ABSTRACT
How plus-strand [+]RNA virus genomes transition from translation templates to replication templates is a matter of much speculation. We have previously proposed that, for Turnip crinkle virus, binding of the encoded RNA-dependent RNA polymerase (RdRp) to the 3′UTR of the [+]RNA template promotes a regional wide-spread conformational switch to an alternative structure that disassembles the cap-independent translation enhancer (CITE) in the 3′UTR. The active 3′CITE folds into a tRNA-like T-shaped structure (TSS) that binds to 80S ribosomes and 60S subunits in the P-site. In this Point-of-View, we discuss the history of our research on the TSS and our recent report combining coarse level single molecule force spectroscopy (optical tweezers) with fine-grain computer simulations of this experimental process and biochemical approaches to obtain a detailed understanding of how RdRp binding in the TSS vicinity might lead to an extensive rearrangement of the RNA structure.
Historical perspective on the Turnip Crinkle Virus (TCV) T-Shaped structure (TSS)
The research that culminated with Le et al., 2017Citation1 began 13 y ago when a well-known RNA biochemist interrupted a presentation being given by A.E. Simon with the pronouncement that a pseudoknot being proposed in the 3′UTR of TCV (Ψ2) was not physically possible due to the location and size of two nearby hairpins (H4b and H5) (). Since genetic data backed up the importance of all 3 elements for accumulation of TCV in single cell protoplasts,Citation2 the question became whether all 3 elements could physically co-exist in the plus-strand [+]RNA genome or if some were present in an alternative conformation (or in the complementary [−]-strand). Using our massively parallel genetic algorithm MPGAfoldCitation3 to predict the secondary structure along with RNA2D3DCitation4 to predict a 3D structure model, and multiple rounds of model stability evaluation with molecular dynamics (MD) simulations,Citation5 we found that the three elements, when combined with the adjacent upstream hairpin H4a and Ψ3, folded into a stable T-shaped structure (TSS) that was superimposable on canonical tRNAsCitation6 (). Binding of the TSS to 80S ribosomes and 60S ribosome subunits in the P-siteCitation7 provided a function for this complex element, since we determined that the TCV 3′UTR contained a strong cap-independent translation enhancer (3′CITE). The TSS significantly enhanced translation of reporter open reading frames (ORFs) in constructs containing the TCV 3′ 400 nt and the 63-nt 5′UTR, and was equally active when associated with 5′UTR of other carmoviruses despite there being little or no sequence similarity. No obvious RNA:RNA interactions connected the two ends of the TCV genomic RNA, unlike the long-distance RNA bridges present in many other viruses in the Tombusviridae.Citation8–10 A binding site for 40S ribosomal subunits in the TCV 5′UTR suggested a model whereby ribosomal subunits binding to the 5′ and 3′ ends might join to circularize the RNA thus contributing to ribosome recycling.Citation11
Figure 1. The TSS serves as a central hub for interactions throughout the TCV 3′UTR. (A) Secondary structure of the TSS. Tertiary interactions (Ψ2 and Ψ3) are shown. Values in parentheses are the predicted contour lengths of H5, H4b, and H4a/Ψ3. This RNA fragment is the one subjected to optical tweezer (OT) experiments in Le et al.Citation1 as denoted by the arrows. (B) Initial RNA2D3D model of the TSS in the absence of the 5 upstream adenylates. (C) An MD state of the new model of the TSS from C5 to A112. This was the fragment subjected to steered molecular dynamics (SMD), as denoted by the arrows. (D) In-line probing of a 3′UTR TCV fragment in the presence and absence of the RdRp. Bands denote that the radiolabeled fragment was self-cleaved due to flexible residues aligning to allow for a nucleophilic attack on the RNA backbone. L, hydroxide-treated ladder; T1, partial RNase T1 digest of a linearized fragment to denote location of guanylates; IL, in-line probed samples that were incubated at 25°C for 14 h. This result was originally reported in Yuan et al.Citation12 (E) Interactions between different elements near the 3′ end of TCV. The stop codon for the 3′-most ORF (the coat protein ORF) is shown. TSS residues are in red. Upstream 5As are in orange. Dark blue lines/arrows connect elements where structural changes are evident (arrowhead) when the connecting element is disrupted. Connections between pseudoknots denote that disrupting one pseudoknot (by altering base-paired residues) affects the connecting pseudoknot (i.e., residues in the connecting pseudoknot gain flexibility). Double arrowhead indicates that alterations in both locations affect the structure in the connecting location. Light blue lines connect compensatory second-site changes (arrowheads) with primary mutations.
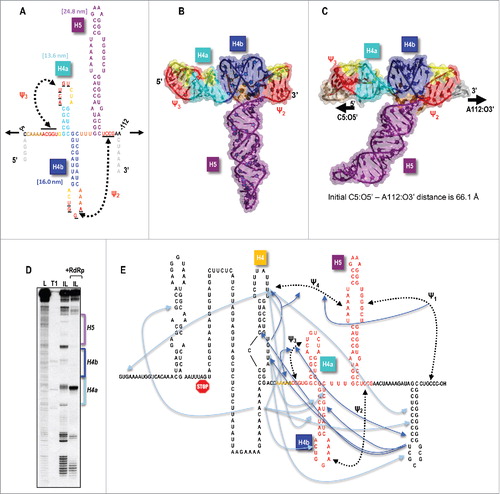
When the TCV-encoded RNA-dependent RNA polymerase (RdRp) was added to fragments containing (at least) the 200-nt 3′UTR of TCV, a major rearrangement of the RNA was discernable by in-line structure probing that extended from near the 3′ end of the UTR to the 5′ end of the longest fragment testedCitation12,13 (). This substantial wide-spread rearrangement, which did not occur when ribosomes were added to the RNA, begged the question of how the RdRp could promote such a substantial change in RNA structure throughout the fragment. This question led to a series of structural and genetic studies where mutations disrupting elements within and outside the TSS were evaluated for effects on structure, translation, replication and genesis of compensatory second-site mutations. Surprisingly, non-canonical interactions between many 3′ proximal RNA elements and the TSS were found, leading to the proposal that the TSS serves as a central hub for interactions throughout the region ().Citation12 Another intriguing finding was that a series of 5 adenylates (5A) just upstream from H4a/Ψ3 (which is located at the 5′ end of the TSS), were critical for [+]RNA accumulation. In addition, altering the adenylates caused structural changes to H4a/Ψ3 and vice-versa.
Since the TSS appeared to play a central role in RNA interactions in the 3′UTR of TCV (), it seemed logical that disruption of the highly stable TSS would be a necessary prelude to any widespread alterations in RNA structure. To gain an understanding of how the different components of the TSS, possibly together with the upstream 5As, interacted to produce a stable T-shaped structure and how this structure might be destabilized, we decided to use two approaches that we hoped would be complementary and inform each other: coarse level single molecule force spectroscopy, which would allow us to examine the unfolding and folding of single TSS molecules using optical tweezers; and fine-grain computational simulations of this experimental approach, which was hoped might reveal more detailed information on specific interactions.
Single molecule force spectroscopy- The coarse grain experimental approach
Single molecule techniques have emerged as powerful tools for understanding behaviors of biologic molecules by directly probing their folding pathways and mechanisms of molecular machinery. Instead of measuring ensemble averaged behaviors, single molecule techniques allow for examination of activities of individual molecules. A recently developed laser machine, “optical tweezers (OT),” can probe the structure of single RNA molecules by repeated rounds of stretching the molecule (by moving apart beads attached to the RNA ends), and then releasing the force and letting the RNA structure reassemble.Citation14–17 The extension lengths of individual elements (e.g., hairpins) within the TSS obtained under the force-induced unfolding/folding process are related to the number of nucleotides participating in unfolding/folding. Disassembly of an element (as the RNA element is physically stretched) appears as a “rip” in the force extension curve (FEC) or as “hops” when the force is held constant and elements that disassemble at that force level are allowed to transition repeatedly between folded and unfolded states.
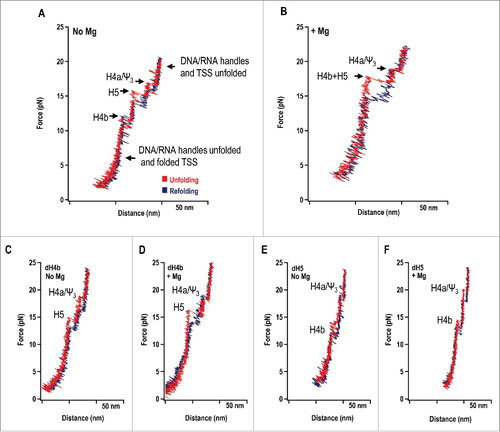
Unexpectedly, the FECs revealed that H4b unfolded first, followed by H5 and then H4a/Ψ3, the latter of which had the highest (least negative) predicted ΔG (). We also determined that H4b and H5 disassembled cooperatively as a single rip in Mg2+ (), but only in the presence of wild-type H4a/Ψ3 (not shown). Assignments of hairpins to corresponding rips were made following hybridization of complementary oligonucleotides to disrupt the hairpins (). For example, oligonucleotides complementary to H4b or H5 eliminated the first and middle rips, respectively in the absence of Mg2+ (Compare and with ). In the presence of Mg2+, disrupting H4b or H5 generated rips in the locations of individual H5 and H4b, respectively, and eliminated the large rip just below that of H4a/Ψ3 (compare and with ). In the case of H4a/Ψ3, the element was too stable to bind the oligonucleotide and mutations were used to disrupt the element. In addition, using truncated wild-type and mutant fragments that terminated just downstream of H4a, we determined that H4a/Ψ3 formed in the absence of H4b/H5, but did not form when the upstream 5As were converted to 5Us.Citation1 OT also revealed that the experimental contour lengths of H4a/Ψ3 and H5 were about 1/3 shorter than predicted, suggesting that the intact, well-studied structures shown in were not the structures unfolding in their assigned rips.Citation1 It is worth noting that the original RNA2D3D modelCitation6 () and the NMR TSS structureCitation18 excluded the 5As, since their importance at the time had not yet been realized. The TSS fragment used in the NMR study was not initially stable, and two additional 5′ guanylates needed to be added that expanded the pseudoknot at the expense of H4a.Citation18
Figure 2. Unfolding/folding of the TSS using optical tweezers. (A) Unfolding/refolding of the TSS RNA in the absence of Mg2+ and (B) presence of Mg2+. The TSS was attached to 500 bp RNA/DNA “handles,” which were attached to the beads. One bead is held in place and the other is in an optical trap, movable using a piezo. The red line in the force extension curve (FEC) represents the unfolding curve, in which force is applied that stretches the TSS and attached handles. The blue line represents the folding curve, in which force is released, allowing the TSS to refold. The specific “rips” in the FECs are labeled. (C) FEC in the presence of an oligonucleotide complementary to the complete H4b hairpin in the absence of Mg2+ or (D) presence of Mg2+. (E) FEC in the presence of an oligonucleotide complementary to the entire H5 hairpin in the absence of Mg2+ or (F) presence of Mg2+.
Computer simulations of single-molecule force spectroscopy: The fine-grain approach
Our hope was that molecular modeling and a form of molecular dynamics (steered molecular dynamics, or SMD) could provide higher resolution insight into the TSS unfolding process (for example, reveal canonical and non-canonical hydrogen bond interactions between nucleotides) thus providing suggestions for the unexpected stability of H4a/Ψ3 and reduced contour lengths for H4a/Ψ3 and H5. Repeating MD simulations using the same fragment as was used for the OT experiments () revealed a potential placement of the five adenylates in the major groove of Ψ3 (). A smaller subset of the fragment (that eliminated unpaired residues at the termini) was then used in SMD-based simulations of the pulling experiments. From the onset, the major issue was that simulated pulling speeds could not match the very slow OT extension rates because the run times for individual simulations would be prohibitively long even on the fastest processors (and multiple runs were required for statistical evaluation). Thus, we needed to establish the simulated pulling speed at which SMDs would reproduce the experimentally observed order of TCV TSS unfolding, as the simulated pulling speed determines the total simulation time (i.e., computation time). Even the slowest practical explicit solvent MD, which accounts for all atoms including water and salt, as implemented in the Amber 14 packageCitation19 was still insufficiently slow, and therefore this computationally more intensive (but possibly more accurate) method turned out to be impractical for simulating pulling of an RNA structure the size of the TCV TSS.
A much faster method based on the implicit solvent Generalized Born model (GB) (i.e., using an analytical approximation of solvent-solute interactions) allowed us to slow down the simulated pulling speed by an order of magnitude, and led to reproduction of the OT experimental results at the macro scale. We do not want to claim that GB simulations are generally better than the explicit solvent dynamics for this purpose, but because the slower simulated pulling speeds were crucial to reproducing the experimental results, the intrinsically faster GB-based simulations extended the practical range of simulated pulling speeds while being sufficiently accurate for use with RNA. The explicit solvent SMD runs, however, did provide reference trajectories helping to validate the GB-based simulations. It should be noted that this is a novel application of implicit solvent molecular dynamics to simulations of OT experiments on nucleic acids.
An illustration of the impact of pulling speed on the unfolding order is shown in . At faster pulling speeds, following the opening of Ψ2, H4b either cleanly separated from the rest of the TSS or formed an alternative pseudoknot (Ψalt) leading to a simulated opening pathway not observed by OT (, right). In contrast, in the majority of the runs with the slowest constant pulling speeds of 0.01 and 0.005 Å/ps, the 5′ end H4a/Ψ3 structure opened last, and the fully correct sequence of unfolding events was reproduced in 40 to 50% of all the simulations (depending on the GB parameters used) (, left and ).
Figure 3. Illustration of the impact of the simulated pulling speed on the unfolding order. The correct sequence of events achieved at lower pulling speeds (SMD run at 0.005 Å/ps illustrated) is marked with green arrows, while red arrows indicate the most common difference in simulations using pulling speeds higher than 0.01 Å/ps. An alternative pseudoknot between H4b and the 5′ side of H5 (and up-stream nucleotides) is more frequent at higher simulated pulling speed. At lower pulling speeds, simulations agreed with OT experiments and predicted that the partly re-arranged H4a/Ψ3 was the last TCV TSS structural element to open.
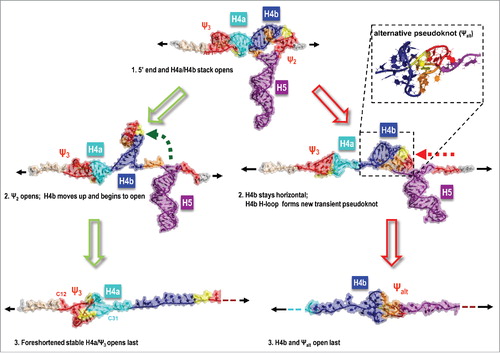
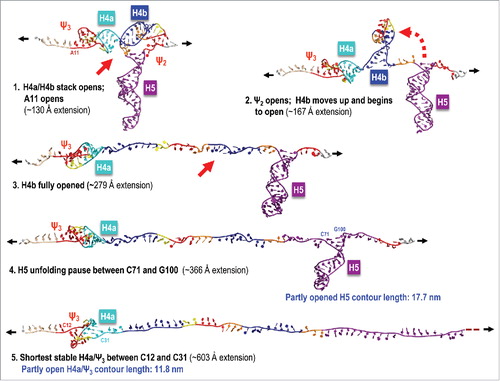
Figure 4. Illustration of a simulated TCV TSS pulling SMD run at 0.005 Å/ps with a correct order of unfolding events. In the majority of simulations, 5As and H4a/H4b stacking opened first,Citation1 followed by pseudoknot Ψ2 openingCitation2 and H4b unravelling in full.Citation3 H5 opens partly to form a substructure between C71 and G100 that offers more resistance to further opening.Citation4 After H5 is opened fully, a partly opened and re-arranged H4a/Ψ3 rapidly opens last.Citation5 The blue labels next to the last 2 states illustrated (4 and 5) indicate partly foreshortened contour lengths that agree well with the experimental measurements.
The parallel orientation of the H4a/Ψ3 helices with respect to the direction of pulling in the simulations provided a logical explanation for the increased stability of H4a/Ψ3 under pulling conditions compared with its calculated free energy, consistent with the OT study by Chen et al.Citation20 The 5′ end 5As and C6 were the first TSS nucleotides to open their triple interactions and thus 5A did not itself stabilize H4a/Ψ3 in the simulations. Whereas modeling and MD provided evidence for intercalation of the 5A in Ψ3 (), their early release from H4a/Ψ3 (without disassembly of H4a/Ψ3) conflicted with OT data that H4a/Ψ3 is unstable in the absence of 5A.Citation1 This suggests that 5A interaction with H4a/Ψ3 may have an unusual structure that will require further analysis by NMR or crystallography.
The SMD simulations that recapitulated the experimentally observed sequence of unfolding provided a detailed picture of individual events, some of which would not lead to discernable rips or hops in OT results. As described above, OT indicated that the contour lengths of H5 and H4a/Ψ3 were about 1/3 shorter than predicted, suggesting that the structures unfolding in observable rips/hops were not the full structures shown in . For H5, simulations showed that the lower stem unzipped before the majority of the stem-loop unraveled, with a delayed opening of the upper (terminal) C-G base pairs. This suggests that the H5 rips/hops did not reflect the entirety of the H5 hairpin. SMD simulations also showed a partial unzipping of H4a and shifts in the Ψ3 pairing (“slipping”) preceding the final opening rip (), which could contribute to the experimental difference observed. This slipping maintained a pseudoknot at the 5′ end and the resulting high resistance to pulling, while not maintaining any of the original paired nucleotides or even canonical Watson-Crick interactions. Whereas the alignment between the motif axis and the direction of pulling leads to what could be termed as “static friction,” the sliding pseudoknots in which new nucleotide interactions are formed as the original interactions open while the general topology is maintained could be thought of as “kinetic friction.” Both static and kinetic friction influenced the order of unfolding in our simulations, with H4a/Ψ3 being only one example.
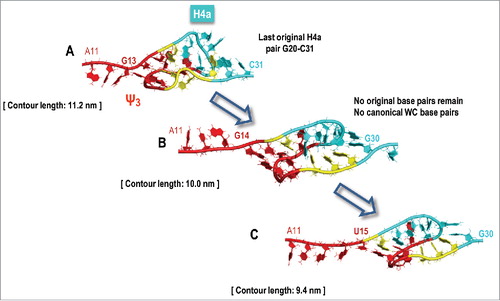
Figure 5. Illustration of a simulated opening of H4a/Ψ3. SMD predicts partial opening of H4a/Ψ3, foreshortening its contour length first to 11.2–11.8 nm (19–20 nts), and then, in some runs, down to 9.4 nm before the remaining structure opens rapidly. (A) Partial opening of H4a/Ψ3 foreshortens its contour length. (B-C) Slipping of H4a/Ψ3 precedes the final opening rip. This slipping maintains the pseudoknot at the 5′ end and the resulting high resistance to pulling. This scenario could explain the difference between the estimated (13.6 nm) and measured (9.1 ± 1.5 nm) H4a/Ψ3 contour length.
Because of the lack of reliable information on the Mg2+ ion placement around the TCV TSS and the prohibitive simulation times that would be required to predict their placement ab initio, we did not attempt to model the influence of magnesium in our explicit solvent runs. However, based on the observed impact of the alternative of tertiary interactions involving H4b in some SMD simulations, we can speculate that Mg2+ could strengthen Ψ2 interactions and potentially alter the order of TSS building blocks opening, possibly leading to the H4b/Ψ2 and H5 opening in parallel, which was observable as one rip by OT (). At the same time the Mg2+-strengthened H4a/Ψ3 could still offer proportionately more resistance to pulling and open last.
Model for the RdRp-mediated conformational switch
OT revealed that the 5As upstream of H4a/Ψ3 were critical for the stability of H4a/Ψ3 and that H4a/Ψ3 was needed for cooperativity between H4b and H5. This suggested that RdRp binding to the 5A/H4a/Ψ3 region could lead to disassembly of the TSS and disrupt its role as a central hub for RNA interactions in the region, leading to the conformational switch. Mutations in the 5A region eliminated one-site RdRp binding to a 3′UTR fragment as did mutations just downstream that disrupted Ψ3.Citation1 Surprisingly, mutations in the Ψ3 partner bases within the H4a loop had the opposite effect, enhancing RdRp binding to the fragment. This indicated that simple disruption of Ψ3 was not the reason for loss of specific binding, but rather that RdRp binding to the adenylates and downstream sequence that also participates in Ψ3 might be the key to the conformational shift.
Conclusions
This study demonstrates the usefulness of combining biochemical, genetic, coarse level (OT) and fine-grain (SMD) approaches to shed light on a complex biologic switch. Our ability to model TSS disassembly that recapitulates the unexpected order of unfolding and the unexpected shortened contour lengths revealed by OT could potentially greatly simplify examination of the folding/unfolding dynamics of other complex RNA structures.
Disclosure of potential conflicts of interest
No potential conflicts of interest were disclosed.
Funding
This work was supported by the National Science Foundation (MCB-1411836) and National Institutes of Health (R21AI117882–01) to A.E.S. Support for this research was also provided by NIH training grant (NIGMS T32GM080201) to M. T. L. This project has been funded in part with federal funds from the Frederick National Laboratory for Cancer Research, National Institutes of Health, under contract HHSN261200800001E for W.K.K. This research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US. Government.
References
- Le M-T, Kasprzak WK, Kim T, Gao F, Young MYL, Yuan X, Shapiro BA, Seog J, Simon AE. Folding behavior of a T-shaped, ribosome-binding translation enhancer implicated in a wide-spread conformational switch. eLIFE. 2017;6:e22883. doi:10.7554/eLife.22883.
- Simon AE, Howell SH. The virulent satellite RNA of turnip crinkle virus has a major domain homologous to the 3′ end of the helper virus genome. EMBO J. 1986;5:3423–8.
- Shapiro BA, Kasprzak W, Grunewald C, Aman J. Graphical exploratory data analysis of RNA secondary structure dynamics predicted by the massively parallel genetic algorithm. J Mol Graph Model. 2006;25:514–31. doi:10.1016/j.jmgm.2006.04.004.
- Martinez HM, Maizel JV, Shapiro BA. RNA2D3D: A program for generating, viewing, and comparing 3-dimensional models of RNA. J Biomol Struct Dyn. 2008;25:669–83. doi:10.1080/07391102.2008.10531240.
- Case DA, Darden TA, Cheatham TEI, Simmerling CL, Wang J, Duke RE, Luo R, Crowley M, Walker RC, Zhang W, et al. 2008. AMBER 10, University of California, San Francisco
- McCormack JC, Yuan X, Yingling YG, Kasprzak W, Zamora RE, Shapiro BA, Simon AE. Structural domains within the 3′ untranslated region of Turnip crinkle virus. J Virol. 2008;82:8706–20. doi:10.1128/JVI.00416-08.
- Stupina VA, Meskauskas A, McCormack JC, Yingling YG, Shapiro BA, Dinman JD, Simon AE. The 3′ proximal translational enhancer of Turnip crinkle virus binds to 60S ribosomal subunits. RNA. 2008;14:2379–93. doi:10.1261/rna.1227808.
- Chattopadhyay M, Shi K, Yuan X, Simon AE. Long-distance kissing loop interactions between a 3 ′ proximal Y-shaped structure and apical loops of 5 ′ hairpins enhance translation of Saguaro cactus virus. Virology. 2011;417:113–25. doi:10.1016/j.virol.2011.05.007.
- Gao F, Kasprzak W, Stupina VA, Shapiro BA, Simon AE. A ribosome-binding, 3 ′ translational enhancer has a T-shaped structure and engages in a long-distance RNA-RNA interaction. J Virol. 2012;86:9828–42. doi:10.1128/JVI.00677-12.
- Fabian MR, White KA. 5′-3′ RNA-RNA interaction facilitates cap- and poly(A) tail-independent translation of tomato bushy stunt virus mrna: a potential common mechanism for tombusviridae. J Biol Chem. 2004;279:28862–72. doi:10.1074/jbc.M401272200.
- Stupina VA, Yuan X, Meskauskas A, Dinman JD, Simon AE. Ribosome binding to a 5 ′ translational enhancer Is altered in the presence of the 3 ′ untranslated region in cap-independent translation of Turnip crinkle virus. J Virol. 2011;85:4638–53. doi:10.1128/JVI.00005-11.
- Yuan XF, Shi KR, Meskauskas A, Simon AE. The 3′ end of Turnip crinkle virus contains a highly interactive structure including a translational enhancer that is disrupted by binding to the RNA-dependent RNA polymerase. RNA. 2009;15:1849–64. doi:10.1261/rna.1708709.
- Yuan X, Shi K, Young MYL, Simon AE. The terminal loop of a 3 ′ proximal hairpin plays a critical role in replication and the structure of the 3 ′ region of Turnip crinkle virus. Virology. 2010;402:271–80. doi:10.1016/j.virol.2010.03.036.
- Green L, Kim C-H, Bustamante C, Tinoco I, Jr. Characterization of the mechanical unfolding of RNA pseudoknots. J Mol Biol. 2008;375:511–28. doi:10.1016/j.jmb.2007.05.058.
- Greenleaf WJ, Frieda KL, Foster DAN, Woodside MT, Block SM. Direct observation of hierarchical folding in single riboswitch aptamers. Science. 2008;319:630–3. doi:10.1126/science.1151298.
- Li PTX, Bustamante C, Tinoco I. Unusual mechanical stability of a minimal RNA kissing complex. Proc Natl Acad Sci USA. 2006;103:15847–52. doi:10.1073/pnas.0607202103.
- Liphardt J, Onoa B, Smith SB, Tinoco I, Bustamante C. Reversible unfolding of single RNA molecules by mechanical force. Science. 2001;292:733–7. doi:10.1126/science.1058498.
- Zuo X, Wang J, Yu P, Eyler D, Xu H, Starich MR, Tiede DM, Simon AE, Kasprzak W, Schwieters CD, et al. Solution structure of the cap-independent translational enhancer and ribosome-binding element in the 3 ′ UTR of turnip crinkle virus. Proc Natl Acad Sci USA. 2010;107:1385–90. doi:10.1073/pnas.0908140107.
- Case DA, Babin V, Berryman JT, Betz RM, Cai Q, Cerutti DS, Cheatham I, Darden RE, Duke RE, Gohlke H, et al. 2014. AMBER 14, University of California San Francisco
- Chen G, Wen J-D, Tinoco I, Jr. Single-molecule mechanical unfolding and folding of a pseudoknot in human telomerase RNA. RNA. 2007;13:2175–88. doi:10.1261/rna.676707.
