p63 is known as a regulator of cell cycle and apoptosis in response to genotoxic insults and for its function in regulation of epithelial morphogenesis. Melino and colleagues now demonstrate that phosphorylation of a conserved threionine (T538) regulates p63 stability and degradation mediated by WW domain containing proteins, Itch and Pin1.
WW domains are protein modules that mediate protein-protein interactions through recognition of proline-rich peptide motifs and phosphorylated serine/threonine-proline (S/T)P sites.Citation1 p63, a member of the p53 family of transcription factors and known to play a crucial role in regulating transcription of genes with distinct biological functions in regulating epithelium development and differentiation, metastasis and chemosensitivity,Citation2,3 contains a (T/S)PPPxY domain which equates to a class IV WW target domain, (T/S)P, followed by a class I WW domain target, PPxY. Indeed p63 is a known target of Class I WW domains interacting with Itch, an ubiquitin E3 ligase, which via its WW2 domain mediates the proteasomal degradation of p63.Citation4,5 More recently, it has been shown that Pin1, a prolyl-peptidyl cis/trans isomerase protein which targets Class IV WW domains, modulates p63 protein stability by inhibiting p63 proteasomal degradation.Citation6 In their latest study, Melino and colleaguesCitation7 elucidate aspects of p63 stability mediated by the WW domain-containing proteins Itch and Pin1, proposing a twofold regulation mechanism: whereas phosphorylation of threonine (T538) within the (T/S)PPPxY motif of p63 increases its binding affinity to the Itch WW domain, a post phosphorylation cis/trans switch catalyzed by Pin1 prolyl-isomerase regulates the final outcome of this interaction.Citation7 This further extends on previous observations that E3 ligases prefer the trans conformer (reviewed in Citation8) showing that the more relaxed trans conformer is indeed favorable for Itch binding. This is likely a consequence of the favorable local structural and chemical complementarity with the WW binding trench (). To maintain steady states of p63 levels in the cell, Melino and colleaguesCitation7 proposed that the cis/trans equilibrium and consequent stability of the Itch-p63 interaction might be regulated by Pin1, which catalyzes the formation of the Itch-unfavorable cis conformation.
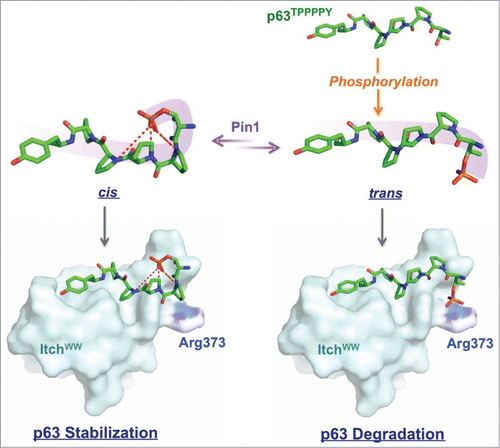
Figure 1. The local structural configuration of the cis isomer, with the phosphate group twisted away from favorable interactions with the WW domain to favor self-interactions with the peptide's own amides, restrains Itch binding leading to p63 stabilization against subsequent ubiquitylation. The trans conformation exposes the negatively charged phosphate group for stabilizing interactions with a conservedCitation7 basic arginine complement within the WW domain.
The twofold post phosphorylation conformational switch mechanism observed by Melino et al.Citation7 is unlikely limited to the Itch-p63 interaction and could also be true for other WW domain interacting partners. Indeed, it has recently been shown that Pin1 directly binds to and stabilizes p63 inhibiting proteasomal degradation mediated by the E3 ligase WWP1.Citation6 Pin1 specifically interacts with T538P and disrupts p63α-WWP1 interaction.Citation6 Likewise, Pin1 binding to phosphorylated p53 or p73 family members has been shown to stabilize both proteins by inhibiting p53 binding to E3 ligase MDM2 or increasing p73 acetylation by p300 (reviewed in Citation8). This study also highlights the essentiality of tightly regulating p(S/T)P motifs involved in pivoting the disastrous switch between normal and pathologic cellular signaling. Considering that the human proteome is highly enriched with (S/T)P motifs,Citation9 it is fair to presume that Pin1, implicated in regulation of cell proliferation, transformation, and tumor growth, interacts with many other phospho-(S/T)P motif containing proteins in order to modulate their function.
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
Acknowledgments
The authors are grateful to Dr. Ailie Marx for critical reading of the manuscript.
Funding
The Aqeilan lab is supported, in part, by funds from the Israel Science Foundation (ISF 12-542).
References
- Salah Z, et al. Front Biosci 2012; 17:331-48; PMID:22201747; http://dx.doi.org/10.2741/3930
- Murray-Zmijewski F, et al. Cell Death Differ 2006; 13:962-72; PMID:16601753; http://dx.doi.org/10.1038/sj.cdd.4401914
- Su X, et al. Nat Rev Cancer 2013; 13:136-43; PMID:23344544; http://dx.doi.org/10.1038/nrc3446
- Rossi M, et al. Proc Natl Acad Sci U S A 2006; 103:12753-8; PMID:16908849; http://dx.doi.org/10.1073/pnas.0603449103
- Rossi M, et al. Cell Cycle 2006; 5:1816-22; PMID:16861923; http://dx.doi.org/10.4161/cc.5.16.2861
- Li C, et al. Cell death & disease 2013; 4:e943; PMID:24309930; http://dx.doi.org/10.1038/cddis.2013.468
- Melino S, et al. Cell Cycle 2014; 13:3207-17; PMID:25485500; http://dx.doi.org/10.4161/15384101.2014.951285
- Lu KP, Zhou XZ. Nat Rev Mol Cell Biol 2007; 8:904-16; PMID:17878917; http://dx.doi.org/10.1038/nrm2261
- Abu-Odeh M, et al. J Biol Chem 2014; 289:8865-80; PMID:24550385; http://dx.doi.org/10.1074/jbc.M113.506790
