Abstract
A conventional fermenter (CF), a single-cathode fermenter (SCF), and a double-cathode fermenter (DCF) were employed to evaluate and compare the effects of H2 and electrochemical reducing power on metabolite production by Clostridium acetobutylicum KCTC1037. The source of the external reducing power for CF was H2, for the SCF was electrochemically reduced neutral red-modified graphite felt electrode (NR-GF), and for the DCF was electrochemically reduced combination of NR-GF and platinum plate electrodes (NR-GF/PtP). The metabolites produced from glucose or CO2 by strain KCTC1037 cultivated in the DCF were butyrate, ethanol, and butanol, but ethanol and butanol were not produced from glucose or CO2 by strain KCTC1037 cultivated in the CF and SCF. It is possible that electrochemically reduced NR-GF/PtP is a more effective source of internal and external reducing power than H2 or NR-GF for strain KCTC1037 to produce metabolites from glucose and CO2. This research might prove useful in developing fermentation technology to actualize direct bioalcohol production of fermentation bacteria from CO2.
Electrochemical bioreactors (fermenters) composed of a neutral red-modified graphite felt electrode (NR-GF), a platinum counter electrode, a cathode compartment, an anode compartment, and a compartmentation membrane are applied to improve metabolite production by the use of various fermentation bacteria.Citation1–3) NR-GF is an effective solid electron mediator transferring electrons from the electrode to bacterial cells and catalyzing NADH regeneration.Citation4–7) The electrochemical reducing power generated by NR-GF can be not only an external reducing power inducing conservation of environmental redox potential, but also an internal reducing power inducing regeneration of NADH for fermentation bacteria.Citation8) On the other hand, H2 can be limited to external reducing power for the conservation of environmental redox potential, and glucose can be limited to internal reducing power for NADH regeneration, because H2 is absorbed in limited amounts into the bacterial cytoplasm and glucose is metabolically oxidized only in the cytoplasm. Accordingly, electrochemical reducing power can substitute for glucose and H2 in the cultivation of autotrophic fermentation bacteria.
In natural ecosystems, CO2 is converted metabolically to methane, acetic acid, butyric acid, ethanol, and butanol with H2 as electron donor by anaerobic respiration or fermentation metabolism.Citation9–13) Clostridium acetobutylicum is a heterotroph that produces free energy and metabolites by fermentative oxidation of sugar, and it is also an autotroph that produces free energy and metabolites through the coupling redox reaction of H2 and CO2 under completely reduced or strict anaerobic conditions.Citation14) When strictly anaerobic bacteria are exposed to incompletely reduced or oxidized conditions, the biochemical reducing power (NADH) and free energy (ATP) that are regenerated by metabolic oxidation of sugar must be wasted in order to recover environmental redox potential.Citation15,16)
Mixotrophic bacteria inhabiting a natural ecosystem can effectively conserve environmental redox potential by H2 and can regenerate NADH and ATP by additional reducing power generated from oxidation metabolism of sugar.Citation17–19) Strictly autotrophic bacteria that depend totally upon H2 are at a disadvantage as compared to mixotrophic bacteria due to the extremely low solubility of H2 in water. This might be a reason for the environmental redox potential not to be effectively conserved and for internal reducing power to regenerate only to a limited extent. This is a weak point of C. acetobutylicum that grows autotrophically with H2 and CO2. In order to remedy this weak point, H2 must be replaced by the electrochemical reducing power generated by a specific electrode or else high pressure H2 (maximal 1479 kPa or 14.6 atm) must be used to fill in the head space of a bacterial culture.Citation20–22)
The goal of this study was to evaluate the effects of H2, electrochemically reduced NR-GF, and electrochemically reduced NR-GF and platinum plate electrodes (NR-GF/PtP) comparatively on metabolite production by C. acetobutylicum KCTC1037. Electrochemically reduced methyl viologen has been reported to cause the redox potential of bacterial culture fall below −350 mV and to cause C. acetobutylicum to produce 26% more butanol but 25% less acetone than conventional culture conditions.Citation23,24) This is a clue that the fermentation metabolism of C. acetobutylicum is controlled by electrochemical reducing power.
In this study, NR-GF was employed in a fermenter as source of electrochemical reducing power, and PtP was placed on the entire bottom of the fermenter as source of auxiliary external reducing power. A conventional fermenter (CF), an single-cathode fermenter (SCF), and a double-cathode fermenter (DCF) were specially designed to cultivate C. acetobutylicum with various sources of external reducing power, H2, NR-GF, and NR-GF/PtP, respectively, in same medium and under the same culture conditions.
Materials and methods
Bacterial culture
C. acetobutylicum KCTC1037 was cultivated in reinforced clostridial medium composed of 10 g/L of beef extract, 10 g/L of peptone, 5 g/L of sodium chloride, 10 g/L of glucose, 3 g/L of yeast extract, 3 g/L of sodium acetate, 1 g/L of soluble starch, and 0.5 g/L of l-cysteine HCl. This medium was used in all the culture tests, but glucose was selectively excluded when CO2 was used as carbon source. C. acetobutylicum was cultivated in anaerobic vials (total volume, 165 mL; working volume, 50 mL) at 37 °C for successive transfer culture and seed culture. Ten percent (v/v) of seed culture grown in the vials was injected by syringe for inoculation. The headspace of the anaerobic vial was filled with 1.2 atm H2 to conserve the reduced condition. Both H2 and CO2 (99.999% each) were continuously sparged into the culture medium in the CF, but CO2 only was sparged into that of the SCF and DCF by membrane filter (pore 0.22 μm, diameter 90 mm; Merck Millipore, Germany) at a rate of 31–32 mL/min, which corresponded to approximate 44.8 L/d.
Conventional fermenter
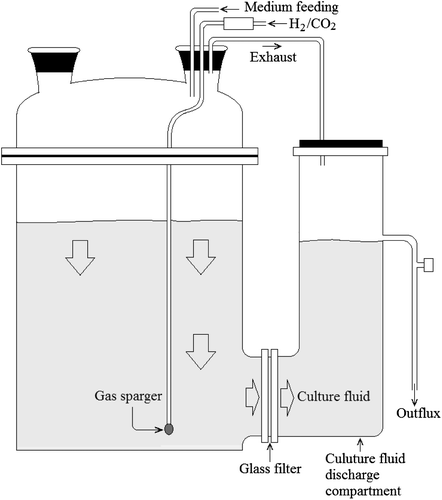
The bacterial culture in the CF was chemically reduced using sparged H2, as shown in Fig. . A glass filter (diameter, 90 mm; pore, 1–1.6 μm; Duran Group, Mainz) was placed between the fermenter and the culture fluid discharge compartment (300 mL) to allow transfer of culture fluid but not bacterial cells. The exhaust was discharged through an outflux to protect the bacterial culture against oxygen and microorganism contamination. The total volume of the fermenter was 2500 mL, and the working volume was 2000 mL.
Single-cathode fermenter
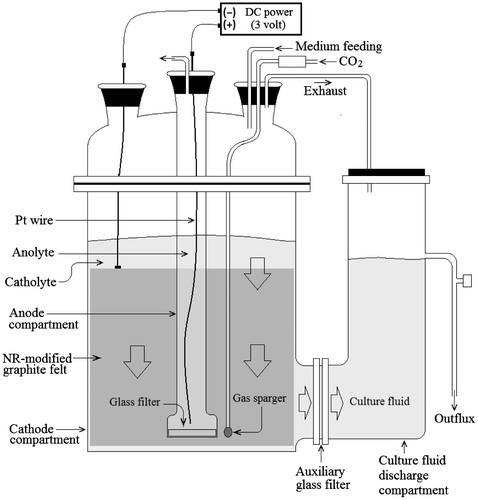
A SCF was prepared by equiping an anode compartment (200 mL) in the CF as shown in Fig. . A glass filter (diameter, 40 mm; pore, 1–1.6 μm; Duran Group), modified with cellulose acetate film (thickness, 35 μm; Electron Microscopy Science, Hatfield, PA), was used to separate the cathode and anode electrically and to limit the transfer of water from the cathode to the anode compartment. An auxiliary glass filter (diameter, 90 mm; pore, 1–1.6 μm; Duran Group), which was not modified with the cellulose acetate film, was employed to prevent the transfer of bacterial cells but to allow that culture fluid. Three-layered rolling-type NR-GF (length, 1000 mm; height, 250 mm; thickness 30 mm; external diameter, 140 mm; internal diameter, 80 mm) was used as cathode, and Pt wire (thickness, 0.5 mm; length, 300 mm) as anode. Electricity (DC 3 V) was charged to the NR-GF cathode and the Pt wire anode to induce the generation of electrochemical reducing power in the bacterial culture.
Double-cathode fermenter
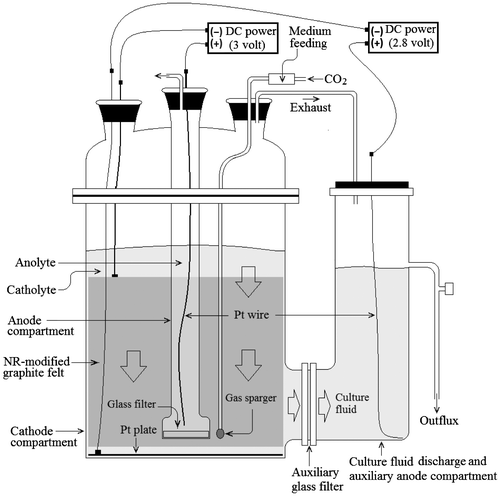
A DCF was prepared by Pt plate (diameter, 140 mm; thickness, 100 μm) on the bottom of the SCF and a Pt wire (thickness, 500 μm; length, 300 mm) in the culture fluid discharge compartment, as shown in Fig. . The Pt plate was an auxiliary cathode and Pt wire equipped in the culture fluid discharge compartment was an auxiliary anode. Electricity (DC 2.8 V) was charged to the auxiliary Pt plate cathode and the auxiliary Pt wire anode to induce the bacterial culture conditions to be reduced. Electricity (DC 3.0 V) was charged to the NR-GF cathode and the Pt wire anode to induce the generation of electrochemical reducing power in the bacterial culture. The DCF, composed of double cathodes, effectively reduced the bacterial culture conditions and effectively activated bacterial metabolism. The PtP cathode was electrically isolated from the NR-GF cathode to block off electrical interference between NR-GF and PtP. The electrical voltage charged to the PtP cathode was slightly lower than that charged to the NR-GF cathode to protect against electrochemical corrosion of the graphite felt.
Analysis
Bacterial metabolites were analyzed by a gas chromatography/mass spectrometry (GC/MS) system (Clarus 600 series + TurboMatrix HSS Trap, PerkinElmer, Waltham, MA) equipped with a capillary column (Elite-FFAP; ID, 0.25 μm; OD, 0.32 μm; length, 30 m; PerkinElmer) and an electron ionization system with He as carrier gas (flow rate, 1.0 mL/min). The injector temperature was adjusted to 250 °C, the initial oven temperature was adjusted to 80 °C, and the oven temperature was increased at a rate of 10 °C min−1. The oven temperature was held for 2 min after it reached 250 °C to clean the column. Initially, the samples were injected in splitless injection mode, and the injector was switched to split mode for 2 min after the sample was injected. The bacterial culture was centrifuged at 10,000 × g at 4 °C for 30 min and filtered with a membrane filter (pore size, 0.22 μm), and then injected directly into the GC/MS. The concentrations of metabolites were determined based on peak areas of standard compounds, and chemical species were determined based on the mass profile database. Acetic acid was not analyzed, since it was included in the culture medium. The biomass of strain KCTC1037 could not be analyzed or compared since it was growing inside the NR-GF cathode.
Continuous culture condition
Fresh reinforced clostridial medium with and without glucose was supplied continuously to the three types of fermenters at a rate of 200 mL/d for 12 weeks. Metabolites produced by C. acetobutylicum KCTC1037 from glucose and from CO2 growing in the fermenters were analyzed daily, and the mean concentrations of daily analyzed metabolites were recorded at one week intervals.
Measurement of biomass
The biomass of KCTC1037 immobilized in NR-GF could not be measured because the NR-GF could not be taken out from the fermenter during operation. Instead, biomass was estimated based on the dry mass of bacterial cells discharged from the outlets of CF, SCF, and DCF. Bacterial culture discharged from the outlet was collected with a cold container (4 °C) during the one week after metabolite production in each fermenter was stabilized. Bacterial cells were harvested from 1000 mL of the collected bacterial culture and washed with distilled water by centrifugation at 5000 × g at 4 °C over 30 min. The washed bacterial cells were lyophilized and precisely weighed to determine dry mass.
Results
Metabolites produced in the CF and SCF
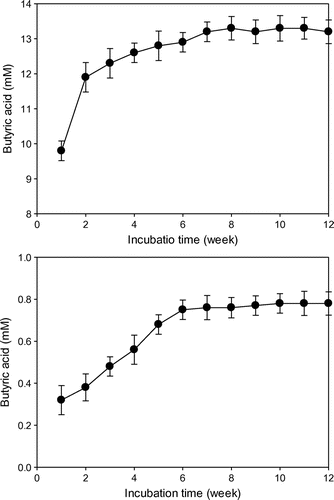
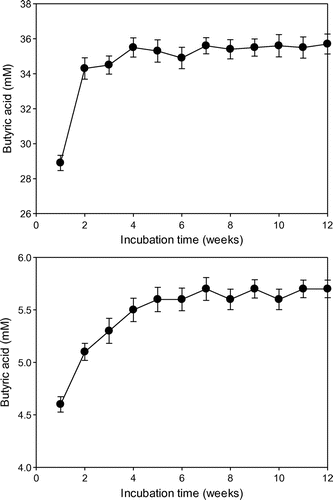
Butyric acid (11–13 mM) was produced from glucose, but only a trace amount of butyric acid was produced from CO2 by C. acetobutylicum KCTC1037 cultivated in CF (Fig. ). In contrast, strain KCTC1037 produced 33–36 and 5–6 mM of butyric acid from glucose and CO2, respectively, in the SCF (Fig. ). The butyric acid produced from glucose by strain KCTC1037 cultivated in SCF was approximately three times as that in the CF. Electrochemically reduced NR-GF-induced strain KCTC1037 to produce butyric acid from CO2, but H2 did not. This result suggests that sparging H2 into the culture medium is less effective as external reducing power to conserve environmental redox potential and to regenerate NADH and ATP than the electrochemically reduced NR-GF.
Metabolites produced in the DCF
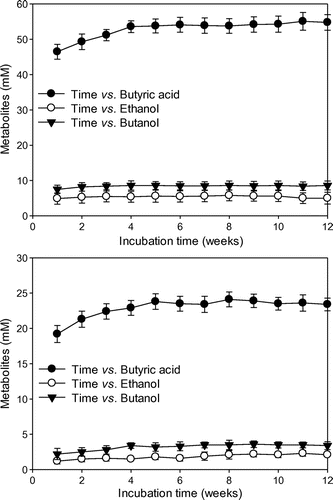
Butyric acid (53–55 and 22–24 mM) was produced from glucose and CO2, 5–6 and 1–2 mM of ethanol was produced from glucose and CO2, and 8–9 and 2–3 mM of butanol was produced from glucose and CO2 in the DCF (Fig. ). Electrochemically reduced NR-GF/PtP-induced strain KCTC1037 to produce ethanol and butanol newly from both glucose and CO2. This is a clue that the electrochemically reduced NR-GF/PtP is a more effective reducing power source for the conservation of environmental redox potential and the regeneration of NADH and ATP than an electrochemically reduced NR-GF.
Biomass of KCTC1037 grown in CF, SCF, and DCF
The KCTC1037 grown in three fermenters with glucose, and in SCF and DCF with CO2 produced butyric acid in proportion to its biomass, but KCTC1037 was not grown and did not produce butyric acid in CF with CO2, as shown in Table . KCTC1037 is assumed not to grow in CF with CO2, considering the inoculation ratio (10%, v/v) of the seed culture (biomass, 0.4 g/L).
Table 1. Dry cell mass of KCTC1037 grown in CF, SCF, and DCF with CO2 and with glucose.
Discussion
In mixed acid fermentation bacteria, the conversion of acyl-CoA (acetyl-CoA and butyryl-CoA) to alkanoic acid and alkanol is a branch point for physiological adjustment of the internal energy state by balancing of the NADH/NAD and ATP/ADP ratios.Citation25) The flux distribution at the branch point can be a metabolic function to conserve the internal energy state in bacterial cells. The internal energy state, as reflected by the NADH/NAD ratio and the ATP pool, is conserved or increased by the environmental redox potential, but the environmental redox potential can be regulated chemically by reduced metabolites or controlled biochemically by NADH.Citation26,27) The correlation between the internal energy state and the environmental redox potential has been positively established, considering that the kind and concentration of metabolites varied with the substrate (glucose or CO2) and the cultivation conditions (CF, SCF, and DCF) (Figs. ).Citation28)
The lower redox potential of H2 (−0.42 V vs. NHE) than of NADH (−0.32 V vs. NHE) is available as an external reducing factor for conserving the internal energy state in C. acetobutylicum and maintaining the environmental redox potential, but the extremely low solubility of H2 in water (0.0014 g/L or 0.7 mM at 37 °C under atmospheric pressure) might limit as an external reducing power.Citation29) Strain KCTC1037 can absorb the H2 sparged into medium to a limited extent and depend completely upon the NADH and ATP generated from glucose to conserve the internal energy state and the environmental redox potential.Citation30) Some of the NADH and ATP generated from glucose in the metabolism of strain KCTC1037 may have been wasted to conserve the cytoplasmic and environmental redox potential under conditions without the available external reducing power. This may have caused the production of ethanol and butanol coupled to NADH oxidation to be limited, as shown in Fig. (CF pathway). The NADH generated from H2 that was absorbed into cytoplasm may not have been enough for the operation of Wood–Ljungdahl pathway. Accordingly, strain KCTC1037 cultivated in the CF with H2 and CO2 neither grew nor produced metabolites (Table ).
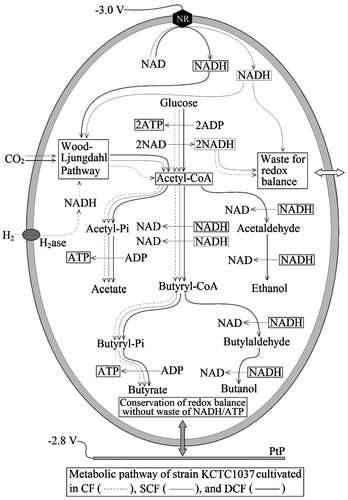
Fig. 7. Proposed metabolic pathway of C. acetobutylicum KCTC1037 cultivated in CF, SCF, and DCF in coupled to redox reaction of NADH/NAD and the regeneration of ATP.
Note: H2ase (hydrogenase) catalyzes the oxidation of H2 coupled to NADH regeneration, but the NADH regenerated by oxidation of H2 might not be sufficient for the conservation of internal reducing power and environmental redox potential due to extremely low solubility. ATP is regenerated, but NADH is not consumed in the pathway from acyl-CoA to acetic acid, whereas ATP is not regenerated but NADH is consumed in the pathway from acyl-CoA to ethanol. Alcohol production might be limited under NADH-deficient conditions (CF, SCF) but possible under NADH-sufficient conditions (DCF). Butyryl-CoA is generated from two acetyl-CoA coupled to consumption of two NADH, by which butyric acid production is increased proportionally to balance NADH/NAD, which is higher in the SCF than in the CF but lower than in DCF. Alcohol production can be proportional to the conservation efficiency of ATP and butyric acid production can be proportional to NADH/NAD ratio.
A suitable environmental redox potential is a signal for the expression of a specific gene that controls metabolite production in fermentation bacteria.Citation31) For example, the ethanol and butanol dehydrogenase-coding genes are adhE and adhE2, respectively, and these are known to be expressed at high NADH/NAD ratios. NADH regeneration is known to be influenced by the environmental redox potential.Citation32–35) Electrochemically reduced NR-GF is a more effective external reducing factor than sparged H2 for conserving a high NADH/NAD ratio and the environmental redox potential. Under these conditions, acetyl-CoA production from CO2 can be induced and acetyl-CoA can be converted to butyryl-CoA coupled to NADH oxidation, as shown in Fig. (SCF pathway). The NADH regenerated by NR-GF can be wasted as to the conservation of cytoplasmic and environmental redox potential but can be consumed in the Wood–Ljungdahl pathway and in the reduction pathway of acyl-CoA (butyryl-CoA) to ethanol (butanol). Alkanoic acid production from acetyl-CoA might be more advantageous thermodynamically than alkanol production from acetyl-CoA, because ATP is regenerated but NADH is not consumed in the pathway from acetyl-CoA to alkanoic acid.Citation36,37) Accordingly, butyric acid was produced from both glucose and CO2 in the SCF and butyric acid production from glucose in the SCF was higher than in the CF. Butyric acid production from glucose in the CF and SCF was proportional to the biomass of KCTC1037 (Table ). This is a clue that alkanoic acid production is coupled to ATP regeneration.
The metabolic branch point where acyl-CoA is converted to alkanoic acid and alkanol can function to control the internal energy state (the ratio of NADH/NAD and ATP/ADP). The conversion of acyl-CoA to alkanoic acids and alkanols must be metabolically controlled for conservation of the NADH/NAD ratio and the ATP pool. Theoretically, KCTC1037 can produce alkanoic acid and alkanol from glucose or CO2 when NADH/NAD ratio is higher than ATP/ADP ratio and environmental redox potential is suitable.Citation38–40) NADH regenerated by NR-GF might be consumed in the pathway from acyl-CoA to aldehyde and aldehyde to alkanol and electrochemical reducing potential generated by PtP might be consumed for conservation of cytoplasmic and environmental redox potential in DCF, as shown in Fig. (DCF pathway). NADH regenerated by electrochemical reducing power can be consumed for the generation of acetyl-CoA in the Wood–Ljungdahl pathway, and the acetyl-CoA can be converted to alkanol to balance NADH/NAD ratio and can be converted to alkanoic acid for ATP regeneration. Alkanol production from glucose is higher than that from CO2 due to the additional NADH generated by glycolysis. This is why alkanol produced from both glucose and CO2 were lower than alkanoic acid in the DCF and why alkanoic acid production from both glucose and CO2 in the DCF was significantly high than in the SCF.
In conclusion, it is possible that the NR-GF/PtP can more actively catalyze the regeneration of NADH and ATP than sparged H2 or NR-GF. The higher NADH/NAD ratio and higher ATP pool can induce a highly conserved internal energy state and a suitably conserved environmental redox potential, which must be a factor for strain KCTC1037 in producing butyric acid, ethanol, and butanol from CO2, and in increasing production of butyric acid, ethanol, and butanol from glucose.
Notes
Abbreviations: CF, conventional fermenter; DCF, double-cathode fermenter; NR, neutral red; NR-GF, neutral red-modified graphite felt electrode; NR-GF/PtP, NR-GF and PtP electrodes in combination; PtP, platinum plate electrode; SCF, single-cathode fermenter.
References
- Park DH, Zeikus JG. J. Bacteriol. 1999;181:2403–2410.
- Park SM, Sang BI, Park DW, Park DH. J. Microbiol. 2005;43:451–455.
- Jeon BY, Yi JY, Jung IL, Park DH. Adv. Microbiol. 2013;3:42–45.10.4236/aim.2013.31006
- Lee WJ, Park DH. J. Microbiol. Biotechnol. 2009;19:836–844.
- Jeon BY, Hwang TS, Park DH. J. Microbiol. Biotechnol. 2009;19:666–674.
- Jeon BY, Seo HN, Kang SW, Park DH. J. Microbiol. Biotechnol. 2010;20:485–493.
- Girbal L, Vasconcelos I, Saint-Amans S, Soucaille P. FEMS Microbiol. Rev. 1995;161:151–162.
- Snoep JL, de Graef MR, Teixeira de Mattos MJ, Neijssel OM. FEMS Microbiol. Lett. 1994;116:263–267.10.1111/fml.1994.116.issue-3
- Cheng L, Dai L, Li X, Zhang H, Lu Y. Appl. Environ. Microbiol. 2011;77:5212–5219.10.1128/AEM.00210-11
- Buschhorn H, Dürre P, Gottschalk G. Appl. Environ. Microbiol. 1989;55:1835–1840.
- Lajoie SF, Bank S, Miller TL, Wolin MJ. Appl. Environ. Microbiol. 1988;54:2723–2727.
- Ohwaki K, Hungate RE. Appl. Environ. Microbiol. 1977;33:1270–1274.
- Tabita FR. Microbiol. Mol. Biol. Rev. 1988;52:155–189.
- Daniel SL, Hsu T, Dean SI, Drake HL. J. Bacteriol. 1990;172:4464–4471.
- Riebe O, Fischer RJ, Wampler DA, Kurtz DM Jr, Bahl H. Microbiology. 2009;155:16–24.10.1099/mic.0.022756-0
- Hillmann F, Fisher RJ, Saint-Prix F, Girbal L, Bahl H. Microbiology. 2008;68:848–860.
- Dwyer DF, Weeg-Aerssens E, Shelton DR, Tiedje JM. Appl. Environ. Microbiol. 1988;54:1354–1359.
- Tomei FA, Maki JS, Mitchell R. Appl. Environ. Microbiol. 1985;50:1244–1250.
- Lee MJ, Zinder SH. Appl. Environ. Microbiol. 1988;54:124–129.
- Jeon BY, Jung IL, Park DH. J. Microbiol. Biotechnol. 2011;21:590–598.
- Hwang TS, Na BK, Tran HT, Ahn DH, Park DH. Bioprocess Eng. 2008;13:677–682.10.1007/s12257-008-0054-z
- Yerushalmi L, Volesky B, Szczesny T. Appl. Microbiol. Biotechnol. 1985;22:103–107.
- Kim TS, Kim BH. Biotechnol. Lett. 1988;10:123–128.10.1007/BF01024638
- Peguin S, Soucaille P. Biotechnol. Bioeng. 1996;51:342–348.10.1002/(ISSN)1097-0290
- Guedon E, Payot S, Desvaux M, Petitdemange H. J. Bacteriol. 1999;181:3262–3269.
- Girbal L, Soucaille P. J. Bacteriol. 1994;176:6433–6438.
- Meyer CL, Papoutsakis ET. Appl. Microbiol. Biotechnol. 1989;30:450–459.
- de Graef MR, Alexeeva S, Snoep JL, Teixeira de Mattos MJ. J. Bacteriol. 1999;181:2351–2357.
- McKinlay JB, Zeikus JG. Appl. Environ. Microbiol. 2004;70:3467–3474.10.1128/AEM.70.6.3467-3474.2004
- Riondet C, Cachon R, Wache Y, Alcaraz G, Divies C. J. Bacteriol. 2000;182:620–626.10.1128/JB.182.3.620-626.2000
- Iuchi S, Lin EC. Mol. Microbiol. 1993;9:9–15.10.1111/mmi.1993.9.issue-1
- Leonardo MR, Cunningham PR, Clark DP. J. Bacteriol. 1993;175:870–878.
- Leonardo MR, Dailly Y, Clark DP. J. Bacteriol. 1996;178:6013–6018.
- Youngleson JS, Jones WA, Jones DT, Woods DR. Gene. 1989;78:355–364.10.1016/0378-1119(89)90238-2
- Fontaine L, Meynial-Salles I, Girbal L, Yang X, Croux C, Soucaille P. J. Bacteriol. 2002;184:821–830.10.1128/JB.184.3.821-830.2002
- Grupe H, Gottschalk G. Appl. Environ. Microbiol. 1992;58:3896–3902.
- Zhao Y, Tomas CA, Rudolph FB, Papoutsakis ET, Bennett GN. Appl. Environ. Microbiol. 2005;71:530–537.10.1128/AEM.71.1.530-537.2005
- Park DH, Zeikus JG. J. Bacteriol. 1999;181:2403–2410.
- Jeon BY, Hwang TS, Park DH. J. Microbiol. Biotechnol. 2009;19:666–674.
- Park SM, Sang BI, Park DW, Park DH. J. Microbiol. 2005;43:451–455.