ABSTRACT
Centrosomes are organelles that function as hubs of microtubule nucleation and organization, with key roles in organelle positioning, asymmetric cell division, ciliogenesis, and signaling. Aberrant centrosome number, structure or function is linked to neurodegenerative diseases, developmental abnormalities, ciliopathies, and tumor development. A major regulator of centrosome biogenesis and function in C. elegans is the conserved Spindle-defective protein 2 (SPD-2), a homolog of the human CEP-192 protein. CeSPD-2 is required for centrosome maturation, centriole duplication, spindle assembly and possibly cell polarity establishment. Despite its importance, the specific molecular mechanism of CeSPD-2 regulation and function is poorly understood. Here, we combined computational analysis with cell biology approaches to uncover possible structure–function relationships of CeSPD-2 that may shed mechanistic light on its function. Domain prediction analysis corroborated and refined previously identified coiled-coils and ASH (Aspm-SPD-2 Hydin) domains and identified new domains: a GEF domain, an Ig-like domain, and a PDZ-like domain. In addition to these predicted structural features, CeSPD-2 is also predicted to be intrinsically disordered. Surface electrostatic maps identified a large basic region unique to the ASH domain of CeSPD-2. This basic region overlaps with most of the residues predicted to be involved in protein–protein interactions. In vivo, ASH::GFP localized to centrosomes and centrosome-associated microtubules. Our analysis groups ASH domains, PapD, Usher chaperone domains, and Major Sperm Protein (MSP) domains into a single superfold within the larger Immunoglobulin superfamily. This study lays the groundwork for designing rational hypothesis-based experiments to uncover the mechanisms of CeSPD-2 function in vivo.
Abbreviations: AIR, Aurora kinase; ASH, Aspm-SPD-2 Hydin; ASP, Abnormal Spindle Protein; ASPM, Abnormal Spindle-like Microcephaly-associated Protein; CC, coiled-coil; CDK, Cyclin-dependent Kinase; Ce, Caenorhabditis elegans; CEP, Centrosomal Protein; CPAP, centrosomal P4.1-associated protein; D, Drosophila; GAP, GTPase activating protein; GEF, GTPase guanine nucleotide exchange factor; Hs, Homo sapiens/Human; Ig, Immunoglobulin; MAP, Microtubule associated Protein; MSP, Major Sperm Protein; MDP, Major Sperm Domain-Containing Protein; OCRL-1, Golgi endocytic trafficking protein Inositol polyphosphate 5-phosphatase; PAR, abnormal embryonic PARtitioning of the cytosol; PCM, Pericentriolar material; PCMD, pericentriolar matrix deficient; PDZ, PSD95/Dlg-1/zo-1; PLK, Polo like kinase; RMSD, Root Mean Square Deviation; SAS, Spindle assembly abnormal proteins; SPD, Spindle-defective protein; TRAPP, TRAnsport Protein Particle; Xe, Xenopus; ZYG, zygote defective protein.
Introduction
Centrosomes are organelles with key roles in the positioning of other organelles in the cell and function as hubs for microtubule nucleation and organization. They are composed of a pair of microtubular centrioles immersed in a protein matrix known as the pericentriolar material (PCM). The centrioles and the PCM are codependent for many of the centrosome functions but have distinct roles and largely associated with different sets of proteins [Citation1,Citation2]. An exception is the C. elegans spindle-defective protein 2 (CeSPD-2) protein, a homolog of the human CEP-192 protein. CeSPD-2 localization to the centrosome or exclusively to the centrioles is likely relevant to its dual role on centriole duplication and centrosome maturation [Citation3,Citation4].
The first mitosis of the C. elegans embryo has been a central model for understanding centrosome biogenesis, function, and structure [Citation5]. At fertilization, the sperm provides a pair of engaged centrioles that separate, duplicate, and recruit PCM proteins present in the oocyte’s cytosol. SPD-2 is required throughout the assembly of the maternally contributed PCM [Citation1]. In the zygote, SPD-2 localizes to the sperm centriole pair and around the sperm DNA [Citation6]. The earliest detectable PCM proteins around the sperm-provided centrioles are the coiled-coil proteins PCM deficient-1 (PCMD-1), SPD-5, SPD-2, and the microtubule anchoring protein γ -tubulin. SPD-2 and SPD-5 are mutually dependent for their localization to the PCM, and PCMD-1 is required for the recruitment of SPD-5 [Citation3],Citation7]. SPD-2 and γ-tubulin accumulation on the centrosome is partially dependent on the Aurora A Kinase (AIR-1 in C. elegans) and cytoplasmic dynein dhc-1 [Citation3]. As the embryo enters mitosis SPD-2 and SPD-5 promote PCM assembly and centrosome maturation [Citation3]. SPD-2 recruits polo-like kinase-1 (PLK-1) and Aurora kinase A (AIR-1), which are required for maturation [Citation8,Citation9]. At metaphase PCM components are restructured. PLK-1 phosphorylates SPD-5 [Citation10] and this initiates polymerization of PCM and the formation of a γ-tubulin ring surrounded by a microtubule ring [Citation11,Citation12].
As the PCM volume and γ-tubulin increase [Citation13,Citation14], the sperm-provided centrioles separate and start to assemble daughter centrioles by sequentially recruiting maternally provided structural and regulatory centriolar components [Citation15–17]. This process is dependent on PCM proteins [Citation2]. Six proteins are required for centriole assembly; the likely ortholog of Polo like kinase 4 (PLK4) in mammals Zygote defective 1 protein (ZYG-1) [Citation18], and coiled-coil proteins: SPD-2 [Citation3,Citation4,Citation15] and the other Spindle assembly abnormal proteins SAS-4 [Citation19], SAS-5 [Citation20], SAS-6 and SAS-7 [Citation21]. SAS-7 directly binds to and recruits SPD-2 to the emerging daughter centrioles [Citation21]. Although CeSPD-2 and its human ortholog CEP192 are essential for centriole duplication [Citation4,Citation22,Citation23], it is not clear whether this is true for the Drosophila orthologue, DsSPD-2 [Citation24,Citation25,].
As the sperm provided centrosome duplicates, the anteroposterior polarity of the zygote is established. This asymmetry breaking depends on the male pronucleus associated centrosome and possibly the centrosome associated microtubules [Citation26–30]. Therefore, polarity in the C. elegans embryo indirectly depends on the centrosome maturation roles of CeSPD-2 and SPD-5. SPD-2 and SPD-5 have also been proposed to have direct roles in breaking symmetry in the embryo [Citation31–33], but this possibility remains contentious.
CeSPD-2 is an excellent entry point for uncovering mechanisms underlying centrosome function and inheritance, because of its multiple roles and intermolecular interactions, and because its poorly understood regulation and mechanisms of function. We took a mostly in silico approach to predict structural features and domains in CeSPD-2 to base future inquiries about its regulation and function. We report that despite being predicted to be a largely intrinsically disordered protein, CeSPD-2 has several functional domains including previously unreported GEF, Ig-like, and PDZ-like domains, as well as a fourth coiled coil. In addition, this thorough analysis provided a refined prediction of previously reported coiled coils and the Aspm-SPD2-Hydin (ASH) domain of CeSPD-2. The modeled three-dimensional structure of the ASH domain reveals that it is a member of a structural superfold that includes Major Sperm Protein (MSP), PapD, and usher-chaperone domains. The CeSPD-2 ASH domain localizes to the centrosome in-vivo as previously shown in-vitro for other ASH/PapD domain containing proteins [Citation34–37]. Based on previous reports and our findings, we speculate on how the domains and structural features of CeSPD-2 agree with its roles, and hint at possible mechanisms of function and its regulation.
Results
CeSPD-2 is a multidomain protein with an intrinsically disordered N-terminus
The predicted secondary structure of C. elegans SPD-2 protein sequence (NP_492414.1) (Figure S1) suggests that it is 50% unstructured. The N-terminal half of CeSPD-2 including the first 460 residues contains only five to seven α-helices ( and S1) and is predicted to be intrinsically disordered (72%) (). CeSPD-2 was previously reported to have three coiled-coils (CC), similarly located to our prediction of CC1 (65–82), CC2 (111–138), and CC3 (291–322) ( and S1) [Citation3]. Our analysis also identified an additional fourth coiled-coil region, CC4 (368–400). CC3 and CC4 lie within a previously unpredicted RhoGEF-like domain motif (258–467) (). This domain is most similar to the RhoGEF domain in the hypothetical Rom1 protein from Encephalitozoon cuniculi (14–218) that has sequence similarity to the DH domains of RhoGEFs Rom1 and Rgf1 in S. cerevisiae and S. pombe, respectively. In contrast to the largely unstructured N-terminus, the C-terminal half of CeSPD-2 is predicted to contain several secondary structure signatures indicating hallmarks of protein domains. It is predicted to contain a previously reported [Citation38] ASH domain (475–564), followed by previously unreported Immunoglobulin (Ig)-like (593–660) and PDZ-like (695–820) domains (). ASH domains are remote homologs of the immunoglobulin (Ig)-like seven-stranded beta sandwich fold superfamily [Citation38]. The Ig-like domain is made up of six antiparallel β-sheets in a sandwich conformation reminiscent of an immunoglobulin fold () [Citation39]. The PDZ-like domain contains the carboxylate-binding loop that includes the highly conserved R/K-X-X-X-Φ-G-Φ motif (R743-F749) [Citation40,Citation41].
Figure 1. Predicted domain architecture of CeSPD-2.
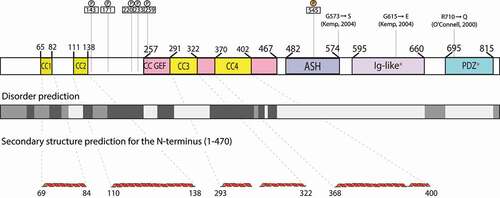
Figure 2. Comparison of sequences of ASH domains from centrosomal proteins.

ASH domains show low primary sequence conservation but have highly conserved structure
A PSI BLAST search (E-value threshold = 0.005) using the CeSPD-2 ASH domain identified only three significant sequence matches: the predicted Caenorhabditis species homologs of SPD-2 from C. brigssae, C. remanei, and a hypothetical protein from the hookworm parasite Necator americanus (Table S1). The third iteration identified seven additional distantly related protein sequences at or above 0.05 E-value threshold. These include Lingula anatine orthologs of SPD-2, a cilia and flagella-associated protein 74 from Spartus aurata, and 5 other predicted proteins of unknown function or proteins with only preliminary sequences from the first round of shotgun sequencing (Table S1).
We wanted to know how conserved the secondary and tertiary structure of ASH domains was, despite their very low primary sequence similarity. To find this, we utilized proteins with known structure and function associated with the centrosome, the cilia, and/or the flagellum, because the PSI-BLAST search yielded only putative or uncharacterized SPD-2 orthologs, predicted proteins, or proteins that are uncharacterized and/or have unknown function and structure. Specifically, we compared the CeSPD-2 ASH domain with its orthologs Centrosomal Protein 192 (Cep192) from Xenopus laevis [Citation42] and Homo sapiens [Citation43,Citation44], the Drosophila melanogaster Abnormal Spindle Protein (ASP) [Citation45], human Abnormal Spindle-like Microcephaly-associated Protein (ASPM) [Citation38,Citation46], the ciliary protein Hydin (Homo sapiens and Mus musculus) [Citation47–49], and the Homo sapiens Golgi endocytic trafficking protein Inositol polyphosphate 5-phosphatase (OCRL-1) [Citation50–52]. Sequence alignment of these ASH domains shows that, as expected, they share little sequence similarity (). An asparagine (N503 in SPD-2) that had been previously defined as a signature feature of ASH domains, is the most highly conserved residue () [Citation38]. Other highly conserved residues include an N-terminal phenylalanine (F486 in CeSPD-2 but absent in XeCEP192), and a proline (P534 in SPD-2, absent in ASP and HsCEP192). In pairwise comparison, CeSPD-2 ASH has higher sequence similarity (33.33%) to the ASH domain of the non-orthologous ASPM protein from humans () than to its orthologs, CEP192 from Xenopus (32.39%) and humans (26.88%). Interestingly, the Xenopus CEP192 ASH domain shares more similarity with the CeSPD-2 ASH domain than it does with the human CEP192 ASH domain (30.98%). Despite having little primary sequence similarity, the secondary structure of these centrosome proteins ASH domains appear to be well conserved. For instance, the 7 β-strands predicted in the CeSPD-2 ASH domain are located at similar positions to β-strands previously determined by X-ray diffraction in the ASH domain from OCRL-1 (3QIS[Citation52]) (). To assess conservation of the tertiary structure of the centrosome protein’s ASH domain, we first generated a 3D model for the ASH domain in CeSPD-2, ASPM, and ASP (). The top ranked 3D model for the CeSPD-2 ASH domain was produced by LOMETS [Citation53] using the C-terminal PapD-like domain of human Hydin as the template (). Although the human Hydin ASH/PapD domains and the CeSPD-2 ASH domain share only about 20% sequence similarity (), these superimposed ASH and Pap-D domains show overall high structural similarity (RMSD = 1.125 Å) (). Further support for the modeled 3D structure is that the most conserved residues (F5, N22, and P53) are in similar location and orientation in the CeSPD-2 and OCRL-1 ASH domains (). Although the topology of Hydin ASH/Pap-D and CeSPD-2 ASH β-sheets differ, both domains form an antiparallel β-sandwich conformation with a predicted classic type parallel β-bulge [Citation54] similarly located in CeSPD-2 ASH and in Hydin PapD-like domain (). The β-bulge in the modeled CeSPD-2 ASH domain forms between residues A4, F5 on β-sheet 1 and W91 on β-sheet 7 (). β-bulges often form between a residue in a β-sheets and two residues from an antiparallel β-sheet, and slightly increases the usual right-handed strand twist of the β-sheets. β-bulges are thought to affect the orientation of a binding site, help with dimerization, and help with accommodating single residue mutations [Citation54,Citation55].
Figure 3. Comparison between the three-dimensional model of the SPD-2 ASH domain and the known structure of ASH domains of HYDIN or OCRL-1.
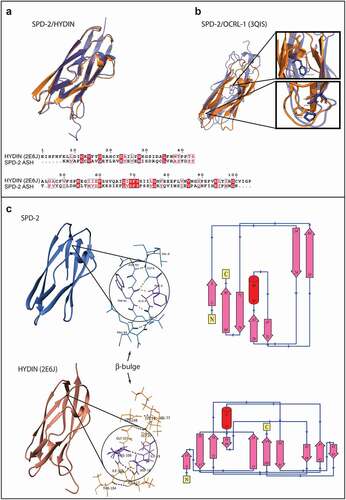
Figure 4. Electrostatic and structural similarities between the ASH domains from SPD-2 and other ASH domain containing proteins.
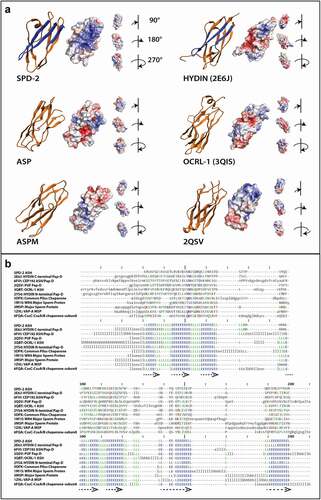
We also generated 3D model predictions for the ASH domains of the centrosome/cilia/flagella proteins ASPM and ASP from the previous alignment in (). The top template selected for the ASP ASH domain 3D model was the OCRL-1 ASH domain in complex with Rab8 (3QBT [Citation52]). Despite showing high structural similarity (RMSD = 2.285 Å), OCRL-1 and ASP ASH domains shared only 20.40% sequence similarity (). Interestingly, the C-terminal domain of a PapD-like domain from a protein of unknown function in Porphyromonas gingivalis (2QSV [Citation56]) was selected as the best template for the ASH domain of ASPM (RMSD = 3.0 Å) ().
ASH domains conserved Immunoglobulin-like structural fold overlaps with that of homologous superfamilies of PapD, MSP and Chaperone Domains
Because the ASH domains from proteins with centrosome/cilia/flagella function or localization show low sequence conservation but higher secondary structure conservation, we performed a DALI structure-based search with the CeSPD-2 ASH domain 3D predicted structure. This search identified 10 domains of known structure analogous to CeSPD-2 ASH. These include at least three PapD domains, three MSP domains, and two bacterial pilus chaperone-usher proteins ( and Table S1). The C-terminal PapD-like domain of HsHYDIN protein (2E6J [Citation47]; Z-score: 15.4) and an ASH domain of HsCEP192 protein (6FVI [Citation43]; Z-score: 11.1), were the two closest neighbors for the CeSPD-2 ASH domain ( and Table S1). This is consistent with the HsHYDIN PapD domain being the chosen template for the top ranked SPD-2 ASH 3D model (). The domains identified based on structure-similarity show little to no conservation of primary sequence between ASH domains but highly conserved secondary structure alignment (), as observed in the sequence alignment of ASH domains from centrosome/cilia/flagella associated proteins (). Superimposition of the modeled CeSPD-2 ASH domain and the 10 closest structural domain neighbors, shown in Movie S1, provides a clear visual confirmation of how closely related the structure of these domains is. All 10 3D structures superimposed with low RMSD (≤2.1 Å) indicating a common structural fold (). Together, the DALI search for structurally similar domains to the CeSDP-2 ASH 3D model and the choice of PapD domains as templates for 3D top models for ASH domains are evidence that ASH, PapD, MSP, and chaperone-usher domains belong to the same Superfold.
CeSPD-2 ASH has a unique biochemical character that may have functional significance
To identify possible functional regions within the ASH superfold, we generated surface electrostatic potential maps for the predicted ASH domains of the centrosomal proteins CeSPD-2, ASP, and ASPM, and the domains of known structure that served as their 3D prediction structural templates HYDIN, OCRL-1, and Porphyromonas gingivalis W8354 protein (). The electrostatic profile of CeSPD-2 ASH shows a large basic groove that is formed by residues K1-C7 and Y71-N93 (). A much smaller basic groove is found at a similar location in the Hydin PapD-like domain (2E6J), and this groove is significantly reduced or absent in ASP, OCRL-1, ASPM, and 2QSV ().
To ask whether the basic groove might be involved in functional interactions, we considered whether the residues predicted to form protein–protein or protein-nucleic acids in the ASH domain overlap with the basic residues (). We found that half of the residues in the basic grove (15/30) are predicted to be involved in protein–protein interactions (), and under a third of the basic residues (7/30) are predicted protein–RNA interaction sites (). There are 19 overlapping residues within the ASH domain that are predicted to bind both protein and RNA, these include the 7 residues predicted to bind RNA located in the basic groove. Comparison of contact site predictions for several ASH containing proteins associated with centrosome/cilia/flagella function or localization and CeSPD-2 ASH (Figure S2) show no overlap between the predicted sites from ASH in SPD-2 and the ASH domains from either Homo sapiens or Xenopus CEP192 (SPD-2 homolog) nor HYDIN (top 3D template for SPD-2). Therefore, neither the electrostatic potential maps nor the contact site predictions define a general ASH domain-specific functional region.
Figure 5. Predicted contact sites for SPD-2 ASH domain.
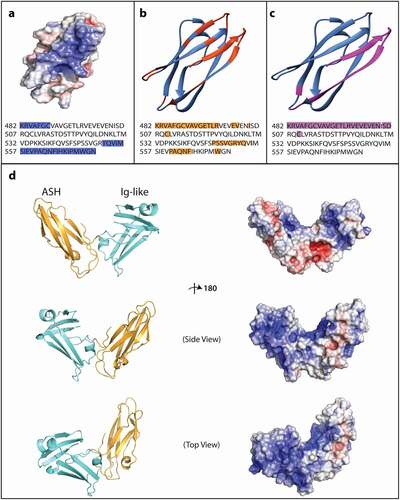
The DALI search for folds analogous to the SPD-2 ASH domain identified bacteria pili common chaperones and chaperone usher proteins (Table S1). These chaperones have two Ig-like or PapD domains oriented to one another in a functional structure that resembles a boomerang shape[Citation39]. Because several of the previously identified proteins have two or more Ig-like and/or ASH/PapD domains (e.g. HYDIN and CEP192), and CeSPD-2 has an Ig-like domain adjacent to the ASH domain, we wondered whether they also may form a boomerang structure analogous to the common and usher chaperones in bacteria. The 3D model of the CeSPD-2 ASH and Ig-like domains together also forms a boomerang-shape structure very similar to the bacterial chaperones identified in the DALI search () [Citation56]. Interestingly, the boomerang shape brings together several basic regions within the CeSPD-2 primary sequence. The basic region on the surface of the ASH domain extends into the cleft between the two domains, to the outward facing surface of the first β-sheet of the Ig-like domain (S2-K32), into the cavity between the first β-sheet and the second β-sheet (K57-V63), and ends at the short α-helix (M37-Q39) ().
The CeSPD-2 ASH domain localizes to centrosomes
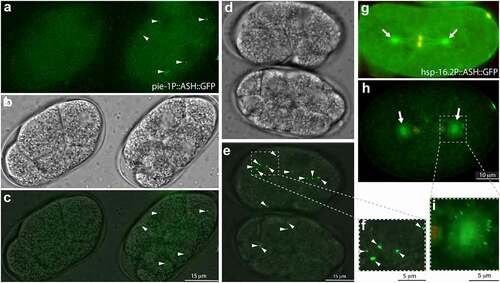
ASH domains have been proposed to have microtubule binding function [Citation38] and have been also shown to localize to centrosomes and cilia basal bodies in vitro [Citation34,Citation37,Citation38,Citation57]. To test whether the ASH domain of CeSPD-2 localizes to the centrosome, we expressed the ASH domain of CeSPD-2 as a fusion protein with GFP (ASH::GFP) in vivo (). Failure to generate a strain expressing GFP from a MosSCI insertion of an ASH::GFP construct driven by a germline promoter (Personal communication T. Mikeladze-Dvali) suggested that expression of the ASH::GFP protein might be toxic. We therefore expressed the ASH::GFP construct in a few cells of the animal and transiently upon induction. Strains expressing an extrachromosomal array of ASH::GFP driven by a germline-specific promoter yielded GFP expressing strains (see materials and methods) [Citation58,Citation59]. Further, suggesting that expression of ASH::GFP might be toxic was the very low germline transmission (≤10%) of the construct in the two strains obtained. In these strains, GFP was detected on the centrosomes and centrosome-associated microtubules in cells of embryos undergoing mitosis (). We also generated strains carrying a heat-shock inducible ASH::GFP transgene (see materials and methods) [Citation58,Citation59]. Brief heat-shock treatment induced expression of GFP that was detected as a single focus adjacent to the interphase nuclei (). Longer heat-shock treatments resulted in cytosolic GFP aggregates of variable size ().
Figure 6. Overexpressing the SPD-2 ASH domain in C. elegans embryos.
Discussion
CeSPD-2 is a bi-functional protein, with essential roles in centriole duplication and centrosome maturation. Although protein interactions required for each of the SPD-2 functions have been identified, the molecular mechanisms of its interactions and its regulation are largely unknown. For instance, it is unknown how specific interactions with SPD-2 that take place either exclusively at the centrioles or at the PCM are regulated.
CeSPD-2 intrinsic disorder may be important for its multifunctionality
We found that CeSPD-2 is highly disordered (50%), with most of the disordered regions being in the N-terminal half of the protein (72%) (). Many intrinsically disordered proteins display phase transition behaviors [Citation60]. One possibility is that CeSPD-2 might aid in the regulation of the phase-separation properties of the PCM [Citation61]. Consistent with this is the finding in PCM reconstitution experiments that CeSPD-2 and PLK-1 have been shown to either initiate or enhance SPD-5 formation of spherical condensates characteristic of proteins with phase transition properties [Citation12]. Additionally, similarly to regulators of transcription, translation, and cell cycle with disordered regions that act as hubs for different protein interactions [Citation59,Citation60], the highly disordered N-terminus of CeSPD-2 might provide the conformational plasticity necessary to accommodate alternative binding partners that operate in different processes [Citation14,Citation62–64]. For example, CDK targets often contain multiple phosphorylation sites within intrinsically disorder regions that allow for alternate phosphorylated outputs that produce different responses [Citation65,Citation66]. The N-terminus of CeSPD-2, expected to be intrinsically disordered, has predicted and experimentally confirmed CDK and PLK-1 phosphorylation sites () [Citation4,Citation14,Citation67]. This region is where PLK-1 binds to promote centrosome maturation [Citation63,Citation68] and ZYG-1/PLK4 binds to initiate centriole duplication [Citation23,Citation64]. We speculate that the intrinsically disordered N-terminus might allow alternate CDK (or other kinase) phosphorylated outputs that prime CeSPD-2 for either PLK-1 and/or ZYG-1/PLK-4 binding.
The SPD-2 domain[Citation4] ASH and Ig-like domains may be important for SPD-2 regulation.
Together, the Ig-like and ASH domains make up most of the “SPD-2 domain”, previously defined based on its shared similarity with the DsSPD-2 and HsCEP192 orthologs [Citation4]. The DALI structure-based search with the SPD-2 ASH 3D model identified bacterial pili PapD/usher chaperones domains as closely related folds. These chaperones are two Ig-like or PapD domains oriented toward each other that form a bilobed boomerang-like structure important for protein interactions during pilus assembly [Citation69]. We generated a 3D model of the SPD-2 ASH domain together with the Ig-like domain because of the similarity of the SPD-2 ASH domain to the PapD chaperones and because several proteins have more than one domain from the Immunoglobulin superfamily (e.g. CEP192, OCRL-1 and HYDIN). This 3D model strongly resembles the boomerang-shape of the PapD-like chaperones in bacteria (). The electrostatic map of the ASH and Ig-like domain 3D model predicts a large basic region that extends from the ASH domain (K1-C7; Y71-N93) to the boomerang shape cleft (S2-K32) and a large region of the exposed surface of the Ig-like domain (M37-Q39; K57-V63). This boomerang structure brings together and exposes several of the primary sequence basic regions in SPD-2 (). Interestingly, Shimanovskaya and colleagues (2014) [Citation64] proposed that ZYG-1 exclusive binding to the acidic N-terminus of CeSPD-2 localized at the centrioles (and not at the cytosol or PCM) might be regulated via intramolecular electrostatic interactions with one or several basic regions within SPD-2. Based on their proposal and the reported centriole duplication phenotypes of phospho-mutations within the ASH domain in polyploid intestinal cells [Citation67], we speculate that the boomerang structure might provide a mechanism to discriminate between cytosolic (or PCM) and centriolar CeSPD-2.
Identification of the ASH/PapD Superfold
Two main observations support our designation of the ASH/PapD superfold. First, top ranked 3D models for SPD-2ʹs and ASPM’s ASH domains utilized as templates PapD folds from the Hydin and W3 proteins with overall high structural similarity (RMSD < 2 Å) (). The SPD-2 ASH 3D model using Hydin PapD as template is also supported by the similar orientation and location of conserved amino acids in the ASH domains of CeSPD-2 and the distantly related protein OCRL1 (). Second, the top 10 matches in an unbiased search for folds structurally similar to CeSPD-2 ASH using DALI identified the ASH, PapD and chaperone usher bacterial pilus, and MSP domains. Despite MSP domains only sharing 11% sequence similarity with PapD/PapD-like domains, they have been classified as members of the PapD-like superfamily [Citation70]. Likeness to PapD chaperone domains [Citation71] is thought to be due to the potential of MSPs to bind interfaces used for filament biogenesis and to alter the rate of filament formation [Citation72]. Here we propose that the ASH domain, the PapD-like domain, the MSP domain, and common and usher pili chaperones domains from bacteria belong to the same superfold within the larger Immunoglobulin superfamily. Currently, there is not sufficient data to assign the Ig-like domain in CeSPD-2 and other ASH/PapD containing proteins to its own structural family.
ASH domains contact site predictions and localization to the centrosome
ASH domains have been proposed to have microtubule binding function [Citation38] and have been shown to localize to centrosomes and cilia basal bodies in vitro [Citation34,Citation37,Citation38,Citation57]. Our observations corroborate CeSPD-2 ASH localizing to the centrosome and centrosome associated microtubules in vivo. This is consistent with prior observations that mutations in ASH domains impair CeSPD-2 orthologue CEP192 localization to the centrosome and cilia [Citation3,Citation73], and similar observations for other ASH domain containing proteins in vitro [Citation34,Citation45,Citation57,Citation73,Citation74]. However, further studies are needed to establish the minimum requirement for its localization pattern of the ASH domain, given that localization might be dependent on its expression level and its expression might be toxic (). It might be useful to compare the localization of ASH on its own and together with the Ig-like domain, since other centrosome/cilia ASH proteins also have additional Ig-like, ASH, or PapD domains (e.g. HYDIN and CEP192) and the Ig-like domain and ASH in SPD-2 is predicted to form a boomerang shape analogous to the PapD and usher chaperones in bacteria pili that bind actin-like fibers.
It was slightly surprising that the predicted electrostatic surface and/or protein interacting residues of ASH domains from centrosomal, cilia, and flagella proteins did not reveal any shared properties that could be associated with their localization or microtubule binding properties. Are there shared properties that promote ASH/PapD folds to associate with filaments/microtubule containing structures? One possibility is that the identification of shared chemical properties is clouded by specific additional protein interaction binding requirements of each member of this superfold [Citation51,Citation75]. Alternatively, only physical structure properties shared within this superfold are required for their localization.
Testable hypotheses to uncover molecular mechanisms of SPD-2 regulation and function
Efforts to uncover molecular mechanisms of SPD-2 function are hindered by its dual roles in centrosome duplication and maturation, two co-dependent processes. The predicted domains and structural features identified in this study can be used to design rational hypothesis-based experiments aimed at uncovering molecular mechanisms of SPD-2 regulation and function. This analysis also hints at possible explanations for the functional diversity of SPD-2, because it predicted features often associated with proteins that are multifunctional or hubs for protein interactions.
Despite the sizable number of isolated spd-2 missense mutants, there are very few mutations affecting either centriole duplication or centrosome maturation exclusively. The structures and properties predicted here provide a logical set of residues in SPD-2 to target for mutational analysis. For instance, residues in the basic region of SPD-2 ASH predicted to be involved in protein interactions might affect the localization of SPD-2 or SPD-2 binding partners at the centrosome. The twenty-five amino acid stretch within the ASH domain predicted to bind RNA might also be a good target to test possible molecular interactions with the RNA binding protein, SZY-20, that antagonizes SPD-2 and ZYG-1 [Citation76].
The newly identified Ig-like, GEF-like, and PDZ-like domains might shed light to these domains’ roles in SPD-2 regulation and function. Several centrosome/cilia ASH proteins have more than one Ig-like/ASH/PapD domain and at least one other of these (OCRL-1) has a GTPase regulatory domain involved in vesicle trafficking. The putative GEF domain as well as the PDZ domains on their own and together in the same protein have been associated with processes that are consistent with known and putative SPD-2 functions. These include providing functional plasticity in multifunctional proteins [Citation41], regulating of polarized exocytosis [Citation77,Citation78] and mediating dynamics and reorganization of the microtubule and actin cytoskeleton [Citation79]. These domains will help identify new protein interactions or elucidate the regulation of known interactions and might provide new insight regarding a possible direct role for SPD-2 in anteroposterior polarization of the C. elegans embryo [Citation31,Citation33,Citation80].
Materials and methods
Secondary structure prediction and sequence analysis
The full-length C. elegans SPD-2 protein sequence was retrieved from the NCBI Reference Sequence (RefSeq) Protein sequence database (NP_492414.1; The C. elegans Sequencing Consortium, 1998) [Citation81]. The domain architecture for SPD-2 was analyzed using SMART (Simple Modular Architecture Research Tool) [Citation82], HHpred (v.2016) [Citation83], Pfam [Citation84], Prosite [Citation85], Interpro [Citation86], and the NCBI Conserved Domain (CD) database [Citation87], and a final consensus domain architecture cartoon was drawn to scale using Adobe Illustrator (Adobe Inc., 2019). Predictive secondary structure analysis of the full-length protein and ASH domain-specific sequence was used to confirm domain boundaries and assess the underlying secondary structure using PSIPRED [Citation88], PSSpred [Citation89], SPIDER2 [Citation90], Jpred4 [Citation91], and SABLE [Citation92]. Based on the domain architecture and secondary structure prediction results, the following sequence analysis was also performed for a more detailed sequence characterization: (a) Sequence disorder was predicted using Genesilico MetaDisorder [Citation93] and PrDOS [Citation94], and (b) Coiled-coils were predicted using COILS [Citation95], DeepCoil [Citation96], Paircoil2 [Citation97], and Multicoil [Citation98]. Potential sequence-based intermolecular interaction sites were assessed with ANCHOR [Citation99], DisoRDPbind [Citation100].
The ASH domain protein sequences of additional proteins were retrieved from NCBI: Human [Hs] hydrocephalus-inducing protein/HYDIN isoforms a/b and c/d (NP_001257903.1:513–613, NP_001185471.1:541–640), abnormal spindle-like microcephaly-associated protein/ASPM isoform 1/2 (NP_060606.3:37–134), inositol polyphosphate 5-phosphatase/OCRL-1 isoform a (NP_000267.2:564–678) and centrosomal protein 192kDa/CEP192 (NP_115518.3:2256–2402); Xenopus laevis [Xe] centrosomal protein/CEP192 (ADP36856.1:1776–1846); Mus musculus [Mm] hydrocephalus inducing protein/HYDIN (NP_766504.3:559–659); Drosophila melanogaster [Dm] abnormal spindle protein (NP_524488.3:25–122). Domain sequence alignments were done using the MUSCLE webserver [Citation101] and annotated with ESPript3 [Citation102]. A sequence similarity matrix was constructed using the SIAS (Sequence Identity and Similarity) webserver [Citation103].
Modeling and structure-based analysis
Human Hydin PapD-like C-terminal ASH domain (2E6J [Citation47]) was the best candidate template identified for modeling the SPD-2 ASH domain using various fold recognition algorithms. Three-dimensional structural models of the CeSPD-2 ASH domain were generated using template-based modeling programs as well as ab initio approaches with Modeller [Citation104], RAPTOR-X [Citation105], Phyre2 [Citation106], I-TASSER [Citation107], LOMETS [Citation53], and Swissmodel [Citation108]. Model quality was evaluated using ProSA-Web [Citation109], Verfiy3D [Citation110], ProQ3 [Citation111], and VoroMQA [Citation112]. The model with the best evaluation profile was further refined using ModRefiner [Citation113] and used for all subsequent analyses. The ab initio modeling algorithm in Robetta [Citation114] was also used to gather predicted structural data for the disordered domains in CeSPD-2. DEMO [Citation115] was used to assemble full-length models. Electrostatic potential maps were produced and visualized using the Adaptive Poisson-Boltzmann Solver (APBS) – PDB2PQR server [Citation116] and the PyMOL (v.1.8) molecular graphics system [Citation117]. Chimera (v1.11) [Citation118] was used for structure alignment and the production of backbone images. Potential contact sites based on the sequence and predicted structure of the model were predicted using SPPIDER (Solvent accessibility-based Protein–Protein Interface iDEntification and Recognition) [Citation119]. The same procedure was done for modeling ASH domains of Drosophila abnormal spindle protein/ASP and human abnormal spindle-like microcephaly-associated protein/ASPM.
Plasmid construct
The pie-1P::ASH::wGFP clone was generated by synthesis (Genewiz). This construct includes the pie-1 gene promoter in the pCM1.127 plasmid (Addgene), driving a translational fusion of the SPD-2 ASH domain (nucleotides 1444–1720) and the wGFP sequence isolated from the pCFJ1848 plasmid (optimized for germline expression [Citation59]). The 3ʹUTR in this construct contains the pie-1 3ʹUTR from the pCM5.47 plasmid (Addgene). The hsp-16.2P::ASH::GFP construct was generated by cloning the same SPD-2 ASH domain sequence (nucleotides 1444–1720) and the wGFP sequence into the PTB11 vector (Andrew Fire plasmid kit) that includes the hsp-16.3 promoter and the unc-54 3ʹUTR.
Culture conditions strains
C. elegans hermaphrodites were cultured using standard techniques [Citation120]. All strains were maintained at 16°C unless otherwise noted. The pie-1p::ASH::GFP plasmid was injected (15 ng/uL) into the OD95 strain – unc-119(ed3) III; Itls37 IV[(pAA64) pie-1p::mCherry::his-58 + unc-119(+)]; ltIs38 III [pie-1p::GFP::PH(PLC1delta1) + unc-119(+)] by NemaMetrix (Murray, UT). A rol-6 construct (pNU406) was used as the injection marker. The MSC25 strain that resulted from these injections transmits the extrachromosomal to about 10% of its progeny. MSC25 roller hermaphrodites were crossed with AV675 [mCherry::H2B] males and the resulting progeny was screened for ASH::GFP expression. The hsp-16.2p::ASH::GFP plasmid (2–10ng/l) was co-injected with the rol-6 (PRF4) injection marker (15–25ng/µL) into the germline of N2 nematodes. The resulting strain (MSC26) expressed ASH::GFP upon heat shock treatment. To reduce sequence repetitiveness in extra chromosomal arrays and prevent suppression of expression of the GFP constructs in the germline, sheered salmon sperm DNA was included to obtain a final injection mix concentration of 100 ng/µl [Citation58].
Heat shock treatment and gonad dissection
Gravid MSC6 hermaphrodites were heat-shocked at 30°C for 4 hours. After heat-shock, nematodes were recovered at 16°C for 1–2 hours. Recovered hermaphrodites were placed in 5 µl drop of M9 on a glass slide cover slip and were dissected to release the embryos. Coverslips with dissected MSC25 or MSC26 hermaphrodites were placed onto microscope slides with a 4% agar cushion sealed with Vaseline, and embryos were immediately imaged.
Imaging
GFP fluorescence was observed in about 1 embryo out of 10 of each the strains carrying an ASH::GFP construct, because of low transmission of the extra-chromosomal array. Given the low germline transmission of the arrays in all strains most embryos show no GFP fluorescence. For each condition 100% of the same pattern ASH::GFP expression was observed. At least seven animals were observed for each expression pattern. Z-stacks images of embryos were collected with the DeltaVision Deconvolution system mounted on an Olympus IX-70 inverted microscope equipped with a CoolSnap HQ CCD camera and a LED arc bulb illumination source, using 60 × 1.514NA oil immersion objective, GFP/mCherry fluorescence filter sets, and DIC optics. Image processing and z-stack projections were done with the SoftWorx software included in the Delta Vision Deconvolution system
Supplemental Material
Download Zip (1.7 MB)Acknowledgments
The authors would like to thank the Caenorhabditis Genetics Center funded by National Institute of Health (NIH) Office of Research Infrastructure Programs (P40OD010440) for strains, and Tamara Mikeladze-Dvali and Elif Nur Firat Karalar for feedback on the manuscript. This work was supported by a PSC-CUNY award TRADB-46-113 and NIH grant 1SC2GM118275-01.
Disclosure statement
No potential conflict of interest was reported by the authors.
Supplementary material
Supplemental data for this article can be accessed online at https://doi.org/10.1080/15384101.2022.2078458
Additional information
Funding
References
- Pintard L, Bowerman B. Mitotic cell division in Caenorhabditis elegans. Genetics . [Internet] 2019; 211:35–73. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=30626640&retmode=ref&cmd=prlinks
- Dammermann A, Müller-Reichert T, Pelletier L, et al. Centriole assembly requires both centriolar and pericentriolar material proteins. Dev Cell. [Internet] 2004; 7:815–829. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=15572125&retmode=ref&cmd=prlinks
- Kemp CA, Kopish KR, Zipperlen P, et al. Centrosome maturation and duplication in C. elegans require the coiled-coil protein SPD-2. Dev Cell. [Internet] 2004; 6:511–523. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=15068791&retmode=ref&cmd=prlinks
- Pelletier L, Özlü N, Hannak E, et al. The Caenorhabditis elegans centrosomal protein SPD-2 is required for both pericentriolar material recruitment and centriole duplication. Curr Biol. [Internet] 2004; 14:863–873. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=15186742&retmode=ref&cmd=prlinks
- Oegema K, Hyman AA. WormBook [Internet]. Available from: http://www.wormbook.org/chapters/www_celldivision/celldivision.html
- McNally KLP, Fabritius AS, Ellefson ML, et al. Kinesin-1 prevents capture of the oocyte meiotic spindle by the sperm aster. Dev Cell. [Internet] 2012; 22:788–798. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=22465668&retmode=ref&cmd=prlinks
- Erpf AC, Stenzel L, Memar N, et al. PCMD-1 organizes centrosome matrix assembly in C. elegans. Curr Biol . [Internet] 2019; 29:1324–1336.e6. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=30982652&retmode=ref&cmd=prlinks
- Hannak E, Kirkham M, Hyman AA, et al. Aurora-A kinase is required for centrosome maturation in Caenorhabditis elegans. J Cell Biol . [Internet] 2001; 155:1109–1116. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=11748251&retmode=ref&cmd=prlinks
- Cabral G, Laos T, Dumont J, et al. Differential requirements for centrioles in mitotic centrosome growth and maintenance. Dev Cell. [Internet] 2019; 50:355–366.e6. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31303441&retmode=ref&cmd=prlinks
- Wueseke O, Zwicker D, Schwager A, et al. Polo-like kinase phosphorylation determines Caenorhabditis elegans centrosome size and density by biasing SPD-5 toward an assembly-competent conformation. Biol Open . [Internet] 2016; 5:1431–1440. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=27591191&retmode=ref&cmd=prlinks
- Woodruff JB, Wueseke O, Viscardi V, et al. Centrosomes. regulated assembly of a supramolecular centrosome scaffold in vitro. Science [Internet] 2015; 348:808–812. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=25977552&retmode=ref&cmd=prlinks
- Woodruff JB, Gomes BF, Widlund PO, et al. The centrosome is a selective condensate that nucleates microtubules by concentrating tubulin. Cell . [Internet] 2017; 169:1066–1077.e10. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=28575670&retmode=ref&cmd=prlinks
- Hannak E, Oegema K, Kirkham M, et al. The kinetically dominant assembly pathway for centrosomal asters in Caenorhabditis elegans is gamma-tubulin dependent. J Cell Biol . [Internet] 2002; 157:591–602. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=12011109&retmode=ref&cmd=prlinks
- Decker M, Jaensch S, Pozniakovsky A, et al. Limiting amounts of centrosome material set centrosome size in C. elegans embryos. Curr Biol . [Internet] 2011; 21:1259–1267. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=21802300&retmode=ref&cmd=prlinks
- Delattre M, Canard C, GOnczy P. Sequential protein recruitment in C. elegans centriole formation. Curr Biol. [Internet] 2006; 16:1844–1849. Available from. http://linkinghub.elsevier.com/retrieve/pii/S0960982206019646
- Pelletier L, O’Toole E, Schwager A, et al. Centriole assembly in Caenorhabditis elegans. Nature [Internet] 2006; 444:619–623. Available from. http://www.nature.com/doifinder/10.1038/nature05318
- Dammermann A, Maddox PS, Desai A, et al. SAS-4 is recruited to a dynamic structure in newly forming centrioles that is stabilized by the gamma-tubulin-mediated addition of centriolar microtubules. J Cell Biol. [Internet] 2008; 180:771–785. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=18299348&retmode=ref&cmd=prlinks
- O’Connell KF, Caron C, Kopish KR, et al. The C. elegans zyg-1 gene encodes a regulator of centrosome duplication with distinct maternal and paternal roles in the embryo. Cell . [Internet] 2001; 105:547–558. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=11371350&retmode=ref&cmd=prlinks
- Leidel S, GOnczy P. SAS-4 is essential for centrosome duplication in C. elegans and is recruited to daughter centrioles once per cell cycle. Dev Cell. [Internet] 2003; 4:431–439. Available from. http://linkinghub.elsevier.com/retrieve/pii/S1534580703000625
- Delattre M, Leidel S, Wani K, et al. Centriolar SAS-5 is required for centrosome duplication in C. elegans. Nat Cell Biol. [Internet] 2004; 6:656–664. Available from. https://www.nature.com/articles/ncb1146
- Sugioka K, Hamill DR, Lowry JB, et al. Centriolar SAS-7 acts upstream of SPD-2 to regulate centriole assembly and pericentriolar material formation. Elife [Internet] 2017; 6. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=28092264&retmode=ref&cmd=prlinks
- Kim T-S, Park J-E, Shukla A, et al. Hierarchical recruitment of Plk4 and regulation of centriole biogenesis by two centrosomal scaffolds, Cep192 and Cep152. Proc Natl Acad Sci USA [Internet] 2013; 110:E4849–57. Available from. :http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=24277814&retmode=ref&cmd=prlinks
- Sonnen KF, Gabryjonczyk A-M, Anselm E, et al. Human Cep192 and Cep152 cooperate in Plk4 recruitment and centriole duplication. J Cell Sci. [Internet] 2013; 126:3223–3233. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=23641073&retmode=ref&cmd=prlinks
- Dix CI and Raff JW. Drosophila Spd-2 Recruits PCM to the Sperm Centriole, but is Dispensable for Centriole Duplication. Curr Biol. 2007;17(20):1759–1764. DOI:10.1016/j.cub.2007.08.065
- Giansanti MG, Bucciarelli E, Bonaccorsi Set al, . Drosophila SPD-2 is an Essential Centriole Component Required for PCM Recruitment and Astral-Microtubule Nucleation. Curr Biol. 2008;18(4):303–309. DOI:10.1016/j.cub.2008.01.058.
- Motegi F, Zonies S, Hao Y, et al. Microtubules induce self-organization of polarized PAR domains in Caenorhabditis elegans zygotes. Nat Cell Biol. 2011;13:1361–1367.
- Tsai M-C, Ahringer J. Microtubules are involved in anterior-posterior axis formation in C. elegans embryos. J Cell Biology 2007; 179:397–402.
- Goldstein B. Embryonic polarity: a role for microtubules. Curr Biol. 2000;10:R820–2.
- Wallenfang MR, Seydoux G. Polarization of the anterior-posterior axis of C. elegans is a microtubule-directed process. Nature . 2000;408:89–92 . . Available from: https://www.nature.com/articles/35040562.
- Gross P, Kumar KV, Goehring NW, et al. Guiding self-organized pattern formation in cell polarity establishment. Nat Phys. 2019;15:293–300.
- Klinkert K, Levernier N, Gross P, et al. Aurora A depletion reveals centrosome-independent polarization mechanism in Caenorhabditis elegans. Elife . [Internet] 2019; 8. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=30801250&retmode=ref&cmd=prlinks
- O’Connell KF. The centrosome of the early C. elegans embryo: inheritance, assembly, replication, and developmental roles. Curr Top Dev Biol [Internet] 2000; 49:365–384. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=11005028&retmode=ref&cmd=prlinks
- Kapoor S, Kotak S. Centrosome Aurora A regulates RhoGEF ECT-2 localisation and ensures a single PAR-2 polarity axis in C. elegans embryos. Development [Internet] 2019; 146. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31636075&retmode=ref&cmd=prlinks
- Schou KB, Morthorst SK, Christensen ST, et al. Identification of conserved, centrosome-targeting ASH domains in TRAPPII complex subunits and TRAPPC8. Cilia [Internet] 2014; 3:6–12. Available from. https://ciliajournal.biomedcentral.com/articles/10.1186/2046-2530-3-6
- Wang B, He W, Prosseda PP, et al. OCRL regulates lysosome positioning and mTORC1 activity through SSX2IP‐mediated microtubule anchoring. Embo Rep. 2021;22:e52173.
- McCrea HJ, Paradise S, Tomasini L, et al. All known patient mutations in the ASH-RhoGAP domains of OCRL affect targeting and APPL1 binding. Biochem Biophys Res Commun. 2008;369:493–499.
- Verdier P, Morthorst SK, Pedersen LB. Targeting of ASH Domain-containing proteins to the centrosome. Methods Mol Biology Clifton N J. 2016;1454:15–33.
- Ponting CP. A novel domain suggests a ciliary function for ASPM, a brain size determining gene. Bioinformatics . [Internet] 2006; 22:1031–1035. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=16443634&retmode=ref&cmd=prlinks
- Bork P, Holm L, Sander C. The immunoglobulin fold. Structural classification, sequence patterns and common core. J Mol Biol. [Internet] 1994; 242:309–320. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=7932691&retmode=ref&cmd=prlinks
- Lee H-J, Zheng JJ. PDZ domains and their binding partners: structure, specificity, and modification. Cell Commun Signal [Internet] 2010; 8:8–18. Available from. https://biosignaling.biomedcentral.com/articles/10.1186/1478-811X-8-8
- Harris BZ, Lim WA. Mechanism and role of PDZ domains in signaling complex assembly. J Cell Sci. [Internet] 2001; 114:3219–3231. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=11591811&retmode=ref&cmd=prlinks
- Joukov V, Nicolo AD, Rodriguez A, et al. Centrosomal protein of 192 kDa (Cep192) promotes centrosome-driven spindle assembly by engaging in organelle-specific Aurora A activation. Proc Natl Acad Sci USA . [Internet] 2010; 107:21022–21027. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=21097701&retmode=ref&cmd=prlinks
- Kovalevskiy OV, Breugel M. ASH/PapD-like domain of human CEP192 [Internet]. 2020; Available from: https://www.wwpdb.org/pdb?id=pdb_00006fvi
- Gomez-Ferreria MA, Rath U, Buster DW, et al. Human Cep192 is required for mitotic centrosome and spindle assembly. Curr Biol. [Internet] 2007; 17:1960–1966. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=17980596&retmode=ref&cmd=prlinks
- Saunders RD, Avides MC, Howard T, et al. The Drosophila gene abnormal spindle encodes a novel microtubule-associated protein that associates with the polar regions of the mitotic spindle. J Cell Biol . [Internet] 1997; 137:881–890. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=9151690&retmode=ref&cmd=prlinks
- Zhong X, Liu L, Zhao A, et al. The abnormal spindle-like, microcephaly-associated (ASPM) gene encodes a centrosomal protein. Cell Cycle . [Internet] 2005; 4:1227–1229. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=16123590&retmode=ref&cmd=prlinks
- Li H, Sato M, Koshiba S, et al. Solution structure of the C-terminal PapD-like domain from human HYDIN protein [Internet]. 2020; Available from: https://ftp.wwpdb.org/pub/pdb/validation_reports/e6/2e6j/2e6j_full_validation.pdf.gz
- Li HTTKSWSHTKTYS. Solution structure of the N-terminal PapD-like domain of HYDIN protein from human [Internet]. Available from: https://ftp.wwpdb.org/pub/pdb/validation_reports/ys/2ys4/2ys4_full_validation.pdf.gz
- Dawe HR, Shaw MK, Farr H, et al. The hydrocephalus inducing gene product, Hydin, positions axonemal central pair microtubules. BMC Biol [Internet] 2007; 5. Available from. https://bmcbiol.biomedcentral.com/articles/10.1186/1741-7007-5-33
- Erdmann KS, Mao Y, McCrea HJ, et al. A role of the Lowe syndrome protein OCRL in early steps of the endocytic pathway. Dev Cell. [Internet] 2007; 13:377–390. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=17765681&retmode=ref&cmd=prlinks
- Hou X, Hagemann N, Schoebel S, et al. A structural basis for Lowe syndrome caused by mutations in the Rab-binding domain of OCRL1. EMBO J [Internet] 2011; 30:1659–1670. Available from. https://www.embopress.org/doi/full/10.1038/emboj.2011.60
- Pirruccello M, Swan LE, Folta-Stogniew E, et al. Recognition of the F&H motif by the Lowe syndrome protein OCRL. Nat Struct Mol Biol . [Internet] 2011; 18:789–795. Available from. https://www.nature.com/articles/nsmb.2071
- Zheng W, Zhang C, Wuyun Q, et al. LOMETS2: improved meta-threading server for fold-recognition and structure-based function annotation for distant-homology proteins. Nucleic Acids Res . [Internet] 2019; 47:W429–36. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31081035&retmode=ref&cmd=prlinks
- Richardson JS, Getzoff ED, Richardson DC. The beta bulge: a common small unit of nonrepetitive protein structure. Proc Natl Acad Sci. 1978;75:2574–2578.
- Chan AWE, Hutchinson EG, Harris D, et al. Identification, classification, and analysis of beta-bulges in proteins. Protein Sci. 1993;2:1574–1590.
- Pakharukova N, Garnett JA, Tuittila M, et al. Structural insight into archaic and alternative chaperone-usher pathways reveals a novel mechanism of pilus biogenesis. PLoS Pathog . [Internet] 2015; 11:e1005269. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=26587649&retmode=ref&cmd=prlinks
- Zhao L, Hou Y, McNeill NA, et al. The unity and diversity of the ciliary central apparatus. Philos Trans R Soc Lond B Biol Sci . [Internet] 2020; 375:20190164. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31884923&retmode=ref&cmd=prlinks
- Kelly WG, Xu S, Montgomery MK, et al. Distinct requirements for somatic and germline expression of a generally expressed Caernorhabditis elegans gene. Genetics. 1997;146:227–238.
- Zeiser E, Frøkjær-Jensen C, Jorgensen E, et al. MosSCI and gateway compatible plasmid toolkit for constitutive and inducible expression of transgenes in the C. elegans germline. PLoS ONE [Internet] 2011; 6:e20082. Available from. http://dx.plos.org/10.1371/journal.pone.0020082
- Quiroz FG, Li NK, Roberts S, et al. Intrinsically disordered proteins access a range of hysteretic phase separation behaviors. Sci Adv . [Internet] 2019; 5:eaax5177. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31667345&retmode=ref&cmd=prlinks
- Zwicker D, Decker M, Jaensch S, et al. Centrosomes are autocatalytic droplets of pericentriolar material organized by centrioles. Proc Natl Acad Sci USA . [Internet] 2014; 111:E2636–45. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=24979791&retmode=ref&cmd=prlinks
- Wright PE, Dyson HJ. Intrinsically disordered proteins in cellular signalling and regulation. Nat Rev Mol Cell Biol . [Internet] 2015; 16:18–29. Available from. https://www.nature.com/articles/nrm3920
- Boxem M, Maliga Z, Klitgord N, et al. A protein domain-based interactome network for C. elegans early embryogenesis. Cell [Internet] 2008; 134:534–545. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=18692475&retmode=ref&cmd=prlinks
- Shimanovskaya E, Viscardi V, Lesigang J, et al. Structure of the C. elegans ZYG-1 cryptic polo box suggests a conserved mechanism for centriolar docking of Plk4 kinases. Structure . [Internet] 2014; 22:1090–1104. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=24980795&retmode=ref&cmd=prlinks
- Kõivomägi M, Ord M, Iofik A, et al. Multisite phosphorylation networks as signal processors for Cdk1. Nat Struct Mol Biol . [Internet] 2013; 20:1415–1424. Available from. https://www.nature.com/articles/nsmb.2706
- Ferrell JE, Ha SH. Ultrasensitivity part II: multisite phosphorylation, stoichiometric inhibitors, and positive feedback. Trends Biochem Sci . [Internet] 2014; 39:556–569. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=25440716&retmode=ref&cmd=prlinks
- Lu Y, Roy R. Centrosome/cell cycle uncoupling and elimination in the endoreduplicating intestinal cells of C. elegans. PLoS ONE [Internet] 2014; 9:e110958–17. Available from. http://dx.plos.org/10.1371/journal.pone.0110958
- Lee KS, Park J-E, Kang YH, et al. Mechanisms of mammalian polo-like kinase 1 (Plk1) localization: self- versus non-self-priming. Cell Cycle . [Internet] 2008; 7:141–145. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=18216497&retmode=ref&cmd=prlinks
- Holmgren A, Kuehn MJ, Brändén CI, et al. Conserved immunoglobulin-like features in a family of periplasmic pilus chaperones in bacteria. EMBO J. [Internet] 1992; 11:1617–1622. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=1348692&retmode=ref&cmd=prlinks
- Bullock TL, Roberts TM, Stewart M. 2.5 A resolution crystal structure of the motile major sperm protein (MSP) of Ascaris suum. J Mol Biol . [Internet] 1996; 263:284–296. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=8913307&retmode=ref&cmd=prlinks
- Sauer FG, Barnhart M, Choudhury D, et al. Chaperone-assisted pilus assembly and bacterial attachment. Curr Opin Struct Biol. [Internet] 2000; 10:548–556. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=11042452&retmode=ref&cmd=prlinks
- Tarr DEK, Scott AL. MSP domain protein-1 from Ascaris suum and its possible role in the regulation of major sperm protein-based crawling motility. Mol Biochem Parasitol . [Internet] 2005; 143:165–172. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=16026867&retmode=ref&cmd=prlinks
- Moser SC, Bensaddek D, Ortmann B, et al. PHD1 links cell-cycle progression to oxygen sensing through hydroxylation of the centrosomal protein Cep192. Dev Cell. [Internet] 2013; 26:381–392. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=23932902&retmode=ref&cmd=prlinks
- Lechtreck K-F, Witman GB. Chlamydomonas reinhardtii hydin is a central pair protein required for flagellar motility. J Cell Biol . [Internet] 2007; 176:473–482. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=17296796&retmode=ref&cmd=prlinks
- Du M, Yuan Z, Werneburg GT, et al. Processive dynamics of the usher assembly platform during uropathogenic Escherichia coli P pilus biogenesis. Nat Commun. 2021;12:5207.
- Song MH, Aravind L, Müller-Reichert T, et al. The conserved protein SZY-20 opposes the Plk4-related kinase ZYG-1 to limit centrosome size. Dev Cell. 2008;15:901–912.
- Nair J, Müller H, Peterson M, et al. Sec2 protein contains a coiled-coil domain essential for vesicular transport and a dispensable carboxy terminal domain. J Cell Biol . [Internet] 1990; 110:1897–1909. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=1693620&retmode=ref&cmd=prlinks
- Walch-Solimena C, Collins RN, Novick PJ. Sec2p mediates nucleotide exchange on Sec4p and is involved in polarized delivery of post-Golgi vesicles. J Cell Biol . [Internet] 1997; 137:1495–1509. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=9199166&retmode=ref&cmd=prlinks
- Longhurst DM, Watanabe M, Rothstein JD, et al. Interaction of PDZRhoGEF with microtubule-associated protein 1 light chains: link between microtubules, actin cytoskeleton, and neuronal polarity. J Biol Chem. [Internet] 2006; 281:12030–12040. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=16478718&retmode=ref&cmd=prlinks
- O’Connell KF, Maxwell KN, White JG. The spd-2 gene is required for polarization of the anteroposterior axis and formation of the sperm asters in the Caenorhabditis elegans zygote. 2000;222:55–70. Available from: http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=10885746&retmode=ref&cmd=prlinks
- Sayers EW, Agarwala R, Bolton EE, et al. Database resources of the national center for biotechnology information. Nucleic Acids Res [Internet] 2019; 47:D23–8. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=30395293&retmode=ref&cmd=prlinks
- Letunic I, Khedkar S, Bork P. SMART: recent updates, new developments and status in 2020. Nucleic Acids Res . [Internet] 2021; 49:D458–60. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=33104802&retmode=ref&cmd=prlinks
- Meier A, Söding J. Automatic prediction of protein 3D structures by probabilistic multi-template homology modeling. PLoS Comp Biol [Internet] 2015; 11:e1004343. Available from. https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1004343
- El-Gebali S, Mistry J, Bateman A, et al. The Pfam protein families database in 2019. Nucleic Acids Res [Internet] 2019; 47:D427–32. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=30357350&retmode=ref&cmd=prlinks
- Sigrist CJA, Cerutti L, de CE, et al. PROSITE, a protein domain database for functional characterization and annotation. Nucleic Acids research. [Internet] 2010; 38:D161–6. Available from. ;(suppl_1). http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=19858104&retmode=ref&cmd=prlinks
- Mitchell AL, Attwood TK, Babbitt PC, et al. InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res [Internet] 2019; 47:D351–60. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=30398656&retmode=ref&cmd=prlinks
- Lu S, Wang J, Chitsaz F, et al. CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res [Internet] 2020; 48:D265–8. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31777944&retmode=ref&cmd=prlinks
- Buchan DWA, Jones DT. The PSIPRED protein analysis workbench: 20 years on. Nucleic Acids Res . [Internet] 2019; 47:W402–7. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31251384&retmode=ref&cmd=prlinks
- Yan R, Xu D, Yang J, et al. A comparative assessment and analysis of 20 representative sequence alignment methods for protein structure prediction. Sci Rep . [Internet] 2013; 3. Available from. https://www.nature.com/articles/srep02619
- Yang Y, Heffernan R, Paliwal K, et al. SPIDER2: a package to predict secondary structure, accessible surface area, and main-chain torsional angles by deep neural networks. Methods Mol Biol [Internet] 2017; 1484:55–63. Available from. https://link.springer.com/protocol/10.1007/978-1-4939-6406-2_6
- Drozdetskiy A, Cole C, Procter J, et al. JPred4: a protein secondary structure prediction server. Nucleic Acids Res . [Internet] 2015; 43:W389–94. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=25883141&retmode=ref&cmd=prlinks
- Adamczak R, Porollo A, Meller J. Combining prediction of secondary structure and solvent accessibility in proteins. Proteins [Internet] 2005; 59:467–475. Available from. https://onlinelibrary.wiley.com/doi/full/10.1002/prot.20441
- Kozlowski LP, Bujnicki JM. MetaDisorder: a meta-server for the prediction of intrinsic disorder in proteins. BMC Bioinformatics [Internet] 2012; 13. Available from. https://bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-13-111
- Ishida T, Kinoshita K. PrDOS: prediction of disordered protein regions from amino acid sequence. Nucleic Acids Res . [Internet] 2007; 35:W460–4. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=17567614&retmode=ref&cmd=prlinks
- Lupas A, Dyke MV, Stock J. Predicting coiled coils from protein sequences. Science . [Internet] 1991; 252:1162–1164. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=2031185&retmode=ref&cmd=prlinks
- Ludwiczak J, Winski A, Szczepaniak K, et al. DeepCoil-a fast and accurate prediction of coiled-coil domains in protein sequences. Bioinformatics . [Internet] 2019; 35:2790–2795. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=30601942&retmode=ref&cmd=prlinks
- McDonnell AV, Jiang T, Keating AE, et al. Paircoil2: improved prediction of coiled coils from sequence. Bioinformatics . [Internet] 2006; 22:356–358. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=16317077&retmode=ref&cmd=prlinks
- Wolf E, Kim PS, Berger B. MultiCoil: a program for predicting two- and three-stranded coiled coils. Protein Sci [Internet] 1997; 6:1179–1189. Available from. https://onlinelibrary.wiley.com/doi/full/10.1002/pro.5560060606
- Dosztányi Z, Mészáros B, Simon I. ANCHOR: web server for predicting protein binding regions in disordered proteins. Bioinformatics . [Internet] 2009; 25:2745–2746. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=19717576&retmode=ref&cmd=prlinks
- Peng Z, Kurgan L. High-throughput prediction of RNA, DNA and protein binding regions mediated by intrinsic disorder. Nucleic Acids Res . [Internet] 2015; 43:e121. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=26109352&retmode=ref&cmd=prlinks
- Edgar RC. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics [Internet] 2004; 5:113–119. Available from. https://bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-5-113
- Robert X, Gouet P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res . [Internet] 2014; 42:W320–4. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=24753421&retmode=ref&cmd=prlinks
- Garcia-Boronat M, Diez-Rivero CM, Reinherz EL, et al. PVS: a web server for protein sequence variability analysis tuned to facilitate conserved epitope discovery. Nucleic Acids Res . [Internet] 2008; 36:W35–41. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=18442995&retmode=ref&cmd=prlinks
- Webb B, Sali A. Comparative protein structure modeling using MODELLER. Curr Protoc Protein Sci [Internet] 2016; 86:2.9.1–2.9.37. Available from. https://currentprotocols.onlinelibrary.wiley.com/doi/full/10.1002/cpps.20
- Adhikari B, Cheng J. CONFOLD2: improved contact-driven ab initio protein structure modeling. BMC Bioinformatics [Internet] 2018; 19:22–25. Available from. https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-018-2032-6
- Kelley LA, Mezulis S, Yates CM, et al. The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc . [Internet] 2015; 10:845–858. Available from. https://www.nature.com/articles/nprot.2015.053
- Yang J, Zhang Y. I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res . [Internet] 2015; 43:W174–81. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=25883148&retmode=ref&cmd=prlinks
- Waterhouse A, Bertoni M, Bienert S, et al. Swiss-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res [Internet] 2018; 46:W296–303. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=29788355&retmode=ref&cmd=prlinks
- Wiederstein M, Sippl MJ. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res . [Internet] 2007; 35:W407–10. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=17517781&retmode=ref&cmd=prlinks
- Lüthy R, Bowie JU, Eisenberg D. Assessment of protein models with three-dimensional profiles. Nature . [Internet] 1992; 356:83–85. Available from. https://www.nature.com/articles/356083a0
- Uziela K, Shu N, Wallner B, et al. ProQ3: improved model quality assessments using Rosetta energy terms. Sci Rep . [Internet] 2016; 6:33509–33510. Available from. https://www.nature.com/articles/srep33509
- Olechnovič K, Venclovas Č. VoroMQA web server for assessing three-dimensional structures of proteins and protein complexes. Nucleic Acids Res . [Internet] 2019; 47:W437–42. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31073605&retmode=ref&cmd=prlinks
- Xu D, Zhang Y. Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys J . [Internet] 2011; 101:2525–2534. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=22098752&retmode=ref&cmd=prlinks
- Song Y, Wang -RY-R, Kim D, et al. High-resolution comparative modeling with RosettaCM. Structure . [Internet] 2013; 21:1735–1742. Available from. https://www.sciencedirect.com/science/article/pii/S0969212613002979
- Zhou X, Hu J, Zhang C, et al. Assembling multidomain protein structures through analogous global structural alignments. Proc Natl Acad Sci USA . [Internet] 2019; 116:15930–15938. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=31341084&retmode=ref&cmd=prlinks
- Dolinsky TJ, Nielsen JE, McCammon JA, et al. PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res . [Internet] 2004; 32:W665–7. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=15215472&retmode=ref&cmd=prlinks
- Shrodinger. The PyMOL molecular graphics system, version 1.8 [Internet]. 2015. Available from: https://www.schrodinger.com/products/pymol
- Pettersen EF, Goddard TD, Huang CC, et al. UCSF Chimera—A visualization system for exploratory research and analysis. J Comput Chem [Internet] 2004; 25:1605–1612. Available from. https://onlinelibrary.wiley.com/doi/full/10.1002/jcc.20084
- Porollo A, Meller J. Prediction-based fingerprints of protein-protein interactions. Proteins [Internet] 2007; 66:630–645. Available from. https://onlinelibrary.wiley.com/doi/full/10.1002/prot.21248
- Brenner S. The genetics of Caenorhabditis elegans. Genetics . [Internet] 1974; 77:71–94. Available from. http://eutils.ncbi.nlm.nih.gov/entrez/eutils/elink.fcgi?dbfrom=pubmed&id=4366476&retmode=ref&cmd=prlinks
- Binkowski TA, Duggan E, Shui M, et al. Crystal structure of protein of unknown function from Porphyromonas gingivalis W83. 2011
