences the nuclear entry of PER/TIM complex in the circadian pacemaker neurons (Martinek et al. Citation2001). In mammals, GSK3β isoform regulates the BMAL1 protein stability through phosphorylation (Sahar et al. Citation2010). Furthermore, many other clock components are rhythmically phosphorylated by multiple kinases and phosphatases to generate rhythmic circadian oscillations (Reischl and Kramer Citation2011).
Central and peripheral clocks
Based on the expression of PER, TIM, and the circadian neuropeptide Pigment Dispersing Factor (PDF), approximately 150 neurons distributed across the fly brain have been identified to constitute the central clock circuitry (Helfrich-Förster et al. Citation2007; Kaneko and Hall Citation2000; Kaneko et al. Citation1997; Shafer et al. Citation2006). This circadian pacemaker neuronal network composed of subsets of small and large ventral lateral neurons (sLNv and lLNv), dorsal neurons (DNs), dorsal lateral neurons (LNd), and lateral posterior neurons (LPN) govern behavioral rhythms in Drosophila () (Helfrich-Förster Citation1997; Kaneko and Hall Citation2000; Kaneko et al. Citation1997; Klarsfeld et al. Citation2004; Shafer et al. Citation2006). The expression of clock proteins is not restricted to the central clock neurons of the fly brain, but also present in the peripheral circadian clocks operating in multiple tissues and organs throughout the body (Emery et al. Citation1997; Giebultowicz and Hege Citation2002; Plautz et al. Citation1997; Saez and Young Citation1988). Although central and peripheral clocks show evolutionarily conserved remarkable similarities, there are significant differences in their resetting mechanism and the relative contributions of the specific clock components vary in different cellular clocks (Ito and Tomioka Citation2016).
Figure 2. Central and peripheral circadian clocks in Drosophila and mammals.
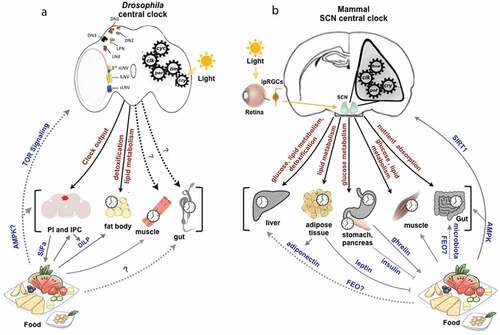
In mammals, the hypothalamic suprachiasmatic nucleus (SCN) composed of a population of about 20,000 neurons and astrocytes, was identified as the central circadian oscillator that drives behavioral rhythms. These SCN neuronal circadian oscillators are coupled to one another via intercellular signaling and this coupling is paramount for the synchrony of circadian rhythms (Hegazi et al. Citation2019). The mammalian circadian system consists of many cellular clocks operating hierarchically across multiple tissues and organs throughout the body (). At the top of the hierarchy is the central clock that synchronizes the peripheral circadian clocks via systemic cues and endocrine signals (Mohawk et al. Citation2012). The peripheral circadian clock drives the circadian expression of diverse genes involved in various physiological processes such as metabolic, cardiovascular, reproductive, immune, and endocrine system functions (Richards and Gumz Citation2012). The interplay between the central clock and peripheral circadian clocks in metabolically active tissues is not well ascertained. Understanding the role of metabolic peripheral clocks and its intricate connections with the central clock would expand our understanding of circadian physiology and how the circadian clock enables an organism to achieve metabolic homeostasis.
Role of central and peripheral clocks in metabolism
Drosophila has served as an excellent model system to understand the molecular basis of the circadian clock and to dissect the neural circuits that drive the behavioral rhythms. In addition to understanding the molecular basis of the clock and behavioral rhythms, studies in the last decade explored the powerful genetic approaches available in Drosophila to understand the still largely unknown circadian regulation of different physiological processes, including metabolism, to provide an in-depth understanding of the circadian physiology of an organism (Allada and Chung Citation2010; Shi and Zheng Citation2013). A number of studies on clock mutants showed that clock function is required for glucose and lipid homeostasis. Loss of function of critical circadian regulator gene per affects the levels of diacylglycerols (DAG) and acylcarnitines (AC) involved in lipid metabolism and the per heterodimer partner tim regulates fat metabolism under dietary restriction in Drosophila (Katewa et al. Citation2016; Schäbler et al. Citation2020; Ulgherait et al. Citation2016). Flies feed primarily during early morning hours resulting in a rapid increase in circulating sugar that is subsequently stored as glycogen. The daily rhythms in feeding behavior and increase in carbohydrate levels were disrupted in cryptochrome null (cryo) flies, suggesting that the circadian photoreceptor cry plays an important role in regulating the feeding behavior and carbohydrate homeostasis (Seay and Thummel Citation2011). Studies also identified sets of metabolite rhythms associated with endogenous clock and light-dark (LD) cycles. Among the measured metabolites, 9% of metabolites exhibited circadian oscillation under constant darkness. Whereas per01 flies displayed strong ultradian rhythms in metabolite oscillations, indicating that the endogenous clock machinery drives the metabolite rhythms in Drosophila (Rhoades et al. Citation2018).
Many of the findings from initial studies on mice gave insights on the role of the clock in governing metabolic physiology and phenotype. For instance, mice carrying a mutation on the Clock gene (ClockΔ19) were obese with hypertriglyceridemia and attenuated expression of transcripts encoding metabolically relevant neuropeptides (Turek et al. Citation2005). A mutation on another essential clock component Bmal1, a heterodimer partner of CLOCK, impaired glucose homeostasis (Rudic et al. Citation2004) and adipogenesis in mice (Shimba et al. Citation2005). Plasma glucose and triglycerides exhibit circadian variation in wild type mice with a peak at ~circadian time 4 (CT4), and this temporal variation was disrupted in Bmal1−/− mutant mice, indicating the importance of core clock component in driving metabolic oscillations (Rudic et al. Citation2004). Other clock gene components implicated in circadian control of metabolism are the downstream components of CLOCK/BMAL1 heterodimer, such as CRY and PER. Disruption of Cry1 and Cry2 resulted in glucose intolerance (Lamia et al. Citation2011) and altered lipogenic pathways (Bur et al. Citation2009), whereas knockdown of Per2 primarily attenuated two critical metabolic rhythms, the glucocorticoid and food intake rhythms (Yang et al. Citation2009). Circadian peak in serum glucocorticoid level is locked with the activity phase in Per2+/+ mice during the early night, whereas this temporal pattern is attenuated in Per2−/− mice (Yang et al. Citation2009). Changes in circadian regulation of corticosteroids may affect carbohydrate and lipid metabolism. These studies showed the important role played by both the clock gene transcription activators and repressors in circadian integration of metabolism and energy homeostasis.
Role of liver and adipose tissue peripheral clocks in metabolism
To better understand how the central clock in the brain and the peripheral clocks in various metabolic tissues contribute to circadian physiology and metabolism, a number of peripheral clock-specific ablation or loss of function studies have been carried out in Drosophila and mammals. Among the various peripheral clocks present in metabolically active tissues, the hepatic clock is of paramount importance to metabolic physiology due to its essential role in glucose and lipid homeostasis. In Drosophila, clock proteins are expressed in various metabolically active tissues, including the fat body, which is the functional equivalent of the mammalian liver and adipose tissue () (Scott et al. Citation2004). Studies focused on understanding the functional significance of fat body peripheral clock in Drosophila revealed that disruption of the fat body clock by expressing a dominant-negative version of CLK abolished tim rhythmicity in the fat body (Xu et al. Citation2008). In addition, disruption of the fat body clock decreased the levels of triglyceride and glycogen storage, whereas perturbing the central clock neurons increased the levels of this nutrient storage, indicating that central and peripheral fat body clock oppose each other to achieve metabolic homeostasis in Drosophila (DiAngelo et al. Citation2011; Xu et al. Citation2008).
Drosophila exhibits 24 h rhythmicity in feeding behavior with the maximum food intake occurring during the early daytime. This clock-controlled feeding rhythm is eliminated in Drosophila lacking clock genes (Seay and Thummel Citation2011; Xu et al. Citation2008). Flies with disrupted fat body circadian clock exhibited enhanced food intake and altered phase of feeding rhythm, suggesting that the fat body circadian clock participates in the regulation of the feeding rhythm (Xu et al. Citation2008). In addition to driving the feeding rhythmicity, the fat body circadian clock drives rhythmic expression of various genes involved in metabolism, detoxification, and immunity (Xu et al. Citation2011). Microarray analysis identified the genes rhythmically expressed in the fat body of wild type flies. Some 40% of the genes peaked in the late night, whereas only a few genes peaked during the mid-late day. This study further analyzed the cycling transcripts in the fat body and identified a large group of metabolic genes rhythmically expressed with a peak during the night, including the transporter genes and the genes implicated in lipid metabolism (Xu et al. Citation2011). In addition, rhythmic expression of 81 transcripts were dampened or were arrhythmic as a result of the disruption of the fat body peripheral clock, indicating that it drives rhythmic expression of genes in the fat body involved in metabolism, detoxification, and immune response (Xu et al. Citation2011). These studies showed the relevance of the fat body peripheral clock in the temporal organization of metabolic processes.
Studies on CLOCK-deficient mice showed that there were changes in the clock gene and protein expression in both the suprachiasmatic nucleus and liver. But, the molecular clock continued to run, and the mice also showed circadian rhythm in locomotor activity (Debruyne et al. Citation2006). Transcription factor Neuronal PAS Domain Protein 2 (NPAS2) maintains the rhythmicity in the central clock in the absence of Clock gene expression, whereas liver peripheral clocks are arrhythmic without the CLOCK, indicating that NPAS2, alone, is unable to sustain the peripheral clock rhythmicity (DeBruyne et al. Citation2007). These findings support the notion that fundamental differences exist in the core clock machinery, depending on their functional context in various tissues.
Subsequent tissue-specific ablation studies were instrumental in demonstrating the circadian autonomous function of the liver clock and its importance in metabolism. Liver-specific deletion of Bmal1 resulted in hypoglycemia and loss of rhythmicity in several hepatic glucose regulatory genes (Lamia et al. Citation2008). In wild type mice liver, glucose transporter 2 (Glut2) circadian expression peaks during the subjective day, corresponding to the fasting phase, and the expression trough occurs during the subjective night, coinciding with the feeding phase. Liver-specific deletion of Bmal1 perturbs the circadian expression of Glut2, indicating the importance of liver clock in glucose homeostasis by regulating the daily rhythm in hepatic glucose transport and storage that counterbalances the feeding-fasting behavior cycle (Lamia et al. Citation2008). Reconstitution of BMAL1 exclusively in the liver of Bmal1 null mice demonstrated the degree of autonomy of the liver clock during the absence of clock function in the other body tissues. This autonomous circadian function of the liver specifically regulates certain metabolic pathways/processes, such as NAD+ salvage pathway and glycogen metabolism (Koronowski et al. Citation2019).
In addition to Bmal1 mediated regulation of glucose homeostasis, other clock machinery components, such as CRY, regulate the hepatic gluconeogenesis, and REV-ERBα integrates hepatic lipid metabolism (Zhang et al. Citation2016, Citation2010). In mouse liver, CRY levels are elevated during morning and it inhibits gluconeogenic gene expression, whereas reduced CRY levels at night promote gluconeogenesis (Zhang et al. Citation2010). Thus, CRY modulates hepatic gluconeogenesis in a temporal manner in accordance with changes in the external environment. Studies on the human osteosarcoma cell line and mouse primary hepatocytes identified and characterized a small molecule-carbazole derivative KL001 that specifically interacts with CRY and prevents its degradation. KL0001-mediated CRY stabilization inhibits gluconeogenesis in primary hepatocytes, and this finding may have relevance to develop therapeutic approaches for diabetes (Hirota et al. Citation2012). In the liver, peroxisome proliferator-activated receptor α (PPARα), which promotes mitochondrial fatty acid β-oxidation, is diurnally regulated (Yang et al. Citation2006). PPARα elevates utilization of fatty acids during the beginning of the night, coinciding with the onset of activity and foraging (Li and Li Citation2012). Thus, circadian clocks temporally orchestrate glucose and lipid metabolism in coordination with the feeding, activity, and daily changes in the environment.
Another dynamic metabolic tissue central to energy homeostasis is the adipose tissue, and the adipocytes exhibit robust oscillations of clock genes (Zvonic et al. Citation2006). Studies focused on characterizing the circadian regulation of adipocyte physiology showed that clock genes regulate the key functions of the adipose tissue, such as lipolysis and lipogenesis in a rhythmic manner (Shostak et al. Citation2013). In wild type mice, triglyceride is transported to adipose tissue predominantly during the night when the animal is active and foraging. Adipose tissue promotes lipolysis during the day, coinciding with the inactive-fasting phase, and circadian clock temporally coordinates lipid metabolism (Shostak et al. Citation2013). Studies also reported that lipoprotein lipase (LPL) activity peaks during the dark phase in the adipose tissue to promote fat accumulation (Benavides et al. Citation1998), whereas mobilization of free fatty acid is increased during the light phase (Suzuki et al. Citation1983). Lipid metabolism is temporally coordinated within adipose and among metabolic tissues. For example, triglyceride synthesis occurs in a rhythmic manner in the liver, with an increase during the actively feeding dark phase (Martin et al. Citation1979). Hepatic triglyceride synthesis is enhanced in phase, with the increased level of LPL activity in adipose tissue during the dark phase. This temporal coordination of lipid metabolism among liver and adipose tissue probably ensures efficient storage of dietary carbon as fat in the adipose tissue.
Adipocyte-specific deletion of Bmal1 was associated with reduced polyunsaturated fatty acids, altered food intake rhythm, and obesity (Paschos et al. Citation2012). The hypothalamus drives the rhythm in food intake by rhythmically expressing the appetite regulating neuropeptide Y (NPY) (Wiater et al. Citation2011). Disruption of the adipocyte clock increases hypothalamic expression of neuropeptides governing appetite and feeding rhythms (Paschos et al. Citation2012). These findings indicate the importance of the adipocyte clock, per se, in the temporal organization of energy regulation and the significance of feedback signals from adipocyte clock to the hypothalamus to achieve energy homeostasis.
Hypothalamic and pancreatic circadian clock-mediated regulation of feeding rhythm and metabolism
Studies in Drosophila investigated the roles of the pars intercerebralis (PI), the functional homolog of mammalian hypothalamus, and the insulin-producing cells (IPCs), functionally analogous to the mammalian pancreatic β-cells, in circadian regulation of metabolism and feeding rhythm. In Drosophila, PI neuronal populations expressing the neuropeptide SIFamide (SIFa) act as circadian output centers to control the feeding rhythm. On the other hand, IPCs, a neuronal subset of PI expressing Drosophila Insulin-Like Peptides (DILPs), affect the amount of food intake without affecting the feeding rhythm () (Dreyer et al. Citation2019). In addition to the homeostatic regulation of feeding behavior, IPCs are connected with the central clock neurons via DN1 neurons (Cavanaugh et al. Citation2014). Inputs from the central clock and the feeding rhythm drive the IPC firing rhythm, and these IPCs act as a metabolic clock output region. The IPC firing rhythm regulates the rhythmic expression of the lipase transcript sxe2 in the fat body via the clock output signaling molecule insulin (Barber et al. Citation2016) (). These studies collectively reveal the importance of PI as a circadian clock output center in integrating feeding behavior and metabolism of Drosophila.
While PI acts as a clock output center without an autonomous clock, the mammalian hypothalamus, itself, hosts autonomous circadian oscillators to essentially generate circadian rhythmicity in a wide range of behavioral rhythms, including feeding. Subdivisions of the arcuate nucleus (ARC) and dorsomedial nuclei (DMH) were found to show intrinsic rhythmicity (Abe et al. Citation2002; Guilding et al. Citation2009). Mammalian hypothalamic neuropeptide Y (NPY) exhibits 24 h rhythmicity in its expression. NPY receptor-expressing neurons in the mediobasal hypothalamic area (MBH) and leptin receptor-expressing neurons in the ARC are essential for integrating the sleep-wake cycle and feeding rhythm in mammals (Li et al. Citation2012; Wiater et al. Citation2011). It was also recently shown that the AgRP neurons in the ARC play an important role in regulating the feeding rhythm (Cedernaes et al. Citation2019). Apart from regulating the feeding rhythmicity, core clock gene Bmal1 expressed in the ventromedial hypothalamus (VMH) regulates energy expenditure (Orozco-Solis et al. Citation2016). These studies shed light on the time-tracking system in the hypothalamus in the temporal regulation of sleep-wake cycle, feeding rhythm, and energy homeostasis.
In addition to the peripheral clocks present in metabolic tissues, such as liver, adipose tissue, and hypothalamus, a self-sustained circadian clock in pancreatic β-cells governs glucose metabolism and energy homeostasis in mice (Sadacca et al. Citation2011). Disruption of pancreatic circadian clock by specifically eliminating Clock or Bmal1 in the islet cells attenuated the insulin level and proliferation of islet cells, indicating the importance of pancreatic clock in insulin production and glucose maintenance (Marcheva et al. Citation2010; Sadacca et al. Citation2011). To address the role of Rev-erbα in pancreatic function, small interfering RNA was used in MIN-6 cells and mouse islet cells. Down regulation of Rev-erbα reduced the mRNA levels of lipogenic genes, such as sterol regulatory element-binding protein 1 c (Srebp-1 c) and its target gene fatty acid synthase (FAS) in mouse islet cells (Vieira et al. Citation2012). In addition, insulin secretion and the genes involved in the exocytotic machinery, such as Vamp3, Munch18, SNAP25, and Syntaxin1a, were reduced in these cell models when Rev-erbα was silenced (Vieira et al. Citation2012). These results suggest that the molecular clock secondary feedback loop component Rev-erbα expressed in the pancreatic islets is important in regulating insulin secretion and lipid metabolism (Allaman-Pillet et al. Citation2004; Frese et al. Citation2007; Vieira et al. Citation2012). Additionally, Rev-erbα agonist GSK41112 increased glucose-induced insulin secretion, and the antagonist SR8278 reduced insulin release (Vieira et al. Citation2012). These synthetic ligands targeting REV-ERB may offer a basis to develop the treatment for diabetes.
Muscle circadian clock regulates metabolic rhythm
Global and tissue-specific clock gene ablation or loss of function studies were carried out in mouse models to understand the physiological importance of the intrinsic muscle timekeeping system. In Drosophila, the role, if any, that the muscle peripheral clock plays in metabolism remains to be studied. Muscle-specific ablation of core clock gene Bmal1 in mice impaired insulin-mediated glucose uptake, glucose oxidation, and the protein levels of insulin-dependent glucose transporter GLUT4 (Dyar et al. Citation2014; Harfmann et al. Citation2016). To understand how the muscle clock drives metabolic rhythms, the genome-wide putative binding sites of the muscle clock transcription factors, such as BMAL1 and REV-ERBα were analyzed in mice. It was found that BMAL1 activates expression of the gene Diacylglycerol acyltransferase 2 (Dgat2) to promote lipid storage, whereas REV-ERBα represses the major target genes involved in lipid catabolism and protein turnover, thus temporally regulating the gene expression and energy fluctuations (Dyar et al. Citation2018a). LPL activity exhibits 24 h variation with an opposite phase between the skeletal muscle and adipose tissues of nocturnal rats (Tsutsumi et al. Citation2002). LPL activity increases in the adipose tissue and decreases in the skeletal muscle during the dark phase, leading to accelerated fat accumulation by suppressing fat oxidation. Its activity increases in the skeletal muscle during the light phase to promote fat uptake and oxidation (Tsutsumi et al. Citation2002). It is possible that this inverse daily circadian rhythm in LPL activity between adipose tissue and skeletal muscle ensures efficient storage of triglyceride and maintains adequate balance of body fat in mammals (Tsutsumi et al. Citation2002).
Signaling between the central and peripheral clocks
Communication between the environment, the central pacemaker, and the downstream peripheral clocks effectively coordinates the physiological functions, metabolism, and behavior of an organism. The internal organization of the central clock and the neuroendocrine mechanisms mediate the clock-to-clock coupling and synchrony. Pigment dispersing factor (PDF) is the most extensively studied circadian neuropeptide in the Drosophila circadian pacemaker network that drives synchrony among the neuronal populations of the clock network (Helfrich-Förster et al. Citation2000; Park and Hall Citation1998; Renn et al. Citation1999). Another important neuropeptide expressed in the circadian pacemaker neuron subset LNds and LNvs is the neuropeptide F (NPF) (He et al. Citation2013; Johard et al. Citation2009). NPF peptidergic signaling in the brain and insulin signaling from the IPCs to the fat body drive the rhythmic expression of metabolic genes in the fat body (Barber et al. Citation2016; Erion et al. Citation2016).
The mammalian central pacemaker SCN synchronizes the peripheral clocks by relaying time signals through neuronal networks and the endocrine system (Buijs et al. Citation2006). While neurons in the SCN communicate among each other essentially by neurotransmitters, such as vasoactive intestinal polypeptide (VIP) and gastrin releasing peptide (GRP) (Aida et al. Citation2002; Meyer-Spasche and Piggins Citation2004), SCN neurons communicate with extra-hypothalamic targets through vasopressin (AVP) and gamma-aminobutyric acid (GABA) (Patton and Hastings Citation2018). Hormones and neurotransmitters play pivotal roles in the temporal control of metabolic functions by the circadian pacemaker. For instance, SCN mediates the nocturnal release of melatonin by the pineal gland (Hermes et al. Citation1996; Perreau-Lenz et al. Citation2004). Melatonin signaling, and more importantly SCN mediated rhythmic food intake, regulate the blood glucose circadian rhythmicity (Nagai et al. Citation1978; Owino et al. Citation2016; Yamamoto et al. Citation1987). The interaction between SCN and arcuate nucleus (ARC) is necessary to maintain the rhythmicity in feeding behavior and metabolic functions (Méndez‐Hernández et al. Citation2020). The paraventricular nucleus (PVN) of the hypothalamus receives projections from the SCN and ARC. NPY concentration in the PVN and ARC exhibits circadian rhythmicity, with a peak coinciding with the onset of activity and enhanced feeding (Jhanwar-Uniyal et al. Citation1990; Stanley and Leibowitz Citation1985). NPY/AgRP promotes food intake, and ablating NPY/AgRP neurons in the ARC disrupts the feeding rhythm (Wiater et al. Citation2011). It is possible that SCN directly regulates the release of NPY, but the exact mechanism remains to be understood. Glucocorticoid (GC) is considered as a potential signal that modulates the NPY/AgRP expression in the ARC (Shimizu et al. Citation2008). Vasopressin released from SCN terminal into the DMH has an inhibitory effect on plasma corticosterone concentration during the light period (Kalsbeek et al. Citation1992, Citation1996).
Glucocorticoid is an adrenal steroid hormone with robust circadian rhythmicity and an important time-giving signal relayed from SCN to peripheral tissue clocks to synchronize the daily metabolic programs (Pezük et al. Citation2012). The core clock components, Crys, mediate the rhythmic expression of glucocorticoid receptors (Lamia et al. Citation2011; Oster et al. Citation2017). GCs also promote lipolysis in adipocytes by increasing the transcription of adipose triglyceride lipase (ATGL) and hormone-sensitive lipase (HSL) (Peckett et al. Citation2011). GCs synchronize the expression of a wide range of circadian relevant genes, including the liver circadian transcriptome, and promote glucose production, utilization, and hepatic gluconeogenesis (Reddy et al. Citation2007). The SCN controls the daily rhythm in plasma glucose concentration. Synergistic effects of suprachiasmatic GABA-ergic inputs into the PVN and glutamatergic signaling balance the autonomic input into the liver to regulate hepatic glucose production and plasma glucose concentration. Thus, GABA-ergic and glutamatergic signaling in the PVN is an important signaling pathway for the SCN to mediate peripheral physiology and metabolism (Kalsbeek et al. Citation2008, Citation2004). In addition, a functional peripheral circadian oscillator in the pancreas is essential for insulin signaling to synchronize the liver clock (Sadacca et al. Citation2011; Yamajuku et al. Citation2012). Orexin is another hypothalamic neuropeptide that regulates appetite, sleep, and wakefulness. Activation of orexinergic neurons during the active phase promotes wakefulness and ensures efficient adjustment of the peripheral glucose metabolism in a temporal manner (Kalsbeek and Fliers Citation2013).
The feedback signal communication from local tissue clocks to the brain is important for energy homeostasis. Ghrelin is one such hormone rhythmically expressed by stomach oxyntic cells, and this hunger hormone acts largely in the brain and other peripheral tissues (LeSauter et al. Citation2009). Ghrelin-mediated regulation of hypothalamic orexinergic neurons is a possible link between energy balance and wakefulness (Yamanaka et al. Citation2003). Leptin hormone secreted from adipocytes is another important hormone that suppresses appetite and mobilizes excess energy storage by stimulating metabolism () (Zigman and Elmquist Citation2003). Plasma leptin levels exhibit circadian oscillation, and this hormone may bind with leptin receptors to activate the target genes in a time-dependent manner (Sukumaran et al. Citation2010). Although these hormones do not affect the SCN rhythms to great extent, they act as communication signals between the extra-SCN areas and peripheral clock to ensure the logical coordination of behavior and metabolic functions through the neuronal and endocrine systems (Begemann and Neumann A-M Citation2020). However, novel signaling pathways involved in achieving the synchrony among peripheral clocks are still a matter of investigation.
Circadian control of cellular metabolism
Circadian regulation of metabolites and NAD+ salvage pathway
Several lines of evidence from metabolomic studies reveal that a wide array of genes associated with various metabolic processes exhibit circadian oscillation with ~24 h rhythmicity. The circadian clock is implicated in rhythmically regulating the intracellular levels of various metabolites, such as lipids, nucleotides, acetyl-CoA, xenobiotic metabolites, glycolytic intermediates, and also the salvage pathway that serves the production of critical metabolite NAD+ (Eckel-Mahan et al. Citation2013; Eckel-Mahan and Sassone-Corsi Citation2009; Panda et al. Citation2002; Sahar et al. Citation2014) NAD+ also functions as a coenzyme to influence the activity of enzymes involved in other metabolic pathways, such as glycolysis, tricarboxylic acid (TCA) cycle, and fatty acid oxidation (Yaku et al. Citation2018). In particular, it modulates the function of many enzymes involved in mitochondrial energy production pathways. NAD+ abundance displays circadian rhythmicity (Nakahata et al. Citation2009), and this cyclic change in NAD+ level is likely to modulate mitochondrial oxidative function and act as a potential link between the circadian clock and metabolism. In addition, Class III histone deacetylase (HDAC) Sirtuin 1 (SIRT1) uses NAD+ as a coenzyme for its enzymatic activity and is tightly linked to the enzymatic feedback loop of the NAD+ salvage pathway. Circadian clock transcriptional activators CLOCK/BMAL1 regulate the circadian expression of a rate limiting enzyme nicotinamide phosphoribosyl transferase (NAMPT) in the NAD+ salvage pathway. This eventually leads to cyclic oscillation in the HDAC activity of the SIRT1 (Nakahata et al. Citation2009; Ramsey et al. Citation2009). SIRT1 transduces the signal back to the circadian clock to regulate the amplitude of the clock gene expression and thus forms a link between metabolism and chromatin remodeling of the circadian timing system (Nakahata et al. Citation2008) (discussed in detail later). It is also interesting to note that 5ʹ-methylthioadenosine (MTA) and S-adenosylmethionine (SAM), important metabolite precursors for chromatin modifications, show rhythmicity in mice that are amplified by high-fat diet (Krishnaiah et al. Citation2017).
Circadian regulation of mitochondrial energetics and redox rhythms
Evidence from recent studies on skeletal muscle tissue and mouse embryonic fibroblasts indicate that mitochondrial morphology, function, and abundance may change as a response to the nutrient cue an organism experiences from the environment (Bach et al. Citation2003; Gomes and Scorrano Citation2011). Nutrient availability during the active phase increases ATP, oxygen content, and reactive oxygen species (ROS) production, whereas nutrient deprivation during the resting phase decreases ATP content. Mitochondrial morphology varies as a response to these metabolic cues (Sardon Puig et al. Citation2018). Mitochondria form extensive elongated tubular networks through fusion, with increased mitochondrial respiration during energy consumption under nutrient deprivation (Gomes and Scorrano Citation2011), and become fragmented through the process of fission during the active phase of nutrient availability (Bach et al. Citation2003). A recent study on human skin fibroblasts and clock-deficient mice showed that these daily changes in mitochondrial dynamics as a response to metabolic cues are aligned to the LD cycle through circadian clock-dependent mechanisms (Schmitt et al. Citation2018). In addition to clock mediated morphological changes, mitochondrial respiration also displays clock-dependent rhythmicity in mammals (Neufeld-Cohen et al. Citation2016).
Mitochondrial respiration is a major source of ROS, and, hence, it is likely that the redox state may follow circadian rhythmicity (Edgar et al. Citation2012; Sena and Chandel Citation2012). Gene expression profiling of cycling genes in mice SCN revealed that several genes involved in carbon utilization and mitochondrial oxidative phosphorylation exhibit circadian rhythmicity, with a peak expression at CT22. Such strong regulation of genes involved in mitochondrial oxidative phosphorylation indicates rhythmicity in the redox state of the pacemaker neurons (Panda et al. Citation2002). Studies on mouse mutant strain and mouse embryo fibroblasts showed that core-clock components CLOCK, BMAL1, and REV-ERBα regulate the rhythmic expression, levels, and activity of antioxidant enzymes, such as catalase and glutathione (Gong et al. Citation2015; Pekovic-Vaughan et al. Citation2014; Sengupta et al. Citation2016; Wang et al. Citation2012; Xu et al. Citation2012). The NAD+/NADH ratio plays a crucial role in the production of major ROS superoxide and the circadian clock gene Per2 regulates the NAD+/NADH ratio and the response to oxidative stress as shown by experiments on mouse fibroblasts (Magnone et al. Citation2014). Sirtuin 3 (SIRT3) regulates various proteins involved in mitochondrial redox homeostasis. This deacetylase SIRT3 activity is dependent on NAD+ and the rate-limiting enzyme NAMPT in the NAD+ salvage pathway. Studies on mice liver also suggest that the circadian clock regulates redox metabolism through NAMPT-NAD+ mediated SIRT3 activity, and acetylation of SIRT3-targetted mitochondrial proteins show temporal oscillation (Mauvoisin et al. Citation2017). Together, these studies suggest that circadian clocks regulate the functioning of mitochondria and redox homeostasis through multiple ways. Although evidence from studies imply that mitochondrial respiration in various tissues is under the control of the circadian timing system, the role of clock genes in regulating mitochondrial function in a tissue-specific manner remains to be studied.
Metabolic feedback signals to the circadian clock
The circadian clock generates rhythmicity in cellular metabolic processes, and the metabolic signals in turn provide substantial feedback to the circadian timekeeping system. For example, dietary restriction increases the mRNA oscillation amplitude of the core clock genes and the clock protein levels in the brain and peripheral tissues of Drosophila (Katewa et al. Citation2016). One of the possible explanations is that protein dietary restriction modulates TOR (target of rapamycin) signaling, which in turn influences the circadian clock (Kapahi et al. Citation2010; Katewa and Kapahi Citation2010). It was also shown that the TOR pathway regulates the activity of GSK-3, the nuclear accumulation of TIM, and the period of circadian rhythmicity in Drosophila (Zheng and Sehgal Citation2010).
In mammals, mTOR (mammalian/mechanistic target of rapamycin) is a major nutrient/energy status sensor likely to serve as an important signal integrating nutrient/metabolic status with circadian system function. mTOR affects the circadian rhythm in both the central SCN clock and the various peripheral tissue clocks, such as hepatocytes and adipocytes (Ramanathan et al. Citation2018). Studies based on genetic and pharmacological approaches in mice showed that mTOR activation accelerates the circadian clock and increases the amplitude of oscillation, whereas mTOR inhibition lengthens the circadian period (Lipton et al. Citation2017; Ramanathan et al. Citation2018). On the contrary, silencing of tor in per-expressing cells decreases the periodicity of the activity/rest rhythm in flies, indicating a possible mechanistic difference in the clock machinery of Drosophila and mammals (Kijak and Pyza Citation2017; Zheng and Sehgal Citation2010).
Multiple pathways contribute to convey the metabolic status signals to the circadian timing system. Adenosine monophosphate (AMP)–activated protein kinase (AMPK) is a central sensor of metabolic cues, and it has been recently recognized as a key regulator of circadian clock function (). In Drosophila, AMPK regulates the circadian rhythms by phosphorylating CLOCK protein (Cho et al. Citation2019). However, in mammals, AMPK regulates the stability of core clock components through phosphorylation of CRY1 and CKIɛ, leading to degradation of PER2 (Lamia et al. Citation2009; Um et al. Citation2007). Thus, AMPK functions as an important link between metabolism and clock to mediate the metabolic entrainment of the peripheral clock and to determine the speed of the circadian clock (Um et al. Citation2007). Further studies are required to understand whether AMPK may act as an effector mechanism in the indirect light entrainment of peripheral clocks. Light entrains the activity/rest rhythm, and activity governs feeding behavior. Food intake-mediated nutrient status change can modulate AMPK activity, and it could function as a potential mechanism in the indirect light entrainment of the metabolic peripheral clocks. In addition to regulating the stability of clock proteins, AMPK increases HDAC activity of SIRT1 by enhancing NAMPT levels, and the mutual regulation of AMPK and SIRT1 is implicated in the crosstalk between the circadian clock and metabolic states (Cantó et al. Citation2009; Ruderman et al. Citation2010). In flies, it remains to be studied whether sirtuins play any role in the interaction between the circadian clock and metabolism.
SIRT1 in circadian chromatin remodeling
Empirical evidence from earlier studies in mammals suggest that the HDAC activity of SIRT1 enzyme is rhythmically regulated, and this NAD+ dependent deacetylase enzyme in turn regulates the oscillation amplitude of various core clock gene transcripts (Asher et al. Citation2008; Nakahata et al. Citation2008). SIRT1 governs the amplitude of clock gene oscillation through site-specific histone acetylation of the clock gene and, thus, contributes to circadian chromatin remodeling (Asher et al. Citation2008; Nakahata et al. Citation2008). SIRT1 binds with CLOCK-BMAL1 chromatin complex, and both CLOCK and SIRT1 regulate the rhythmic acetylation of BMAL1 (Hirayama et al. Citation2007; Nakahata et al. Citation2008). SIRT1 also mediates the stability of clock proteins and circadian clock gene expression through deacetylation and degradation of PER2. In addition to the circadian control of post-translational modification of BMAL1 and PER2, SIRT1 loss of function attenuates the magnitude of transcript oscillation of other clock genes, such as Cry1, Per1, and Rorγ (Asher et al. Citation2008). Given that SIRT1 uses NAD+, the critical metabolite of salvage pathway, as a coenzyme for its enzymatic activity, it is likely that SIRT1 acts as a transducing signal connecting the cellular metabolism to circadian timekeeping system circuitry ().
Food entrainment
Among the various environmental zeitgebers that entrain the circadian timing system, food availability is considered as a weaker time cue for the central clock in the brain. Prior studies assessed whether food can entrain the circadian rhythm in Drosophila, and the results showed that time-restricted feeding does not entrain either behavioral rhythm or the expression profile of clock genes in the peripheral clock (Oishi et al. Citation2004). However, a subsequent study reported that restricted feeding drives the rhythmic expression of clock genes in the fat body, but not in the brain central clock (Xu et al. Citation2011). Food entrainment in mammals is characterized by gradual increase in the locomotor activity prior to the feeding time. This food-anticipatory activity (FAA) is mediated by a self-sustained food-entrainable oscillator (FEO) (Mistlberger Citation2009). Numerous studies attempted to decode the molecular components of the FEO, and the results suggest that canonical circadian genes are not necessary for FEO and that a noncanonical timekeeping mechanism may drive FEOs (Feillet et al. Citation2006; Iijima et al. Citation2005; Mistlberger et al. Citation2008; Pendergast et al. Citation2009, Citation2017; Storch and Weitz Citation2009). Nevertheless, some of the studies have also shown that clock genes, indeed, modulate FAA. For instance, Bmal1 null mice lacked FAA and injecting viral vector expressing Bmal1 in the SCN did not restore the FAA, whereas rescue of Bmal1 expression in the DMH restored the food entrainment (Fuller et al. Citation2009).
DMH may not be the only locus in the brain for FEO, other sites in the stomach can act as FEOs to drive FAA. Recent studies indicate that stomach oxyntic cells co-express ghrelin and circadian clock proteins PER1 and PER2. Food can entrain the rhythmicity of this hunger hormone and clock proteins (). FAA is reduced in ghrelin receptor knockout mice, indicating that stomach oxyntic cells contain FEOs that anticipate the food intake time (LeSauter et al. Citation2009). Although continuous investigations during the past decades identified the discrete FEO sites in the brain and peripheral organs, the interactions between circadian oscillators involved in the timekeeping mechanism of the FEOs are yet to be discerned.
Feeding time and circadian misalignment
Timing of food consumption appears to be an effective zeitgeber for the entrainment of the peripheral circadian clocks. Feeding at inappropriate times disrupts the phase of the liver metabolic rhythm and uncouples it from the phase of the physiological rhythms dictated by the central SCN clock (Adamovich et al. Citation2014; Mauvoisin et al. Citation2014). A recent study compared the effect of two time-restricted feeding protocols on liver circadian clock oscillation. One involved time-restricted feeding (TRF) with a hypocaloric intake (HCT) with 2 h of access to food for 21 days. The other protocol involved TRF with a gradual increase in calories to achieve a normocaloric intake (NCT) at the end of the protocol (García-Gaytán et al. Citation2020). To assess the impact of HCT and NCT on liver circadian clock, daily oscillation of liver Per1 and Bmal1 were quantified. Both TRF protocols induced a similar phase shift in the liver Per1 and Bmal1 mRNA expression compared to mice provided with ad libitum food (AL). This result suggests that TRF, regardless of hypocaloric and normocalorie intake, elicits a similar phase shift in liver circadian clock compared to AL (García-Gaytán et al. Citation2020). Studies also showed that restricted feeding antiphasic to the LD cycles in mammals decouples the peripheral circadian clocks in tissues, such as pancreas, kidney, liver, and heart from the central pacemaker in the brain with no effect on the SCN-generated rhythms (Cassone and Stephan Citation2002; Damiola et al. Citation2000; Hara et al. Citation2001; Hirota and Fukada Citation2004; Oishi et al. Citation2002). This desynchronization of the central and peripheral clocks increases the risk of metabolic disorders (Mattson et al. Citation2014). These studies collectively indicate the dominant zeitgeber effect of light over food in the SCN. However, food can act as an effective entrainment cue on peripheral oscillators without affecting the central oscillators in the SCN. Misalignment of the timing of food intake with the activity phase governed by the central clock may result in the decoupling of peripheral oscillators, which will have profound metabolic consequences (Damiola et al. Citation2000). Recent studies also showed that feeding time significantly contributes to the rhythmic expression of metabolic genes in the liver, with mild effects on the peripheral clock gene oscillations. These studies indicate that feeding time directly regulates the metabolic rhythm in the liver, possibly by excluding the hepatic circadian clock (de Goede et al. Citation2018; Greenwell et al. Citation2019).
Apart from the timing of food intake, dietary composition also alters peripheral clock oscillations. However, the effect of calorie restriction on liver-specific clock gene expression is distinct from the effects of time-restricted feeding (Patel et al. Citation2016). In contrast to time-restricted feeding, which affects the peripheral clocks without affecting the SCN, metabolic cues associated with calorie restriction can entrain the SCN clock (Challet Citation2010). On the other hand, high-fat diet-mediated changes in the energy balance alters the periodicity of locomotor activity rhythms and the expression of canonical clock genes in the liver (Kohsaka et al. Citation2007). Nutrient challenge by a high-fat diet also disrupts the temporal correlation of metabolites among tissues (Dyar et al. Citation2018b). Further studies also show that a high-fat diet phase delays the daily rhythm of clock genes and adiponectin signaling pathway components in the liver, indicating that the high-calorie diet-mediated phase shift of the mammalian liver circadian timing system may cause misalignment of endogenous rhythms and perturb metabolic homeostasis (Barnea et al. Citation2009). Thus, diet composition and timing of food intake are likely to be potential environmental cues to perpetuate synchrony between the circadian rhythms and metabolic homeostasis.
Perspectives and conclusions
In recent years, substantial advancement has been made in our understanding of the crosstalk between the circadian clock and metabolism. In this review, we summarized the studies implicating the importance of the circadian timing system in the temporal coherence of glucose and lipid metabolic processes within and among tissues in both Drosophila and mammals. Furthermore, we discussed the role of the clock in NAD+ metabolism and highlighted the metabolic feedback signals that maintain the robustness of the circadian rhythms. SIRT1 relays nutritional cues to the circadian clock to shape the transcriptional profile of clock genes, and, in addition, AMPK also acts as a clock resetting signal by transmitting the nutritional input into the clock (Asher et al. Citation2008; Cho et al. Citation2019; Lamia et al. Citation2009; Nakahata et al. Citation2008; Um et al. Citation2007). Although studies suggest that these nutritional sensors transmit the nutrient information to both the brain and peripheral clocks (Lamia et al. Citation2009; Sun et al. Citation2007; Zhou et al. Citation2014), the extent to which SIRT1 and AMPK regulate the central and peripheral clocks remains to be largely understood. Further studies focusing on the tissue and organ-specific roles of AMPK, SIRT1, and other nutritional sensors would be insightful to obtain a comprehensive understanding on the significance of these metabolic signals in synchronizing the brain and tissue clocks.
In this review, we also emphasized the role of food as a potential zeitgeber that exerts a greater influence on the peripheral clock than the SCN central clock. However, the molecular components and the multiple loci of food-entrainable oscillators (FEOs) remain to be studied, and it is likely that FEOs are localized in other tissues in addition to the brain and stomach oxyntic cells (Fuller et al. Citation2009; LeSauter et al. Citation2009). It would be, indeed, worthwhile to explore the possibility of the presence of peripheral FEOs in the metabolically active tissues, such as liver and gut. Apart from this, the specific contribution and the clear-cut underlying mechanisms through which light and food act as zeitgebers to reset various peripheral circadian clocks to achieve metabolic homeostasis require more systematic empirical investigation. Metabolomics studies in the past decade have characterized the circadian clock-driven gene expressions in various tissues and organs throughout the body in mammals. Future studies incorporating higher temporal resolution in transcriptomics and metabolomics under various light and food environmental cues may enable us to elucidate how various metabolic pathways in the tissues and organs are differentially modulated by environmental time cues during different times of the day.
Evidence from recent studies indicates that the gut microbiome influences sleep in Drosophila and regulates the amplitude of circadian oscillators in peripheral clocks (Bi et al. Citation2018; Morioka et al. Citation2018; Selkrig et al. Citation2018). The increase in sleep associated with Wolbachia endosymbiotic bacterial infection is presumably due to the altered expression of essential genes involved in the dopamine biosynthesis pathway in Drosophila (Bi et al. Citation2018). It is also likely that the increase in sleep is an adaptive strategy to conserve the energy for the host. More studies in the future may decipher the importance of the interaction between gut microbiota and circadian clocks in metabolism.
The role of circadian clock component REV-ERB in lipid and glucose metabolism is well characterized. Subsequent identification of endogenous ligands for REV-ERB has made possible the design of synthetic ligands regulating the activity of these receptors (Reinking et al. Citation2005). Synthetic ligand GSK4112 exerts a repressive effect on gluconeogenic REV-ERB target genes and reduces hepatic glucose output. (Grant et al. Citation2010). In addition, studies also demonstrated the PPARγ agonist rosiglitazone and GSK4112 synergistically induce expression of genes involved in adipogenesis (Kumar et al. Citation2010). These developments in synthetic ligand targeting REV-ERB may be useful in the treatment of metabolic disorders. Identifying novel pathways that link circadian clock and metabolism would not only provide a comprehensive and accurate understanding of the mechanisms underlying metabolic disorders, but also have profound implications on improving therapeutic strategies for metabolic diseases.
Acknowledgements
The authors thank Drs. Pawan Kumar Jha, Antara Das and Pankaj Yadav for their careful reading and inputs to improve the manuscript. This work was supported by the DBT/Wellcome Trust India Alliance Fellowship [IA/E/15/1/502329] awarded to NNK and intramural fund from Indian Institute of Science Education and Research, Thiruvananthapuram. We thank the four anonymous reviewers for carefully reading the manuscript and suggesting some very useful changes.
Disclosure statement
The authors report no conflicts of interest.
Additional information
Funding
References
- Abe M, Herzog ED, Yamazaki S, Straume M, Tei H, Sakaki Y, Menaker M, Block GD. 2002. Circadian Rhythms in Isolated Brain Regions. J Neurosci. 22(1):350–356. doi:https://doi.org/10.1523/JNEUROSCI.22-01-00350.2002
- Adamovich Y, Rousso-Noori L, Zwighaft Z, Neufeld-Cohen A, Golik M, Kraut-Cohen J, Wang M, Han X, Asher G. 2014. Circadian clocks and feeding time regulate the oscillations and levels of hepatic triglycerides. Cell Metab. 19(2):319–330. doi:https://doi.org/10.1016/j.cmet.2013.12.016
- Aida R, Moriya T, Araki M, Akiyama M, Wada K, Wada E, Shibata S. 2002. Gastrin-releasing peptide mediates photic entrainable signals to dorsal subsets of suprachiasmatic nucleus via induction of Period gene in mice. Mol Pharmacol. Jan;61(1):26–34.
- Akten B, Jauch E, Genova GK, Kim EY, Edery I, Raabe T, Jackson FR. 2003. A role for CK2 in the Drosophila circadian oscillator. Nat Neurosci. 6(3):251–257. doi:https://doi.org/10.1038/nn1007
- Albrecht U. 2012. Timing to perfection: the biology of central and peripheral circadian clocks. Neuron. 74(2):246–260. doi:https://doi.org/10.1016/j.neuron.2012.04.006
- Allada R, Chung BY. 2010. Circadian organization of behavior and physiology in Drosophila. Annu Rev Physiol. 72(1):605–624. doi:https://doi.org/10.1146/annurev-physiol-021909-135815
- Allada R, White NE, So WV, Hall JC, Rosbash M. 1998. A mutant Drosophila homolog of mammalian Clock disrupts circadian rhythms and transcription of period and timeless. Cell. 93(5):791–804. doi:https://doi.org/10.1016/S0092-8674(00)81440-3
- Allaman-Pillet N, Roduit R, Oberson A, Abdelli S, Ruiz J, Beckmann JS, Schorderet DF, Bonny C. 2004. Circadian regulation of islet genes involved in insulin production and secretion. Mol Cell Endocrinol. 226(1–2):59–66. doi:https://doi.org/10.1016/j.mce.2004.06.001
- Asher G, Gatfield D, Stratmann M, Reinke H, Dibner C, Kreppel F, Mostoslavsky R, Alt FW, Schibler U. 2008. SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell. 134(2):317–328. doi:https://doi.org/10.1016/j.cell.2008.06.050
- Bach D, Pich S, Soriano FX, Vega N, Baumgartner B, Oriola J, Daugaard JR, Lloberas J, Camps M, Zierath JR, et al. 2003. Mitofusin-2 determines mitochondrial network architecture and mitochondrial metabolism. A novel regulatory mechanism altered in obesity. J Biol Chem. 278(19):17190–17197. doi:https://doi.org/10.1074/jbc.M212754200
- Barber AF, Erion R, Holmes TC, Sehgal A. 2016. Circadian and feeding cues integrate to drive rhythms of physiology in Drosophila insulin-producing cells. Genes Dev. 30(23):2596–2606. doi:https://doi.org/10.1101/gad.288258.116
- Barnea B, Madar Z, Froy O. 2009. High-fat diet delays and fasting advances the circadian expression of adiponectin signaling components in mouse liver. Endocrinology. 150(1):161–168. doi:https://doi.org/10.1210/en.2008-0944
- Begemann K, Neumann A-M OH. 2020. Regulation and function of extra-SCN circadian oscillators in the brain. Acta Physiologica. 229(1):e13446. doi:https://doi.org/10.1111/apha.13446
- Benavides A, Siches M, Llobera M. 1998. Circadian rhythms of lipoprotein lipase and hepatic lipase activities in intermediate metabolism of adult rat. Am J Physiol. 275(3):R811–817. doi:https://doi.org/10.1152/ajpregu.1998.275.3.R811
- Benito J, Zheng H, Ng FS, Hardin PE. 2007. Transcriptional feedback loop regulation, function, and ontogeny in Drosophila. Cold Spring Harb Symp Quant Biol. 72:437–444. doi:https://doi.org/10.1101/sqb.2007.72.009
- Bi J, Sehgal A, Williams JA, Wang Y-F. 2018. Wolbachia affects sleep behavior in Drosophila melanogaster. J Insect Physiol. 107:81–88. doi:https://doi.org/10.1016/j.jinsphys.2018.02.011
- Brown SA, Kowalska E, Dallmann R. 2012. (Re)inventing the circadian feedback loop. Dev Cell. 22(3):477–487. doi:https://doi.org/10.1016/j.devcel.2012.02.007
- Buijs RM, Scheer FA, Kreier F, Yi C, Bos N, Goncharuk VD, Kalsbeek A. 2006. Organization of circadian functions: interaction with the body. Prog Brain Res. 153:341–360. doi:https://doi.org/10.1016/S0079-6123(06)53020-1
- Bur IM, Cohen-Solal AM, Carmignac D, Abecassis P-Y, Chauvet N, Martin AO, van der Horst GTJ, Robinson ICAF, Maurel P, Mollard P, et al. 2009. The circadian clock components CRY1 and CRY2 are necessary to sustain sex dimorphism in mouse liver metabolism. J Biol Chem. 284(14):9066–9073. doi:https://doi.org/10.1074/jbc.M808360200
- Cantó C, Gerhart-Hines Z, Feige JN, Lagouge M, Noriega L, Milne JC, Elliott PJ, Puigserver P, Auwerx J. 2009. AMPK regulates energy expenditure by modulating NAD+ metabolism and SIRT1 activity. Nature. 458(7241):1056–1060. doi:https://doi.org/10.1038/nature07813
- Cassone VM, Stephan FK. 2002. Central and peripheral regulation of feeding and nutrition by the mammalian circadian clock: implications for nutrition during manned space flight. Nutrition. 18(10):814–819. doi:https://doi.org/10.1016/S0899-9007(02)00937-1
- Cavanaugh DJ, Geratowski JD, Wooltorton JRA, Spaethling JM, Hector CE, Zheng X, Johnson EC, Eberwine JH, Sehgal A. 2014. Identification of a circadian output circuit for rest: activity rhythms in Drosophila. Cell. 157(3):689–701. doi:https://doi.org/10.1016/j.cell.2014.02.024
- Cedernaes J, Huang W, Ramsey KM, Waldeck N, Cheng L, Marcheva B, Omura C, Kobayashi Y, Peek CB, Levine DC, et al. 2019. Transcriptional basis for rhythmic control of hunger and metabolism within the AgRP neuron. Cell Metab. 29(5):1078–1091.e5. doi:https://doi.org/10.1016/j.cmet.2019.01.023
- Challet E. 2010. Interactions between light, mealtime and calorie restriction to control daily timing in mammals. J Comp Physiol B. 180(5):631–644. doi:https://doi.org/10.1007/s00360-010-0451-4.
- Cho E, Kwon M, Jung J, Kang DH, Jin S, Choi SE, Kang Y, Kim EY. 2019. AMP-activated protein kinase regulates circadian rhythm by affecting CLOCK in Drosophila. J Neurosci. 39(18):3537–3550. doi:https://doi.org/10.1523/JNEUROSCI.2344-18.2019
- Cyran SA, Buchsbaum AM, Reddy KL, Lin MC, Glossop NR, Hardin PE, Young MW, Storti RV, Blau J. 2003. vrille, Pdp1, and Clock form a second feedback loop in the Drosophila circadian clock. Cell. 112:329–341. doi:https://doi.org/10.1016/S0092-8674(03)00074-6
- Cyran SA, Yiannoulos G, Buchsbaum AM, Saez L, Young MW, Blau J. 2005. The double-time protein kinase regulates the subcellular localization of the Drosophila clock protein period. J Neurosci. 25(22):5430–5437. doi:https://doi.org/10.1523/JNEUROSCI.0263-05.2005
- Damiola F, Le Minh N, Preitner N, Kornmann B, Fleury-Olela F, Schibler U. 2000. Restricted feeding uncouples circadian oscillators in peripheral tissues from the central pacemaker in the suprachiasmatic nucleus. Genes Dev. 14(23):2950–2961. doi:https://doi.org/10.1101/gad.183500
- Darlington TK, Wager-Smith K, Ceriani MF, Staknis D, Gekakis N, Steeves TD, Weitz CJ, Takahashi JS, Kay SA. 1998. Closing the circadian loop: CLOCK-induced transcription of its own inhibitors per and tim. Science. 280(5369):1599–1603. doi:https://doi.org/10.1126/science.280.5369.1599
- de Goede P, Sen S, Su Y, Foppen E, Poirel V-J, Challet E, Kalsbeek A. 2018. An Ultradian feeding schedule in rats affects metabolic gene expression in liver, brown adipose tissue and skeletal muscle with only mild effects on circadian clocks. Int J Mol Sci. 19:10. doi:https://doi.org/10.3390/ijms19103171
- Debruyne JP, Noton E, Lambert CM, Maywood ES, Weaver DR, Reppert SM. 2006. A clock shock: mouse CLOCK is not required for circadian oscillator function. Neuron. 50(3):465–477. doi:https://doi.org/10.1016/j.neuron.2006.03.041
- DeBruyne JP, Weaver DR, Reppert SM. 2007. Peripheral circadian oscillators require CLOCK. Curr Biol. 17(14):R538–539. doi:https://doi.org/10.1016/j.cub.2007.05.067
- DiAngelo JR, Erion R, Crocker A, Sehgal A. 2011. The central clock neurons regulate lipid storage in Drosophila. PLoS ONE. 6(5):e19921. doi:https://doi.org/10.1371/journal.pone.0019921
- Dreyer AP, Martin MM, Fulgham CV, Jabr DA, Bai L, Beshel J, Cavanaugh DJ. 2019. A circadian output center controlling feeding: fasting rhythms in Drosophila. PLoS Genet. 15(11):e1008478. doi:https://doi.org/10.1371/journal.pgen.1008478
- Dunlap JC, Loros JJ. 2017. Making time: conservation of biological clocks from fungi to animals. Microbiol Spectr. 5:3. doi:https://doi.org/10.1128/microbiolspec.FUNK-0039-2016
- Dyar KA, Ciciliot S, Wright LE, Biensø RS, Tagliazucchi GM, Patel VR, Forcato M, Paz MIP, Gudiksen A, Solagna F, et al. 2014. Muscle insulin sensitivity and glucose metabolism are controlled by the intrinsic muscle clock. Mol Metab. 3(1):29–41. doi:https://doi.org/10.1016/j.molmet.2013.10.005
- Dyar KA, Hubert MJ, Mir AA, Ciciliot S, Lutter D, Greulich F, Quagliarini F, Kleinert M, Fischer K, Eichmann TO, et al. 2018a. Transcriptional programming of lipid and amino acid metabolism by the skeletal muscle circadian clock. PLoS Biol. 16(8):e2005886. doi:https://doi.org/10.1371/journal.pbio.2005886
- Dyar KA, Lutter D, Artati A, Ceglia NJ, Liu Y, Armenta D, Jastroch M, Schneider S, de Mateo S, Cervantes M, et al. 2018b. Atlas of circadian metabolism reveals system-wide coordination and communication between clocks. Cell. 174(6):1571–1585.e11. doi:https://doi.org/10.1016/j.cell.2018.08.042
- Eckel-Mahan K, Sassone-Corsi P. 2009. Metabolism control by the circadian clock and vice versa. Nat Struct Mol Biol. 16(5):462–467. doi:https://doi.org/10.1038/nsmb.1595
- Eckel-Mahan KL, Patel VR, Mateo SD. 2013. Reprogramming of the circadian clock by nutritional challenge. Cell. 155(7):1464–1478. doi:https://doi.org/10.1016/j.cell.2013.11.034
- Edgar RS, Green EW, Zhao Y, van Ooijen G, Olmedo M, Qin X, Xu Y, Pan M, Valekunja UK, Feeney KA, et al. 2012. Peroxiredoxins are conserved markers of circadian rhythms. Nature. 485(7399):459–464. doi:https://doi.org/10.1038/nature11088
- Emery IF, Noveral JM, Jamison CF, Siwicki KK. 1997. Rhythms of Drosophila period gene expression in culture. Proc Natl Acad Sci USA. 94(8):4092–4096. doi:https://doi.org/10.1073/pnas.94.8.4092
- Erion R, King AN, Wu G, Hogenesch JB, Sehgal A. 2016. Neural clocks and Neuropeptide F/Y regulate circadian gene expression in a peripheral metabolic tissue. Elife. 5. doi:https://doi.org/10.7554/eLife.13552
- Feillet C, Ripperger J, Magnone M, Dulloo A, Albrecht U, Challet E. 2006. Lack of food anticipation in Per2 mutant mice. Curr Biol. 16:2016–2022. doi:https://doi.org/10.1016/j.cub.2006.08.053
- Frese T, Bazwinsky I, Mühlbauer E, Peschke E. 2007. Circadian and age-dependent expression patterns of GLUT2 and glucokinase in the pancreatic beta-cell of diabetic and nondiabetic rats. Horm Metab Res. 39(8):567–574. doi:https://doi.org/10.1055/s-2007-984471
- Fuller PM, Lu J, Saper CB. 2009. Standards of evidence in chronobiology: A response. J Circadian Rhythms. 7:9. doi:https://doi.org/10.1186/1740-3391-7-9
- García-Gaytán AC, Miranda-Anaya M, Turrubiate I, López-De Portugal L, Bocanegra-Botello GN, López-Islas A, Díaz-Muñoz M, Méndez I. 2020. Synchronization of the circadian clock by time-restricted feeding with progressive increasing calorie intake. Resemblances and differences regarding a sustained hypocaloric restriction. Sci Rep. 10(1):10036. doi:https://doi.org/10.1038/s41598-020-66538-0
- Giebultowicz JM, Hege DM. 2002. Circadian clock in Malpighian tubules. Nature. 386:664. doi:https://doi.org/10.1038/386664a0
- Glossop NRJ, Hardin PE. 2002. Central and peripheral circadian oscillator mechanisms in flies and mammals. J Cell Sci. 115(Pt 17):3369–3377. PMID: 12154068
- Glossop NRJ, Houl JH, Zheng H, Ng FS, Dudek SM, Hardin PE. 2003. VRILLE feeds back to control circadian transcription of Clock in the Drosophila circadian oscillator. Neuron. 37(2):249–261. doi:https://doi.org/10.1016/S0896-6273(03)00002-3
- Gomes LC, Scorrano L. 2011. Mitochondrial elongation during autophagy: a stereotypical response to survive in difficult times. Autophagy. 7(10):1251–1253. doi:https://doi.org/10.4161/auto.7.10.16771
- Gong C, Li C, Qi X, Song Z, Wu J, Hughes ME, Li X. 2015. The daily rhythms of mitochondrial gene expression and oxidative stress regulation are altered by aging in the mouse liver. Chronobiol Int. 32(9):1254–1263. doi:https://doi.org/10.3109/07420528.2015.1085388
- Grant D, Yin L, Collins JL, Parks DJ, Orband-Miller LA, Wisely GB, Joshi S, Lazar MA, Willson TM, Zuercher WJ. 2010. GSK4112, a Small molecule chemical probe for the cell biology of the nuclear heme receptor Rev-erbα. ACS Chem Biol. 5(10):925–932. doi:https://doi.org/10.1021/cb100141y
- Greenwell BJ, Trott AJ, Beytebiere JR, Pao S, Bosley A, Beach E, Finegan P, Hernandez C, Menet JS. 2019. Rhythmic food intake drives rhythmic gene expression more potently than the hepatic circadian clock in mice. Cell Rep. 27(3):649–657.e5. doi:https://doi.org/10.1016/j.celrep.2019.03.064
- Griffin EA, Staknis D, Weitz CJ. 1999. Light-independent role of CRY1 and CRY2 in the mammalian circadian clock. Science. 286(5440):768–771. doi:https://doi.org/10.1126/science.286.5440.768
- Guilding C, Hughes ATL, Brown TM, Namvar S, Piggins HD. 2009. A riot of rhythms: neuronal and glial circadian oscillators in the mediobasal hypothalamus. Mol Brain. 2:28. doi:https://doi.org/10.1186/1756-6606-2-28
- Guillaumond F, Dardente H, Giguère V, Cermakian N. 2005. Differential control of Bmal1 circadian transcription by REV-ERB and ROR nuclear receptors. J Biol Rhythms. 20(5):391–403. doi:https://doi.org/10.1177/0748730405277232
- Hara R, Wan K, Wakamatsu H, Aida R, Moriya T, Akiyama M, Shibata S. 2001. Restricted feeding entrains liver clock without participation of the suprachiasmatic nucleus. Genes Cells. 6(3):269–278. doi:https://doi.org/10.1046/j.1365-2443.2001.00419.x
- Hardin PE. 2005. The circadian time keeping system of Drosophila. Curr Biol. 15. doi:https://doi.org/10.1016/j.cub.2005.08.019
- Harfmann BD, Schroder EA, Kachman MT, Hodge BA, Zhang X, Esser KA. 2016. Muscle-specific loss of Bmal1 leads to disrupted tissue glucose metabolism and systemic glucose homeostasis. Skelet Muscle. 6:12. doi:https://doi.org/10.1186/s13395-016-0082-x
- Hastings MH, Maywood ES, Brancaccio M. 2019. The mammalian circadian timing system and the suprachiasmatic nucleus as its pacemaker. Biology (Basel). 8:1. doi:https://doi.org/10.3390/biology8010013
- He C, Cong X, Zhang R, Wu D, An C, Zhao Z. 2013. Regulation of circadian locomotor rhythm by neuropeptide Y-like system in Drosophila melanogaster. Insect Mol Biol. 22(4):376–388. doi:https://doi.org/10.1111/imb.12027
- Hegazi S, Lowden C, Rios Garcia J, Cheng AH, Obrietan K, Levine JD, Cheng H-YM. 2019. A symphony of signals: intercellular and intracellular signaling mechanisms underlying circadian timekeeping in mice and flies. Int J Mol Sci. 20:9. doi:https://doi.org/10.3390/ijms20092363
- Helfrich-Förster C. 1997. Development of pigment-dispersing hormone-immunoreactive neurons in the nervous system of Drosophila melanogaster. J Comp Neurol. 380. doi:https://doi.org/10.1002/(sici)1096-9861(19970414)380:3<335::aid-cne4>3.0.co;2-3
- Helfrich-Förster C, Shafer OT, Wülbeck C, Grieshaber E, Rieger D, Taghert P. 2007. Development and morphology of the clock-gene-expressing lateral neurons of Drosophila melanogaster. J Comp Neurol. 500(1):47–70. doi:https://doi.org/10.1002/cne.21146
- Helfrich-Förster C, Täuber M, Park JH, Mühlig-Versen M, Schneuwly S, Hofbauer A. 2000. Ectopic expression of the neuropeptide pigment-dispersing factor alters behavioral rhythms in Drosophila melanogaster. J Neurosci. 20(9):3339–3353. doi:https://doi.org/10.1523/JNEUROSCI.20-09-03339.2000
- Hermes ML, Coderre EM, Buijs RM, Renaud LP. 1996. GABA and glutamate mediate rapid neurotransmission from suprachiasmatic nucleus to hypothalamic paraventricular nucleus in rat. J Physiol. 496(Pt 3):749–757. doi:https://doi.org/10.1113/jphysiol.1996.sp021724
- Hirayama J, Sahar S, Grimaldi B, Tamaru T, Takamatsu K, Nakahata Y, Sassone-Corsi P. 2007. CLOCK-mediated acetylation of BMAL1 controls circadian function. Nature. 450(7172):1086–1090. doi:https://doi.org/10.1038/nature06394
- Hirota T, Fukada Y. 2004. Resetting mechanism of central and peripheral circadian clocks in mammals. Zool Sci. 21(4):359–368. doi:https://doi.org/10.2108/zsj.21.359
- Hirota T, Lee JW, St John PC, Sawa M, Iwaisako K, Noguchi T, Pongsawakul PY, Sonntag T, Welsh DK, Brenner DA, et al. 2012. Identification of small molecule activators of cryptochrome. Science. 337(6098):1094–1097. doi:https://doi.org/10.1126/science.1223710
- Hurley JM, Loros JJ, Dunlap JC. 2016 10. Circadian oscillators: around the transcription-translation feedback loop and on to output. Trends Biochem Sci. 41(10):834–846. doi:https://doi.org/10.1016/j.tibs.2016.07.009.
- Iijima M, Yamaguchi S, van der Horst G, Bonnefont X, Okamura H, Shibata S. 2005. Altered food-anticipatory activity rhythm in Cryptochrome-deficient mice. Neurosci Res. 52:166–173. doi:https://doi.org/10.1016/j.neures.2005.03.003
- Ito C, Tomioka K. 2016. Heterogeneity of the peripheral circadian systems in Drosophila melanogaster: a review. Front Physiol. 7:1–7. doi:https://doi.org/10.3389/fphys.2016.00008
- Jhanwar-Uniyal M, Beck B, Burlet C, Leibowitz SF. 1990. Diurnal rhythm of neuropeptide Y-like immunoreactivity in the suprachiasmatic, arcuate and paraventricular nuclei and other hypothalamic sites. Brain Res. 536(1):331–334. doi:https://doi.org/10.1016/0006-8993(90)90045-D
- Johard HAD, Yoishii T, Dircksen H, Cusumano P, Rouyer F, Helfrich-Förster C, Nässel DR. 2009. Peptidergic clock neurons in Drosophila: ion transport peptide and short neuropeptide F in subsets of dorsal and ventral lateral neurons. J Comp Neurol. 516(1):59–73. doi:https://doi.org/10.1002/cne.22099
- Kalsbeek A, Buijs RM, van Heerikhuize JJ, Arts M, van der Woude TP. 1992. Vasopressin-containing neurons of the suprachiasmatic nuclei inhibit corticosterone release. Brain Res. 580(1–2):62–67. doi:https://doi.org/10.1016/0006-8993(92)90927-2
- Kalsbeek A, Fliers E. 2013. Daily regulation of hormone profiles. Handb Exp Pharmacol. 217:185–226. doi:https://doi.org/10.1007/978-3-642-25950-0_8
- Kalsbeek A, Foppen E, Schalij I, Van Heijningen C, van der Vliet J, Fliers E, Buijs RM. 2008. Circadian control of the daily plasma glucose rhythm: an interplay of GABA and glutamate. PLoS ONE. 3(9):e3194. doi:https://doi.org/10.1371/journal.pone.0003194
- Kalsbeek A, La Fleur S, Van Heijningen C, Buijs RM. 2004. Suprachiasmatic GABAergic inputs to the paraventricular nucleus control plasma glucose concentrations in the rat via sympathetic innervation of the liver. J Neurosci. 24(35):7604–7613. doi:https://doi.org/10.1523/JNEUROSCI.5328-03.2004
- Kalsbeek A, van Heerikhuize JJ, Wortel J, Buijs RM. 1996. A diurnal rhythm of stimulatory input to the hypothalamo-pituitary-adrenal system as revealed by timed intrahypothalamic administration of the vasopressin V1 antagonist. J Neurosci. 16(17):5555–5565. doi:https://doi.org/10.1523/JNEUROSCI.16-17-05555.1996
- Kaneko M, Hall JC. 2000. Neuroanatomy of cells expressing clock genes in Drosophila: transgenic manipulation of the period and timeless genes to mark the perikarya of circadian pacemaker neurons and their projections. J Comp Neurol. 422(1):66–94. doi:https://doi.org/10.1002/(SICI)1096-9861(20000619)422:1<66::AID-CNE5>3.0.CO;2-2
- Kaneko M, Helfrich-Förster C, Hall JC. 1997. Spatial and temporal expression of the period and timeless genes in the developing nervous system of Drosophila: newly identified pacemaker candidates and novel features of clock gene product cycling. J Neurosci. 17(17):6745–6760. doi:https://doi.org/10.1523/JNEUROSCI.17-17-06745.1997
- Kapahi P, Chen D, Rogers AN, Katewa SD, Li PW-L, Thomas EL, Kockel L. 2010. With TOR, less is more: a key role for the conserved nutrient-sensing TOR pathway in aging. Cell Metab. 11(6):453–465. doi:https://doi.org/10.1016/j.cmet.2010.05.001
- Katewa SD, Akagi K, Bose N, Rakshit K, Camarella T, Zheng X, Hall D, Davis S, Nelson CS, Brem RB, et al. 2016. Peripheral circadian clocks mediate dietary restriction-dependent changes in lifespan and fat metabolism in Drosophila. Cell Metab. 23(1):143–154. doi:https://doi.org/10.1016/j.cmet.2015.10.014
- Katewa SD, Kapahi P. 2010. Dietary restriction and aging. Aging Cell. 9(2):105–112. doi:https://doi.org/10.1111/j.1474-9726.2010.00552.x
- Kijak E, Pyza E. 2017. TOR signaling pathway and autophagy are involved in the regulation of circadian rhythms in behavior and plasticity of L2 interneurons in the brain of Drosophila melanogaster. PLoS ONE. 12(2):e0171848. doi:https://doi.org/10.1371/journal.pone.0171848
- Klarsfeld A, Malpel S, Michard-Vanhee C, Picot M, Chelot E, Rouyer F. 2004. Novel features of cryptochrome-mediated photoreception in the brain circadian clock of Drosophila. J Neurosci. 24. doi:https://doi.org/10.1523/JNEUROSCI.3661-03.2004
- Kloss B, Price JL, Saez L, Blau J, Rothenfluh A, Wesley CS, Young MW. 1998. The Drosophila clock gene double-time encodes a protein closely related to human casein kinase I epsilon. Cell. 94(1):97–107. doi:https://doi.org/10.1016/S0092-8674(00)81225-8
- Knippschild U, Gocht A, Wolff S, Huber N, Löhler J, Stöter M. 2005. The casein kinase 1 family: participation in multiple cellular processes in eukaryotes. Cell Signal. 17(6):675–689. doi:https://doi.org/10.1016/j.cellsig.2004.12.011
- Kohsaka A, Laposky AD, Ramsey KM, Estrada C, Joshu C, Kobayashi Y, Turek FW, Bass J. 2007. High-fat diet disrupts behavioral and molecular circadian rhythms in mice. Cell Metab. 6(5):414–421. doi:https://doi.org/10.1016/j.cmet.2007.09.006
- Kondratov RV, Chernov MV, Kondratova AA, Gorbacheva VY, Gudkov AV, Antoch MP. 2003. BMAL1-dependent circadian oscillation of nuclear CLOCK: posttranslational events induced by dimerization of transcriptional activators of the mammalian clock system. Genes Dev. 17(15):1921–1932. doi:https://doi.org/10.1101/gad.1099503
- Konopka RJ, Benzer S. 1971. Clock mutants of Drosophila melanogaster. Proc Natl Acad Sci USA. 68. doi:https://doi.org/10.1073/pnas.68.9.2112
- Koronowski KB, Kinouchi K, Welz P-S, Smith JG, Zinna VM, Shi J, Samad M, Chen S, Magnan CN, Kinchen JM, et al. 2019. Defining the independence of the liver circadian clock. Cell. 177(6):1448–1462.e14. doi:https://doi.org/10.1016/j.cell.2019.04.025
- Krishnaiah SY, Wu G, Altman BJ, Growe J, Rhoades SD, Coldren F, Venkataraman A, Olarerin-George AO, Francey LJ, Mukherjee S, et al. 2017. Clock regulation of metabolites reveals coupling between transcription and metabolism. Cell Metab. 25(4):961–974.e4. doi:https://doi.org/10.1016/j.cmet.2017.03.019
- Kumar N, Solt LA, Wang Y, Rogers PM, Bhattacharyya G, Kamenecka TM, Stayrook KR, Crumbley C, Floyd ZE, Gimble JM, et al. 2010. Regulation of adipogenesis by natural and synthetic REV-ERB ligands. Endocrinology. 151(7):3015–3025. doi:https://doi.org/10.1210/en.2009-0800
- Kume K, Zylka MJ, Sriram S, Shearman LP, Weaver DR, Jin X, Maywood ES, Hastings MH, Reppert SM. 1999. mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell. 98(2):193–205. doi:https://doi.org/10.1016/S0092-8674(00)81014-4
- Kwon I, Lee J, Chang SH, Jung NC, Lee BJ, Son GH, Kim K, Lee KH. 2006. BMAL1 shuttling controls transactivation and degradation of the CLOCK/BMAL1 heterodimer. Mol Cell Biol. 26(19):7318–7330. doi:https://doi.org/10.1128/MCB.00337-06
- Lamia KA, Papp SJ, Yu RT, Barish GD, Uhlenhaut NH, Jonker JW, Downes M, Evans RM. 2011. Cryptochromes mediate rhythmic repression of the glucocorticoid receptor. Nature. 480(7378):552–556. doi:https://doi.org/10.1038/nature10700
- Lamia KA, Sachdeva UM, DiTacchio L, Williams EC, Alvarez JG, Egan DF, Vasquez DS, Juguilon H, Panda S, Shaw RJ, et al. 2009. AMPK regulates the circadian clock by cryptochrome phosphorylation and degradation. Science. 326(5951):437–440. doi:https://doi.org/10.1126/science.1172156
- Lamia KA, Storch K-F, Weitz CJ. 2008. Physiological significance of a peripheral tissue circadian clock. Proc Natl Acad Sci USA. 105(39):15172–15177. doi:https://doi.org/10.1073/pnas.0806717105
- LeSauter J, Hoque N, Weintraub M, Pfaff DW, Silver R. 2009. Stomach ghrelin-secreting cells as food-entrainable circadian clocks. Proc Natl Acad Sci USA. 106(32):13582–13587. doi:https://doi.org/10.1073/pnas.0906426106
- Li A-J, Wiater MF, Oostrom MT, Smith BR, Wang Q, Dinh TT, Roberts BL, Jansen HT, Ritter S. 2012. Leptin-sensitive neurons in the arcuate nuclei contribute to endogenous feeding rhythms. Am J Physiol Regul Integr Comp Physiol. 302(11):R1313–1326. doi:https://doi.org/10.1152/ajpregu.00086.2012
- Li M-D, Li C-M WZ. 2012. The role of circadian clocks in metabolic disease. Yale J Biol Med. 85(3):387–401. PMID: 23012586
- Lin J-M, Schroeder A, Allada R. 2005. In vivo circadian function of casein kinase 2 phosphorylation sites in Drosophila PERIOD. J Neurosci. 25(48):11175–11183. doi:https://doi.org/10.1523/JNEUROSCI.2159-05.2005
- Lipton JO, Boyle LM, Yuan ED, Hochstrasser KJ, Chifamba FF, Nathan A, Tsai PT, Davis F, Sahin M. 2017. Aberrant proteostasis of BMAL1 underlies circadian abnormalities in a paradigmatic mTOR-opathy. Cell Rep. 20(4):868–880. doi:https://doi.org/10.1016/j.celrep.2017.07.008
- Magnone MC, Langmesser S, Bezdek AC, Tallone T, Rusconi S, Albrecht U. 2014. The Mammalian circadian clock gene per2 modulates cell death in response to oxidative stress. Front Neurol. 5:289. doi:https://doi.org/10.3389/fneur.2014.00289
- Maier B, Wendt S, Vanselow JT, Wallach T, Reischl S, Oehmke S, Schlosser A, Kramer A. 2009. A large-scale functional RNAi screen reveals a role for CK2 in the mammalian circadian clock. Genes Dev. 23(6):708–718. doi:https://doi.org/10.1101/gad.512209
- Marcheva B, Ramsey KM, Buhr ED, Kobayashi Y, Su H, Ko CH, Ivanova G, Omura C, Mo S, Vitaterna MH, et al. 2010. Disruption of the clock components CLOCK and BMAL1 leads to hypoinsulinaemia and diabetes. Nature. 466(7306):627–631. doi:https://doi.org/10.1038/nature09253
- Martin RJ, Stolz DJ, Buck DC. 1979. Diurnal changes in adipose and liver tissue metabolism of lean and obese Zucker rats. J Nutr. 109(3):412–417. doi:https://doi.org/10.1093/jn/109.3.412
- Martinek S, Inonog S, Manoukian AS, Young MW. 2001. A role for the segment polarity gene shaggy/GSK-3 in the Drosophila circadian clock. Cell. 105(6):769–779. doi:https://doi.org/10.1016/S0092-8674(01)00383-X
- Mattson MP, Allison DB, Fontana L, Harvie M, Longo VD, Malaisse WJ, Mosley M, Notterpek L, Ravussin E, Scheer FAJL, et al. 2014. Meal frequency and timing in health and disease. Proc Natl Acad Sci USA. 111(47):16647–16653. doi:https://doi.org/10.1073/pnas.1413965111
- Mauvoisin D, Atger F, Dayon L, Núñez Galindo A, Wang J, Martin E, Da Silva L, Montoliu I, Collino S, Martin FP, et al. 2017. Circadian and feeding rhythms orchestrate the diurnal liver acetylome. Cell Rep. 20(7):1729–1743. doi:https://doi.org/10.1016/j.celrep.2017.07.065
- Mauvoisin D, Wang J, Jouffe C, Martin E, Atger F, Waridel P, Quadroni M, Gachon F, Naef F. 2014. Circadian clock-dependent and -independent rhythmic proteomes implement distinct diurnal functions in mouse liver. Proc Natl Acad Sci USA. 111(1):167–172. doi:https://doi.org/10.1073/pnas.1314066111
- Méndez‐Hernández R, Escobar C, Buijs RM. 2020. Suprachiasmatic nucleus–arcuate nucleus axis: interaction between time and metabolism essential for health. Obesity. 28(S1):S10–S17. doi:https://doi.org/10.1002/oby.22774
- Meyer-Spasche A, Piggins HD. 2004. Vasoactive intestinal polypeptide phase-advances the rat suprachiasmatic nuclei circadian pacemaker in vitro via protein kinase A and mitogen-activated protein kinase. Neurosci Lett. 358(2):91–94. doi:https://doi.org/10.1016/j.neulet.2003.12.114
- Mistlberger RE. 2009. Food-anticipatory circadian rhythms: concepts and methods. Eur J Neurosci. 30(9):1718–1729. doi:https://doi.org/10.1111/j.1460-9568.2009.06965.x
- Mistlberger RE, Yamazaki S, Pendergast JS, Landry GJ, Takumi T, Nakamura W. 2008. Comment on Differential rescue of light- and food-entrainable circadian rhythms. Science. 322(5902):675. doi:https://doi.org/10.1126/science.1161284
- Mohawk JA, Green CB, Takahashi JS. 2012. Central and peripheral circadian clocks in mammals. Annu Rev Neurosci. 35:445–462. doi:https://doi.org/10.1146/annurev-neuro-060909-153128
- Morioka E, Oida M, Tsuchida T, Ikeda M. 2018. Nighttime activities and peripheral clock oscillations depend on Wolbachia endosymbionts in flies. Sci Rep. 8(1):15432. doi:https://doi.org/10.1038/s41598-018-33522-8
- Nagai K, Nishio T, Nakagawa H, Nakamura S, Fukuda Y. 1978. Effect of bilateral lesions of the suprachiasmatic nuclei on the circadian rhythm of food-intake. Brain Res. 142(2):384–389. doi:https://doi.org/10.1016/0006-8993(78)90648-0
- Nakahata Y, Kaluzova M, Grimaldi B, Sahar S, Hirayama J, Chen D, Guarente LP, Sassone-Corsi P. 2008. The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control. Cell. 134(2):329–340. doi:https://doi.org/10.1016/j.cell.2008.07.002
- Nakahata Y, Sahar S, Astarita G, Kaluzova M, Sassone-Corsi P. 2009. Circadian control of the NAD+ salvage pathway by CLOCK-SIRT1. Science. 324(5927):654–657. doi:https://doi.org/10.1126/science.1170803
- Neufeld-Cohen A, Robles MS, Aviram R, Manella G, Adamovich Y, Ladeuix B, Nir D, Rousso-Noori L, Kuperman Y, Golik M, et al. 2016. Circadian control of oscillations in mitochondrial rate-limiting enzymes and nutrient utilization by PERIOD proteins. Proc Natl Acad Sci USA. 113(12):E1673–E1682. doi:https://doi.org/10.1073/pnas.1519650113
- Oishi K, Miyazaki K, Ishida N. 2002. Functional CLOCK is not involved in the entrainment of peripheral clocks to the restricted feeding: entrainable expression of mPer2 and BMAL1 mRNAs in the heart of Clock mutant mice on Jcl: iCRbackground. Biochem Biophys Res Commun. 298(2):198–202. doi:https://doi.org/10.1016/S0006-291X(02)02427-0
- Oishi K, Shiota M, Sakamoto K, Kasamatsu M, Ishida N. 2004. Feeding is not a more potent Zeitgeber than the light-dark cycle in Drosophila. Neuroreport. 15(4):739–743. doi:https://doi.org/10.1097/00001756-200403220-00034
- Orozco-Solis R, Aguilar-Arnal L, Murakami M, Peruquetti R, Ramadori G, Coppari R, Sassone-Corsi P. 2016. The circadian clock in the ventromedial hypothalamus controls cyclic energy expenditure. Cell Metab. 23(3):467–478. doi:https://doi.org/10.1016/j.cmet.2016.02.003
- Oster H, Challet E, Ott V, Arvat E, de Kloet ER, Dijk D-J, Lightman S, Vgontzas A, Van Cauter E. 2017. The functional and clinical significance of the 24-hour rhythm of circulating glucocorticoids. Endocr Rev. 38(1):3–45. doi:https://doi.org/10.1210/er.2015-1080
- Owino S, Contreras-Alcantara S, Baba K, Tosini G. 2016. Melatonin signaling controls the daily rhythm in blood glucose levels independent of peripheral clocks. PLoS ONE. 11(1):e0148214. doi:https://doi.org/10.1371/journal.pone.0148214
- Panda S, Antoch MP, Miller BH, Su AI, Schook AB, Straume M, Schultz PG, Kay SA, Takahashi JS, Hogenesch JB. 2002. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell. 109(3):307–320. doi:https://doi.org/10.1016/S0092-8674(02)00722-5
- Park JH, Hall JC. 1998. Isolation and chronobiological analysis of a neuropeptide pigment-dispersing factor gene in Drosophila melanogaster. J Biol Rhythms. 13(3):219–228. doi:https://doi.org/10.1177/074873098129000066
- Paschos GK, Ibrahim S, Song W-L, Kunieda T, Grant G, Reyes TM, Bradfield CA, Vaughan CH, Eiden M, Masoodi M, et al. 2012. Obesity in mice with adipocyte-specific deletion of clock component Arntl. Nat Med. 18(12):1768–1777. doi:https://doi.org/10.1038/nm.2979
- Patel SA, Velingkaar N, Makwana K, Chaudhari A, Kondratov R. 2016. Calorie restriction regulates circadian clock gene expression through BMAL1 dependent and independent mechanisms. Sci Rep. 6(6):25970. doi:https://doi.org/10.1038/srep25970
- Patton AP, Hastings MH. 2018. The suprachiasmatic nucleus. Curr Biol. 28(15):R816–R822. doi:https://doi.org/10.1016/j.cub.2018.06.052
- Peckett AJ, Wright DC, Riddell MC. 2011. The effects of glucocorticoids on adipose tissue lipid metabolism. Metabolism. 60(11):1500–1510. doi:https://doi.org/10.1016/j.metabol.2011.06.012
- Pekovic-Vaughan V, Gibbs J, Yoshitane H, Yang N, Pathiranage D, Guo B, Sagami A, Taguchi K, Bechtold D, Loudon A, et al. 2014. The circadian clock regulates rhythmic activation of the NRF2/glutathione-mediated antioxidant defense pathway to modulate pulmonary fibrosis. Genes Dev. 28(6):548–560. doi:https://doi.org/10.1101/gad.237081.113
- Pendergast J, Wendroth R, Stenner R, Keil C, Yamazaki S. 2017. mPeriod2 (Brdm1) and other single Period mutant mice have normal food anticipatory activity. Sci Rep. 7:15510. doi:https://doi.org/10.1038/s41598-017-15332-6
- Pendergast JS, Nakamura W, Friday RC, Hatanaka F, Takumi T, Yamazaki S. 2009. Robust food anticipatory activity in BMAL1-deficient mice. PLoS ONE. 4(3):e4860. doi:https://doi.org/10.1371/journal.pone.0004860
- Perreau-Lenz S, Kalsbeek A, Pévet P, Buijs RM. 2004. Glutamatergic clock output stimulates melatonin synthesis at night. Eur J Neurosci. 19(2):318–324. doi:https://doi.org/10.1111/j.0953-816X.2003.03132.x
- Pezük P, Mohawk JA, Wang LA, Menaker M. 2012. Glucocorticoids as entraining signals for peripheral circadian oscillators. Endocrinology. 153(10):4775–4783. doi:https://doi.org/10.1210/en.2012-1486
- Plautz JD, Kaneko M, Hall JC, Kay SA. 1997. Independent photoreceptive circadian clocks throughout Drosophila. Science. 278(5343):1632–1635. doi:https://doi.org/10.1126/science.278.5343.1632
- Preitner N, Damiola F, Lopez-Molina L, Zakany J, Duboule D, Albrecht U, Schibler U. 2002. The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell. 110(2):251–260. doi:https://doi.org/10.1016/S0092-8674(02)00825-5
- Price JL, Blau J, Rothenfluh A, Abodeely M, Kloss B, Young MW. 1998. double-time is a novel Drosophila clock gene that regulates PERIOD protein accumulation. Cell. 94(1):83–95. doi:https://doi.org/10.1016/S0092-8674(00)81224-6
- Ramanathan C, Kathale ND, Liu D, Lee C, Freeman DA, Hogenesch JB, Cao R, Liu AC. 2018. mTOR signaling regulates central and peripheral circadian clock function. PLoS Genet. 14(5):e1007369. doi:https://doi.org/10.1371/journal.pgen.1007369
- Ramsey KM, Yoshino J, Brace CS, Abrassart D, Kobayashi Y, Marcheva B, Hong H-K, Chong JL, Buhr ED, Lee C, et al. 2009. Circadian clock feedback cycle through NAMPT-mediated NAD+ biosynthesis. Science. 324(5927):651–654. doi:https://doi.org/10.1126/science.1171641
- Reddy AB, Maywood ES, Karp NA, King VM, Inoue Y, Gonzalez FJ, Lilley KS, Kyriacou CP, Hastings MH. 2007. Glucocorticoid signaling synchronizes the liver circadian transcriptome. Hepatology. 45(6):1478–1488. doi:https://doi.org/10.1002/hep.21571
- Reinke H, Asher G. 2019. Crosstalk between metabolism and circadian clocks. Nat Rev Mol Cell Biol. 20(4):227–241. doi:https://doi.org/10.1038/s41580-018-0096-9
- Reinking J, Lam MMS, Pardee K, Sampson HM, Liu S, Yang P, Williams S, White W, Lajoie G, Edwards A, et al. 2005. The Drosophila nuclear receptor e75 contains heme and is gas responsive. Cell. 122(2):195–207. doi:https://doi.org/10.1016/j.cell.2005.07.005
- Reischl S, Kramer A. 2011. Kinases and phosphatases in the mammalian circadian clock. FEBS Lett. 585(10):1393–1399. doi:https://doi.org/10.1016/j.febslet.2011.02.038
- Renn SC, Park JH, Rosbash M, Hall JC, Taghert PH. 1999. A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila. Cell. 99(7):791–802. doi:https://doi.org/10.1016/s0092-8674(00)81676-1
- Rhoades SD, Nayak K, Zhang SL, Sehgal A, Weljie AM. 2018. Circadian- and light-driven metabolic rhythms in Drosophila melanogaster. J Biol Rhythms. 33(2):126–136. doi:https://doi.org/10.1177/0748730417753003
- Richards J, Gumz ML. 2012. Advances in understanding the peripheral circadian clocks. Faseb J. 26(9):3602–3613. doi:https://doi.org/10.1096/fj.12-203554
- Ruderman NB, Xu XJ, Nelson L, Cacicedo JM, Saha AK, Lan F, Ido Y. 2010. AMPK and SIRT1: a long-standing partnership? Am J Physiol Endocrinol Metab. 298(4):E751–760. doi:https://doi.org/10.1152/ajpendo.00745.2009
- Rudic RD, McNamara P, Curtis A-M, Boston RC, Panda S, Hogenesch JB, Fitzgerald GA. 2004. BMAL1 and CLOCK, two essential components of the circadian clock, are involved in glucose homeostasis. PLoS Biol. 2(11):e377. doi:https://doi.org/10.1371/journal.pbio.0020377
- Rutila JE, Suri V, Le M, So WV, Rosbash M, Hall JC. 1998. CYCLE is a second bHLH-PAS clock protein essential for circadian rhythmicity and transcription of Drosophila period and timeless. Cell. 93(5):805–814. doi:https://doi.org/10.1016/S0092-8674(00)81441-5
- Sadacca LA, Lamia KA, deLemos AS, Blum B, Weitz CJ. 2011. An intrinsic circadian clock of the pancreas is required for normal insulin release and glucose homeostasis in mice. Diabetologia. 54(1):120–124. doi:https://doi.org/10.1007/s00125-010-1920-8
- Saez L, Young MW. 1988. In situ localization of the per clock protein during development of Drosophila melanogaster. Mol Cell Biol. 8(12):5378–5385. doi:https://doi.org/10.1128/MCB.8.12.5378
- Sahar S, Masubuchi S, Eckel-Mahan K, Vollmer S, Galla L, Ceglia N, Masri S, Barth TK, Grimaldi B, Oluyemi O, et al. 2014. Circadian control of fatty acid elongation by SIRT1 protein-mediated deacetylation of acetyl-coenzyme A synthetase 1. J Biol Chem. 289(9):6091–6097. doi:https://doi.org/10.1074/jbc.M113.537191
- Sahar S, Zocchi L, Kinoshita C, Borrelli E, Sassone-Corsi P. 2010. Regulation of BMAL1 protein stability and circadian function by GSK3beta-mediated phosphorylation. PLoS ONE. 5(1):e8561. doi:https://doi.org/10.1371/journal.pone.0008561
- Sardon Puig L, Valera-Alberni M, Cantó C, Pillon NJ. 2018. Circadian rhythms and mitochondria: connecting the dots. Front Genet. 9:452. doi:https://doi.org/10.3389/fgene.2018.00452
- Schäbler S, Amatobi KM, Horn M, Rieger D, Helfrich-Förster C, Mueller MJ, Wegener C, Fekete A. 2020. Loss of function in the Drosophila clock gene period results in altered intermediary lipid metabolism and increased susceptibility to starvation. Cell Mol Life Sci. 1–18. doi:https://doi.org/10.1007/s00018-019-03441-6
- Schmitt K, Grimm A, Dallmann R, Oettinghaus B, Restelli LM, Witzig M, Ishihara N, Mihara K, Ripperger JA, Albrecht U, et al. 2018. Circadian control of DRP1 activity regulates mitochondrial dynamics and bioenergetics. Cell Metab. 27(3):657–666.e5. doi:https://doi.org/10.1016/j.cmet.2018.01.011
- Scott RC, Schuldiner O, Neufeld TP. 2004. Role and regulation of starvation-induced autophagy in the Drosophila fat body. Dev Cell. 7(2):167–178. doi:https://doi.org/10.1016/j.devcel.2004.07.009
- Seay DJ, Thummel CS. 2011. The circadian clock, light, and cryptochrome regulate feeding and metabolism in Drosophila. J Biol Rhythms. 26(6):497–506. doi:https://doi.org/10.1177/0748730411420080
- Sehgal A, Price JL, Man B, Young MW. 1994. Loss of circadian behavioral rhythms and per RNA oscillations in the Drosophila mutant timeless. Science. 263(5153):1603–1606. doi:https://doi.org/10.1126/science.8128246
- Selkrig J, Mohammad F, Ng SH, Chua JY, Tumkaya T, Ho J, Chiang YN, Rieger D, Pettersson S, Helfrich-Förster C, et al. 2018. The Drosophila microbiome has a limited influence on sleep, activity, and courtship behaviors. Sci Rep. 8(1):10646. doi:https://doi.org/10.1038/s41598-018-28764-5
- Sena LA, Chandel NS. 2012. Physiological roles of mitochondrial reactive oxygen species. Mol Cell. 48(2):158–167. doi:https://doi.org/10.1016/j.molcel.2012.09.025
- Sengupta S, Yang G, O’Donnell JC, Hinson MD, McCormack SE, Falk MJ, La P, Robinson MB, Williams ML, Yohannes MT, et al. 2016. The circadian gene Rev-erbα improves cellular bioenergetics and provides preconditioning for protection against oxidative stress. Free Radic Biol Med. 93:177–189. doi:https://doi.org/10.1016/j.freeradbiomed.2016.02.004
- Shafer OT, Helfrich-Förster C, Renn SCP, Taghert PH. 2006. Reevaluation of Drosophila melanogaster’s neuronal circadian pacemakers reveals new neuronal classes. J Comp Neurol. 498(2):180–193. doi:https://doi.org/10.1002/cne.21021
- Sharma VK, Chandrashekaran MK. 2005. Zeitgebers (time cues) for biological clocks. Curr Sci. 89(7):1136–1146. https://www.jstor.org/stable/24110966
- Shi M, Zheng X. 2013. Interactions between the circadian clock and metabolism: there are good times and bad times. Acta Biochim Biophys Sin (Shanghai). 45(1):61–69. doi:https://doi.org/10.1093/abbs/gms110
- Shimba S, Ishii N, Ohta Y, Ohno T, Watabe Y, Hayashi M, Wada T, Aoyagi T, Tezuka M. 2005. Brain and muscle Arnt-like protein-1 (BMAL1), a component of the molecular clock, regulates adipogenesis. Proc Natl Acad Sci USA. 102(34):12071–12076. doi:https://doi.org/10.1073/pnas.0502383102
- Shimizu H, Arima H, Watanabe M, Goto M, Banno R, Sato I, Ozaki N, Nagasaki H, Oiso Y. 2008. Glucocorticoids increase neuropeptide Y and agouti-related peptide gene expression via adenosine monophosphate-activated protein kinase signaling in the arcuate nucleus of rats. Endocrinology. 149(9):4544–4553. doi:https://doi.org/10.1210/en.2008-0229
- Shostak A, Husse J, Oster H. 2013. Circadian regulation of adipose function. Adipocyte. 2(4):201–206. doi:https://doi.org/10.4161/adip.26007
- Stanewsky R, Kaneko M, Emery P, Beretta B, Wager-Smith K, Kay SA, Rosbash M, Hall JC. 1998. The cryb mutation identifies cryptochrome as a circadian photoreceptor in Drosophila. Cell. 95(5):681–692. doi:https://doi.org/10.1016/S0092-8674(00)81638-4
- Stanley BG, Leibowitz SF. 1985. Neuropeptide Y injected in the paraventricular hypothalamus: a powerful stimulant of feeding behavior. Proc Natl Acad Sci USA. 82(11):3940–3943. doi:https://doi.org/10.1073/pnas.82.11.3940
- Storch K, Weitz C. 2009. Daily rhythms of food-anticipatory behavioral activity do not require the known circadian clock. Proc Natl Acad Sci USA. 106:6808–6813. doi:https://doi.org/10.1073/pnas.0902063106
- Sukumaran S, Xue B, Jusko WJ, Dubois DC, Almon RR. 2010. Circadian variations in gene expression in rat abdominal adipose tissue and relationship to physiology. Physiol Genomics. 42A(2):141–152. doi:https://doi.org/10.1152/physiolgenomics.00106.2010
- Sun C, Zhang F, Ge X, Yan T, Chen X, Shi X, Zhai Q. 2007. SIRT1 improves insulin sensitivity under insulin-resistant conditions by repressing PTP1B. Cell Metab. 6(4):307–319. doi:https://doi.org/10.1016/j.cmet.2007.08.014
- Suzuki M, Shimomura Y, Satoh Y. 1983. Diurnal changes in lipolytic activity of isolated fat cells and their increased responsiveness to epinephrine and theophylline with meal feeding in rats. J Nutr Sci Vitaminol. 29(4):399–411. doi:https://doi.org/10.3177/jnsv.29.399
- Tsuchiya Y, Akashi M, Matsuda M, Goto K, Miyata Y, Node K, Nishida E. 2009. Involvement of the protein kinase CK2 in the regulation of mammalian circadian rhythms. Sci Signal. 2. doi:https://doi.org/10.1126/scisignal.2000305
- Tsutsumi K, Inoue Y, Kondo Y. 2002. The relationship between lipoprotein lipase activity and respiratory quotient of rats in circadian rhythms. Biol Pharm Bull. 25(10):1360–1363. doi:https://doi.org/10.1248/bpb.25.1360
- Turek FW, Joshu C, Kohsaka A, Lin E, Ivanova G, McDearmon E, Laposky A, Losee-Olson S, Easton A, Jensen DR, et al. 2005. Obesity and metabolic syndrome in circadian Clock mutant mice. Science. 308(5724):1043–1045. doi:https://doi.org/10.1126/science.1108750
- Ulgherait M, Chen A, Oliva MK, Kim HX, Canman JC, Ja WW, Shirasu-Hiza M. 2016. Dietary restriction extends the lifespan of circadian mutants tim and per. Cell Metab. 24(6):763–764. doi:https://doi.org/10.1016/j.cmet.2016.11.002
- Um JH, Yang S, Yamazaki S, Kang H, Viollet B, Foretz M, Chung JH. 2007. Activation of 5ʹ-AMP-activated kinase with diabetes drug metformin induces casein kinase I epsilon (CKI epsilon)-dependent degradation of clock protein mPer2. J Biol Chem. 282(29):20794–20798. doi:https://doi.org/10.1074/jbc.C700070200
- Vieira E, Marroquí L, Batista TM, Caballero-Garrido E, Carneiro EM, Boschero AC, Nadal A, Quesada I. 2012. The clock gene Rev-erbα regulates pancreatic β-cell function: modulation by leptin and high-fat diet. Endocrinology. 153(2):592–601. doi:https://doi.org/10.1210/en.2011-1595
- Vitaterna MH, King DP, Chang AM, Kornhauser JM, Lowrey PL, McDonald JD, Dove WF, Pinto LH, Turek FW, Takahashi JS. 1994. Mutagenesis and mapping of a mouse gene, Clock, essential for circadian behavior. Science. 264(5159):719–725. doi:https://doi.org/10.1126/science.8171325
- Wang TA, Yu YV, Govindaiah G, Ye X, Artinian L, Coleman TP, Sweedler JV, Cox CL, Gillette MU. 2012. Circadian rhythm of redox state regulates excitability in suprachiasmatic nucleus neurons. Science. 337(6096):839–842. doi:https://doi.org/10.1126/science.1222826
- Wiater MF, Mukherjee S, Li A-J, Dinh TT, Rooney EM, Simasko SM, Ritter S. 2011. Circadian integration of sleep-wake and feeding requires NPY receptor-expressing neurons in the mediobasal hypothalamus. Am J Physiol Regul Integr Comp Physiol. 301(5):R1569–1583. doi:https://doi.org/10.1152/ajpregu.00168.2011
- Xu K, DiAngelo JR, Hughes ME, Hogenesch JB, Sehgal A. 2011. The circadian clock interacts with metabolic physiology to influence reproductive fitness. Cell Metab. 13(6):639–654. doi:https://doi.org/10.1016/j.cmet.2011.05.001
- Xu K, Zheng X, Sehgal A. 2008. Regulation of feeding and metabolism by neuronal and peripheral clocks in Drosophila. Cell Metab. 8(4):289–300. doi:https://doi.org/10.1016/j.cmet.2008.09.006
- Xu Y-Q, Zhang D, Jin T, Cai D-J, Wu Q, Lu Y, Liu J, Klaassen CD. 2012. Diurnal variation of hepatic antioxidant gene expression in mice. PLoS ONE. 7(8):e44237. doi:https://doi.org/10.1371/journal.pone.0044237
- Yaku K, Okabe K, Nakagawa T. 2018. NAD metabolism: implications in aging and longevity. Ageing Res Rev. 47:1–17. doi:https://doi.org/10.1016/j.arr.2018.05.006
- Yamajuku D, Inagaki T, Haruma T, Okubo S, Kataoka Y, Kobayashi S, Ikegami K, Laurent T, Kojima T, Noutomi K, et al. 2012. Real-time monitoring in three-dimensional hepatocytes reveals that insulin acts as a synchronizer for liver clock. Sci Rep. 2:439. doi:https://doi.org/10.1038/srep00439
- Yamamoto H, Nagai K, Nakagawa H. 1987. Role of SCN in daily rhythms of plasma glucose, FFA, insulin and glucagon. Chronobiol Int. 4(4):483–491. doi:https://doi.org/10.3109/07420528709078539
- Yamanaka A, Beuckmann CT, Willie JT, Hara J, Tsujino N, Mieda M, Tominaga M, Yagami K, Sugiyama F, Goto K, et al. 2003. Hypothalamic orexin neurons regulate arousal according to energy balance in mice. Neuron. 38(5):701–713. doi:https://doi.org/10.1016/s0896-6273(03)00331-3
- Yang S, Liu A, Weidenhammer A, Cooksey RC, McClain D, Kim MK, Aguilera G, Abel ED, Chung JH. 2009. The role of mPer2 clock gene in glucocorticoid and feeding rhythms. Endocrinology. 150(5):2153–2160. doi:https://doi.org/10.1210/en.2008-0705
- Yang X, Downes M, Yu RT, Bookout AL, He W, Straume M, Mangelsdorf DJ, Evans RM. 2006. Nuclear receptor expression links the circadian clock to metabolism. Cell. 126(4):801–810. doi:https://doi.org/10.1016/j.cell.2006.06.050
- Zhang EE, Liu Y, Dentin R, Pongsawakul PY, Liu AC, Hirota T, Nusinow DA, Sun X, Landais S, Kodama Y, et al. 2010. Cryptochrome mediates circadian regulation of cAMP signaling and hepatic gluconeogenesis. Nat Med. 16(10):1152–1156. doi:https://doi.org/10.1038/nm.2214
- Zhang Y, Fang B, Damle M, Guan D, Li Z, Kim YH, Gannon M, Lazar MA. 2016. HNF6 and Rev-erbα integrate hepatic lipid metabolism by overlapping and distinct transcriptional mechanisms. Genes Dev. 30(14):1636–1644. doi:https://doi.org/10.1101/gad.281972.116
- Zheng X, Sehgal A. 2010. AKT and TOR signaling set the pace of the circadian pacemaker. Curr Biol. 20:1203–1208. doi:https://doi.org/10.1016/j.cub.2010.05.027
- Zhou B, Zhang Y, Zhang F, Xia Y, Liu J, Huang R, Yuangao W, Hu Y, Wu J, Dai C, et al. 2014. CLOCK/BMAL1 regulates circadian change of mouse hepatic insulin sensitivity by SIRT1. Hepatology. 59(6):2196–2206. doi:https://doi.org/10.1002/hep.26992
- Zigman JM, Elmquist JK. 2003. Minireview: from anorexia to obesity–the yin and yang of body weight control. Endocrinology. 144(9):3749–3756. doi:https://doi.org/10.1210/en.2003-0241
- Zvonic S, Ptitsyn AA, Conrad SA, Scott LK, Floyd ZE, Kilroy G, Wu X, Goh BC, Mynatt RL, Gimble JM. 2006. Characterization of peripheral circadian clocks in adipose tissues. Diabetes. 55(4):962–970. doi:https://doi.org/10.2337/diabetes.55.04.06.db05-0873
