SUMMARY
Maternal diabetes and/or obesity in pregnancy are undoubtedly associated with later disease-risk in the offspring. The placenta, interposed between the mother and the foetus, is a potential mediator of this risk through epigenetic mechanisms, including DNA methylation. In recent years, multiple studies have identified differentially methylated CpG sites in the placental tissue DNA in pregnancies complicated by diabetes and obesity. We reviewed all published original research relevant to this topic and analysed our findings with the focus of identifying overlaps, contradictions, and gaps. Most studies focused on the association of gestational diabetes and/or hyperglycaemia in pregnancy and DNA methylation in placental tissue at term. We identified overlaps in results related to specific candidate genes, but also observed a large research gap of pregnancies affected by type 1 diabetes. Other unanswered questions relate to analysis of specific placental cell types and the timing of DNA methylation change in response to diabetes and obesity during pregnancy. Maternal metabolism is altered already in the first trimester involving structural and functional changes in the placenta, but studies into its effects on placental DNA methylation during this period are lacking and urgently needed. Foetal sex is also an important determinant of pregnancy outcome, but only few studies have taken this into account. Collectively, we provide a reference work for researchers working in this large and evolving field. Based on the results of the literature review, we formulate suggestions for future focus of placental DNA methylation studies in pregnancies complicated by diabetes and obesity.
Introduction
Diabetes and obesity in pregnancy
The incidence of pregnancies with disturbed metabolic homoeostasis such as in women with Type 1 diabetes mellitus (T1DM), Type 2 diabetes mellitus (T2DM), gestational diabetes mellitus (GDM), and in overweight or obese women has been increasing worldwide [Citation1–5]. Metabolism in these women is often characterized by hyperglycaemia and hyperlipidaemia because of various degrees of and/or combinations of insufficient or absent beta cell function, or insulin resistance. In most instances insulin resistance is associated with elevated concentrations of circulating insulin [Citation6].
The risk of short-term complications of these pregnancies has been known for long. More recently, evidence has accumulated to demonstrate also long term consequences for mother and offspring throughout the life course [Citation6–8]. Specifically, in utero exposure to disturbed metabolism of the mother increases offspring risk for adiposity, T2DM, metabolic syndrome and cardiovascular problems, also with an influence of foetal sex [Citation9].
While the mechanisms underlying these adverse health conditions have not been fully understood, (epi)genetic transmission of risks has been demonstrated and the placenta has been implicated [Citation10,Citation11].
The placenta
The placenta is interposed between the maternal and foetal circulation and, hence, exposed to perturbations in both compartments albeit with different surfaces. The syncytiotrophoblast bathes in maternal blood in the intervillous space, whereas endothelial cells line the feto-placental vasculature and are under the influence of foetal circulating factors [Citation12]. The placenta’s broad array of diverse functions has been long recognized to play a major role in regulating maternal metabolism as well as foetal growth and development. Structural and functional changes of the placenta in T1DM, T2DM, GDM and in obese women have been regarded as contributing to foetal phenotype, mostly foetal overgrowth, in these conditions either directly by modifying foetal metabolism or indirectly by changing maternal insulin secretion or insulin resistance or both [Citation13,Citation14]. Evidence suggests that maternal metabolic adaptations during pregnancy, especially of glucose metabolism, are also partly regulated by foetal sex, as the risk of developing GDM and GDM severity is increased in women with male newborns [Citation9,Citation15]. Moreover, mothers of male offspring have increased risk of developing T2DM later in life when compared to mothers of female offspring [Citation16].
Epigenetic mechanisms in the placenta
Epigenetics is the study of ‘structural adaptations of chromosomal regions so as to register, signal or perpetuate altered activity states’[Citation17]. ‘Epigenetic marks’ is a broad term that includes DNA methylation (DNAm), histone post-translational modifications, RNA modifications, and non-coding RNAs. These marks are modified by specific enzymes which are recruited by transcription factors [Citation18].
The human genome contains ~28 million cytosine-phosphate-guanine (CpG) sites, comprising less than ~1% of the total genome. CpG content in the genome is lower than the expected ~4% due to natural deamination of methylated cytosine to thymine [Citation19]. Therefore, most CpG sites are scattered at low density across the genome, while a subset occur at high density in hypomethylated regions known as CpG Islands [Citation20]. Techniques that measure DNAm range from global (e.g., High-performance liquid chromatography (HPLC) to detect methylated cytosines as a percentage of all cytosines, or whole-genome sequencing), genome-wide arrays (e.g., Illumina arrays, which cover ~3% of all CpG sites) and locus-specific (e.g., bisulphite amplicon sequencing to detect DNAm at a specific genomic region) [Citation21–24]. All of these, except global measurement by HPLC, require bisulphite conversion of DNA as the first step [Citation25]. Due to the multiple different methodological approaches of studying DNAm, we specify throughout the review and in the approach that was used to generate the data.
Table 1. Literature search on studies of DNA methylation in placenta tissue and diabetes and obesity in pregnancy.
Table 2. Literature search on studies of DNA methylation in placenta tissue and hyperglycaemia and dyslipidemia in pregnancy.
Due to its early separation from the embryo at the blastocyst stage, the placenta has a unique DNAm profile, with several similarities to human tumours [Citation26], such as low global methylation [Citation27], partially methylated domains [Citation28], and tumour-suppressor methylation [Citation29]. Alterations of DNAm signatures often parallel transcriptional, morphological and functional changes of the placenta [Citation30–32]. Importantly, specific placental DNAm patterns have been associated with maternal exposures [Citation33] and offspring outcomes [Citation34]. Hence, measuring placental DNAm, including cell-free DNA in maternal circulation, is useful for assessing placental health [Citation35].
In this review we summarize current knowledge on variation of placental epigenetic profiles in pregnancies in women with diabetes and in obese women. Since maternal hyperglycaemia and hyperlipidaemia are hallmarks of metabolic changes in these conditions, we have included placental DNAm related to these perturbations. As the influence of foetal sex is increasingly being acknowledged, we also note how foetal sex was integrated into the statistical models. We focused on DNAm as the most stable epigenetic variation, which can be measured with high reproducibility. The list of genes with identified changes in DNAm in any of these conditions provides an up to date overview and shall help readers to easily assess whether their gene of interest shows DNAm changes. We conclude by highlighting areas for future research that have emerged from conducting this review.
Results
Literature search results
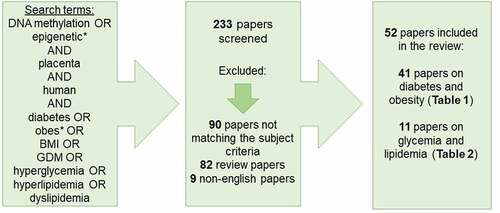
Using PubMed, we conducted a literature search covering publications between 1960 and 12 July 2022 of peer-reviewed original research of placental DNAm in pregnancies complicated by diabetes or obesity. We used following search terms: DNA methylation OR epigenetic* AND placenta AND human AND diabetes OR obes* OR BMI OR GDM OR hyperglycaemia OR hyperlipidaemia OR dyslipidemia. In total, 233 papers were identified using these search terms. All were screened for suitability to be covered in this review (). We included all studies regardless of aim and sample size with following inclusion criteria: original research papers, in English language and matching subject criteria. The screening excluded 90 papers not within the subject area, 82 review papers, and nine non-English language papers. In the end, 52 papers were found suitable and were included (). These were divided into two groups: case-control studies () and studies of continuous glucose/lipid measurements (). Finally, we merged all data on DNAm differences of specific genes/gene regions, and sorted on gene annotations to provide an overview of candidate genes investigated in multiple studies, as well as to assess whether specific genes and differential DNAm associated with exposures were replicated across studies ().
Table 3. Differentially methylated genes identified in literature search.
Diabetes and obesity phenotypes
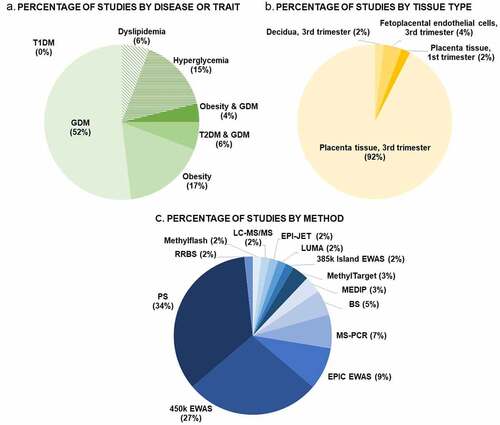
As outlined in , more than half (52%) of the 52 included studies focused on GDM. When combing with the other glucose intolerance phenotypes (T2DM and GDM combined studies (6%) and hyperglycaemia studies (15%)) 73% percent of the studies covered were focusing on aspects of glucose levels in pregnancy. A smaller proportion of studies (17%) focused on obesity, and only 4% of the papers focused on GDM and obesity combined. Important to notice, we were unable to find a study on placental DNAm with a focus on T1DM in pregnancy.
Figure 2. Percental overviews of number of studies conducted stratifying: A. by disease or trait, B. by placental tissue type, and C. by methodology. Abbreviations: GDM: Gestational Diabetes Mellitus, T1DM: Type 1 Diabetes Mellitus, T2DM: Type 2 Diabetes Mellitus, RRBS: reduced representation bisulphite sequencing, LC-MS/MS: liquid chromatography with tandem mass spectrometry, LUMA: Luminometric Methylation Assay, EWAS: epigenome wide association study, MEDIP: methylated DNA immunoprecipitation, BS, bisulphite sequencing, MS-PCR: methylation-specific PCR, PS: pyrosequencing.
Tissue and cell type specificity in Placenta DNAm
Of the 52 papers included, the vast majority (92%) studied DNAm in total placental tissue, mainly collected at term (). Hence, the DNAm results conducted in these studies provide an average DNAm percentage for all placental cell types [Citation36]. Only one study had been performed in first trimester placenta whole tissue biopsies. Only three studies had focused on specific cell types (all at term); one study of DNAm in decidua and two studies in feto-placental endothelial cells (). No studies investigated DNAm in trophoblasts, although this placental cell type is the primary target of alterations in the maternal circulation and has crucial functions for placental growth and development.
Methods of DNAm measurements
Almost half of DNAm studies in the placenta that were included in this review were performed with genome-wide methods. The two most common approaches were pyrosequencing (34% of studies) and the Illumina 450 K array (27% of studies). ().
Sex-specific differences
Only nine of the 52 studies included sex differences as their outcome in the analyses (+2), which is unfortunate since the foetal sex has an impact on placental DNAm. Indeed, DNAm plays a key role in X-chromosome inactivation, a process that achieves dosage compensation for X-encoded gene products between female and male cells [Citation37]. However, differential sex chromosome dosage complicates genome-wide epigenomic assessments as sex-specific methylation patterns on the X chromosome largely reflect the effects of X-chromosome inactivation [Citation38]. Therefore, the sex chromosomes are frequently excluded from statistical analyses to avoid sex bias. Almost half of the EWAS studies investigating GDM removed both X-and Y-chromosome probes prior to statistical analysis (). One study, which segregated the influence of foetal sex on placental DNAm in GDM, removed only X-chromosome associated probes [Citation39]. Another study included all probes in the analysis and report in total 10,424 differentially methylated regions (DMRs) in GDM placenta, out of which only 5% were annotated to autosomal chromosomes [Citation40]. In none of the EWAS studies investigating DNAm in dyslipidemia were X-and Y-chromosomes probes removed, although all studies report adjusted analyses for foetal sex. In contrast, in EWAS studies of hyperglycaemia effects X-and Y-chromosome probes were always removed prior to statistical analysis and, except one, all adjusted for foetal sex ().
Discussion and future perspectives
Similarities and differences of results across maternal phenotypes
Four different genes (PGC1A, PPARA, LEP and ADIPOQ) were studied in three or more separate papers (). Both PPARA (nuclear receptor peroxisome proliferator activated receptor-alpha) and PGC1A (PPAR-gamma coactivator-1-alpha) play important roles in transcriptional regulation of energy metabolism including regulation of mitochondrial biogenesis and liver gluconeogenesis [Citation41]. Indeed, in multiple studies PGC1A DNAm was directly associated with PGC1A mRNA expression and increased PGC1A promoter DNAm is positively associated with T2DM and physical inactivity [Citation42,Citation43]. Regarding PGC1A, four different studies were identified, of which two examined GDM pregnancies, one examined both GDM and T2DM and one examined various levels of glycaemia. Interestingly, all four studies showed an increase in placenta PGC1A promotor DNAm with hyperglycaemia and/or GDM compared to controls [Citation44–47], however, one study observed increased PGC1A DNAm in only placentas linked to male offspring [Citation46]. Regarding PPARA, notably all three studies conducted in GDM versus control cohorts consistently showed decreased PPARA DNAm. Even though cohort size in these studies ranged from rather small sample size of 40 placentas to up to 233 samples, the results demonstrate a reproducible and consistent effect of hyperglycaemia in pregnancy on placenta PGC1A and PPARA promotor DNAm.
Two other candidate genes, LEP (leptin) and ADIPOQ (adiponectin), were also targets in several studies. LEP was investigated in placentas from both GDM and obese pregnancies compared to controls. In two studies, LEP DNAm are found to be increased in GDM pregnancies independent of obesity [Citation48], however, other studies did not find differences [Citation49], or even contradictory results of decreased DNAm associated with glucose levels in 2nd trimester [Citation50]. In obese versus lean pregnancies, one study found no differences in whole placenta tissue [Citation51], whereas another study observed increased LEP DNAm when studying the foetal side of the placenta only [Citation52]. Two studies have investigated ADIPOQ DNAm in obese versus lean pregnancies with apparently contradictory results: Nogues et al. found decreased DNAm at the maternal side only, but the average DNAm percentage was less than 5% [Citation52] raising doubts about the presence of ADIPOQ in placenta. This was indeed concluded in another study, which failed to detect any DNAm of ADIPOQ in placenta [Citation51]. A third study conducted in a significantly larger cohort (n = 98) found ADIPOQ DNAm at foetal side negatively associated with 2 hr glucose post OGTT, ADIPOQ DNAm at maternal side negatively associated with HOMA-IR, and that ADIPOQ DNAm at both sides negatively associated with maternal serum levels of adiponectin [Citation53].
The Horvath epigenetic age acceleration model takes advantage of 62 CpGs in blood cells that are known to be highly associated with biological age. Indeed it has been speculated that offspring of hyperglycaemic and obese pregnancies have an older biological age as compared to their nominal age [Citation54]. We observed that one study of dyslipidemia in women due to altered HDL cholesterol concentrations was associated with accelerated placental epigenetic ageing among women with normal pre-pregnancy weight and a female foetus [Citation55]. This suggests an association between dyslipidemia and placental ageing that may vary by maternal obesity status and foetal sex. In addition, another study observed a negative association between placental epigenetic age acceleration and maternal pre-pregnancy BMI in male offspring only [Citation56]. Whether this can be due to the male sex-associated placentas being more premature remains to be further investigated.
Placenta and imprinted genes
Imprinted genes are characterized by monoallelic expression as a result of epigenetic silencing of one allele based on its parent of origin [Citation57]. Different from all other genes, epigenetic marks of imprinted genes escape erasure during the early stages of blastocyst development and their DNAm levels are stable throughout pregnancy [Citation58]. Based on offspring phenotypes in human imprinting disorders such as Beckwith-Wiedemann or Russel-Silver syndrome, paternally expressed genes are considered to favour foetal growth, whereas maternally expressed genes restrict foetal growth [Citation59,Citation60].
To date close to 100 imprinted genes have been identified in humans [Citation61]. The specific number expressed in human placenta in a strictly monoallelic fashion is unknown, but lower than originally thought [Citation62] and maybe in the range of about 50 to 70. Also the C19MC gene cluster of 52 miRNAs is imprinted in human placenta exclusively expressed from the paternal allele [Citation63]. These imprinted genes and gene clusters play key roles in placental development and function [Citation64].
Pregnancies in women with diabetes or elevated BMI are often associated with altered placental and foetal phenotypes compared to pregnancies in healthy women. Hence, one could predict DNAm changes in imprinted genes with these conditions as a result of both maternal and foetal metabolic changes. Studies have so far limited themselves to maternal exposures and have focused on GDM. They have included only 10 genes imprinted in placenta (). Whereas the majority of maternally expressed genes were unaltered except for DLG associated protein 2 (DLGAP2) with increased DNAm levels in GDM, the three other imprinted genes affected by GDM were paternally expressed [Citation65] (). In addition, reduced MEST DNAm was strongly associated with GDM [Citation66] (). MEST is thought to be involved in angiogenesis regulation [Citation67]. Hence, its lower DNAm in GDM may contribute to placental hypervascularization in some pregnancies in women with GDM.
Table 4. Differentially methylated imprinted genes.
The placenta is not only under the influence of maternal and foetal exposures, but itself can also modulate maternal and foetal metabolic, endocrine and inflammatory conditions thereby establishing a feedforward/feedback loop between mother/placenta and foetus/placenta [Citation68]. Thus, in general DNAm of placental genes may have the potential to also influence maternal conditions. Interestingly, among 188 CpGs, whose DNAm levels associated with maternal insulin sensitivity in an EWAS, were 14 CpGs at 12 imprinted genes, nine maternally and three paternally expressed, respectively [Citation65]. Mendelian randomization found five of these negatively associated with maternal insulin sensitivity among which was DLGAP2 [Citation65] (). Therefore, higher placental DNAm of DLGAP2 contributes to insulin resistance in the pregnant woman, which may explain DLGAP2 increased DNAm in placentas of women with GDM [Citation69].
Cellular composition and methods for placental DNAm data analysis
Placental phenotypes in adverse metabolic conditions are often accompanied by changes in cellular composition of the placenta. Hypervascularization is a common adaptive response to feto-placental transient or chronic hypoxia often found in pregnancies complicated by maternal diabetes or obesity [Citation6,Citation70,Citation71]. Thus, cellular heterogeneity of the placenta may give rise to differential epigenetic patterns in the tissue sample obtained [Citation72]. Variation of cellular composition, but also of position dependent environmental effects on the tissue within placental tissue are major confounders making selection of representative samples important [Citation31]. Position effects of samples have been clearly shown in the imprinted IGF2/H19 region, with increasing methylation the further away the placenta sample was obtained from cord insertion [Citation33]. Cell heterogeneity of the placenta may also be gene-specific, as previously documented for the repetitive LINE-1 region, where DNAm were similar across sampling sites [Citation73]. At present, there is still no consensus on how placenta samples preferably should be obtained, Therefore, the study design of sample positioning, regarding both foetal versus maternal side, central versus posterior location, as well as single versus pooled multiple samples from each placenta, is important to report. This has already been emphasized [Citation35,Citation70,Citation71,Citation74], but positions of sampling sites have not been documented in most studies, which is a considerable limitation of studies using total placental tissue. Bioinformatic methods have been developed to account for potential alterations of cellular composition using deconvolution/cell type specific methods [Citation31]. Reference-based algorithms have been developed to correct for cellular heterogeneity and have also been applied to human placental tissue [Citation75]. Further, a recent study of purified placental cell types allows estimation of cell composition from whole placenta EWAS data [Citation76].
Diabetes or obesity-associated changes in placental cellular composition certainly vary between individual pregnancies adding to confounding. Thus, deconvolution of data that is appropriate in a normal pregnancy may not be suitable for situations with more complex changes in cell composition and cellular phenotype. It remains to be demonstrated whether above or any future methods based on bioinformatics can fully capture the complexity of these changes and correct for them properly.
Perspectives
The focus of studies has so far been on the end of gestation, likely because of easy tissue availability and the association of DNAm with placental health [Citation35].
The early pregnancy period, in particular the first trimester, is understudied. At the molecular and cellular level the placenta responds to maternal diabetes and obesity already at this early stage in pregnancy [Citation77,Citation78]. One can predict changes in DNAm associated with these conditions and, hence, there is an urgent need for these studies. However, early pregnancy placenta samples are difficult to avail. Usually, they are obtained from spontaneous or planned pregnancy terminations, which, for obvious ethical, and in some countries also legal reasons, are very restricted. Even when sampling is possible, general tissue availability is limited, pregnancies are often clinically and metabolically poorly characterized, and pregnancy outcome is unknown. Placental biopsies are normally only obtained by chorionic villus sampling on medical indications (e.g., suspicion of chromosomal/genetic abnormalities) and the amount of tissue is very limited. Whenever feasible, such studies will help to understand how placental trajectories are established that ultimately contribute to foetal development and neonatal outcome [Citation79–81]. Notably, the IGF2/IGF2R axis including H19 is an important target to study, because their transcript levels associated not only with crown-rump length of the foetus in the first trimester, but these associations also track throughout pregnancy to include birth weight [Citation57].
Causal effects of placental DNAm on maternal or foetal phenotype have been hypothesized, but only tested in one study employing Mendelian randomization [Citation65]. This method of genetic epidemiology based on genetic variation needs to be used more widely in order to avoid over-interpretation of statistical exposure-phenotype associations [Citation10]. Associations cannot establish causality and also do not allow for determining directionality. This is particularly important, because of potential bidirectional and distinct effects at the maternal-placental and foetal-placental interface. Quantifying the degree of DNAm of placental genes in the total cell free DNA pool in the maternal circulation may hold promise for being developed into a suitable early biomarker of GDM, perhaps combined with other anamnestic or laboratory parameters predictive of GDM [Citation82].
DNA can not only be methylated to 5-methylcytosine within CpG dinucleotides, but also 5-hydroxymethylated to form 5-hydroxy-methylcytosine. Hydroxymethylation has its own epigenetic function and, in collaboration with 5-methylcytosine, regulates gene transcription in the human placenta [Citation83,Citation84]. Placental hydroxymethylation levels are higher than in most somatic tissues [Citation85] and allelic placental hydroxymethylation is enriched in imprinted domains [Citation84]. Nothing is known about potential placental gene modifications by hydroxymethylation in diabetes and obesity, despite their enrichment in genes involved in regulation of metabolic processes in the placenta [Citation84].
Although the influence of foetal sex on placental responses during pregnancy as well as pregnancy outcome and disease risk later in life is highly suggested, only few studies performed DNAm analysis stratified by foetal sex. Such analyses would provide insights into in utero events driven by foetal sex and potentially shed light on different disease risk development between male and female adults. Besides molecular causes such as X-and Y-chromosome regulated processes, sexual dimorphism might arise due to maternal, placental and/or foetal hormonal differences during pregnancy, which should be taken into account. Specifically, early placental choriogonadotropin (hCG), maternal leptin, oestrogen and progesterone have been associated with risk for GDM and differ between pregnancies of male vs female foetuses [Citation86].
Conclusion
With this review, we have summarized current knowledge on variation of placental DNAm profiles in pregnancies affected by diabetes, obesity, hyperglycaemia and hyperlipidaemia. We observe interesting overlaps in DNAm variation between several studies including a consistent higher DNAm degree at the PGC1A promoter, and lower DNAm degree at the PPARA gene region. Also, the DNAm of the imprinted gene DLGAP2 was found increased both with GDM, and when examining the association by continuous glucose measurements. In addition, available evidence suggests that GDM is associated with higher LEP DNAm, independent of obesity, reinforcing the complexity of GDM effects including different mechanisms linked to hyperglycaemia versus maternal obesity. To the best of our knowledge the effect of maternal T1DM on placental DNAm has not been investigated so far despite established alterations in placental phenotype in T1DM [Citation87]. We furthermore identified missing, yet highly relevant, research of specific placental cell types and in samples obtained at earlier time points than at delivery. Maternal and foetal outcomes directly linked to placental DNAm variation need to be established with consideration of foetal sex. For future approaches there is great potential in conducting Mendelian randomization studies in large sample sizes, to identify causal pathways linking maternal metabolic health during pregnancy with placental DNAm and short- as well as long-term offspring outcome. Introduction of uniformed statistical protocols for DNAm analysis i.e., removal or inclusion of X-and Y-linked probes and adjustment for foetal sex and other known confounders is also a point for improvement as it would enable better comparisons of results between the studies and potentially increase reproducibility. The few studies reporting DNAm variations stratified by foetal sex indeed show sex-specific alterations although we acknowledge the small sample size which is a frequent limitation in these studies. Finally, importance of placental sampling positioning and the overlap between maternal versus foetal DNAm patterns across placenta, cord blood and maternal blood remains unclear, and should be prioritized in future research.
Author Contributions
LH and GD developed the ideas presented in this review, with contributions from BN, SC, RS and PD. LH, BN, SC and GD wrote the manuscript, with contributions from RS and PD. All authors critically revised the manuscript and had access to the final version.
Disclosure statement
No potential conflict of interest was reported by the author(s).
Data Availability Statement
Data is contained within the article and can be accessed from the corresponding authors on reasonable request.
Additional information
Funding
References
- Ben-Haroush A, Yogev Y, Hod M. Epidemiology of gestational diabetes mellitus and its association with type 2 diabetes. Diabet Med. [Internet]. 2004;21(2):103–2208. Available from: http://www.ncbi.nlm.nih.gov/pubmed/14984444
- Damm P, Kühl C, Bertelsen A, et al. Predictive factors for the development of diabetes in women with previous gestational diabetes mellitus. Am J Obstet Gynecol. [Internet]. 1992;167(3):607–616. Available from: http://www.ncbi.nlm.nih.gov/pubmed/1530012
- Wang H, Li N, Chivese T, et al. IDF diabetes atlas: estimation of global and regional gestational diabetes. Diabetes Res Clin Pract. [Internet] 2022;183:109050. Available fromhttp://www.ncbi.nlm.nih.gov/pubmed/34883186
- Chivese T, Hoegfeldt CA, Werfalli M, et al. IDF Diabetes Atlas: the prevalence of pre-existing diabetes in pregnancy - A systematic reviewand meta-analysis of studies published during 2010-2020. Diabetes Res Clin Pract. [Internet]. 2022;183:109049. http://www.ncbi.nlm.nih.gov/pubmed/34883190
- Poston L, Caleyachetty R, Cnattingius S, et al. Preconceptional and maternal obesity: epidemiology and health consequences. Lancet Diabetes Endocrinol. [Internet]. 2016;4(12):1025–1036. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27743975
- McIntyre HD, Catalano P, Zhang C, et al. Gestational diabetes mellitus. Nat Rev Dis Prim. [Internet]. 2019;5:47. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31296866
- Fernandez-Twinn DS, Hjort L, Novakovic B, et al. Intrauterine programming of obesity and type 2 diabetes. Diabetologia. [Internet]. 2019;62(10):1789–1801. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31451874
- Damm P, Houshmand-Oeregaard A, Kelstrup L, et al. Gestational diabetes mellitus and long-term consequences for mother and offspring: a view from Denmark. Diabetologia. [Internet]. 2016;59(7):1396–1399. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27174368
- Retnakaran R, Shah BR. Fetal sex and the natural history of maternal risk of diabetes during and after pregnancy. J Clin Endocrinol Metab. [Internet]. 2015;100(7):2574–2580. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25993641
- Richmond RC, Timpson NJ, Felix JF, et al. Using genetic variation to explore the causal effect of maternal pregnancy adiposity on future offspring adiposity: a mendelian randomisation study. PLoS Med. [Internet]. 2017;14(1):e1002221. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28118352
- Hjort L, Novakovic B, Grunnet LG, et al. Diabetes in pregnancy and epigenetic mechanisms-how the first 9 months from conception might affect the child’s epigenome and later risk of disease. lancet Diabetes Endocrinol [Internet] 2019; Available from: http://www.ncbi.nlm.nih.gov/pubmed/31128973
- Gauster M, Desoye G, Tötsch M, et al. The placenta and gestational diabetes mellitus. Curr Diab Rep. [Internet]. 2012;12(1):16–23. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22102097
- Ringholm L, Mathiesen ER, Kelstrup L, et al. Managing type 1 diabetes mellitus in pregnancy--from planning to breastfeeding. Nat Rev Endocrinol. [Internet]. 2012;8(11):659–667. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22965164
- Metzger BE. Long-term outcomes in mothers diagnosed with gestational diabetes mellitus and their offspring. Clin Obs Gynecol. [Internet]. 2007;50(4):972–979. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17982340
- Verburg PE, Tucker G, Scheil W, et al. Sexual dimorphism in adverse pregnancy outcomes - a retrospective Australian population study 1981-2011. PLoS One. [Internet]. 2016;11:e0158807. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27398996
- Retnakaran R, Shah BR. Sex of the baby and future maternal risk of Type 2 diabetes in women who had gestational diabetes. Diabet Med. [Internet]. 2016;33(7):956–960. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26470996
- Bird A. Perceptions of epigenetics.Nat. 2007;447(7143):396–398.
- Hyun K, Jeon J, Park K, et al. Writing, erasing and reading histone lysine methylations. Exp Mol Med. [Internet]. 2017;49(4):e324. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28450737
- Lander ES, Linton LM, Birren B, et al. Initial sequencing and analysis of the human genome. Nature. [Internet]. 2001;409(6822):860–921. Available from: http://www.ncbi.nlm.nih.gov/pubmed/11237011
- Bird AP. CpG-rich Islands and the function of DNA methylation. Nature. [Internet]. 1986;321(6067):209–213. Available from: http://www.ncbi.nlm.nih.gov/pubmed/2423876
- Pidsley R, Wong CC Y, Volta M, et al. A data-driven approach to preprocessing Illumina 450K methylation array data. BMC Genomics. [Internet]. 2013;14(1):293. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23631413
- Novakovic B, Wong NC, Sibson M, et al. DNA methylation-mediated down-regulation of DNA methyltransferase-1 (DNMT1) is coincident with, but not essential for, global hypomethylation in human placenta. J Biol Chem. [Internet]. 2010;285(13):9583–9593. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20071334
- Lister R, Pelizzola M, Dowen RH, et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature. [Internet]. 2009;462(7271):315–322. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19829295
- Masser DR, Berg AS, Freeman WM. Focused, high accuracy 5-methylcytosine quantitation with base resolution by benchtop next-generation sequencing. Epigenetics Chromatin. [Internet]. 2013;6(1):33. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24279302
- Clark SJ, Harrison J, Paul CL, et al. High sensitivity mapping of methylated cytosines. Nucleic Acids Res. [Internet]. 1994;22(15):2990–2997. Available from: http://www.ncbi.nlm.nih.gov/pubmed/8065911
- Smith ZD, Shi J, Gu H, et al. Epigenetic restriction of extraembryonic lineages mirrors the somatic transition to cancer. Nature. [Internet]. 2017;549(7673):543–547. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28959968
- Ehrlich M, Gama-Sosa MA, Huang LH, et al. Amount and distribution of 5-methylcytosine in human DNA from different types of tissues of cells. Nucleic Acids Res. [Internet]. 1982;10(8):2709–2721. Available from: http://www.ncbi.nlm.nih.gov/pubmed/7079182
- Schroeder DI, Blair JD, Lott P, et al. The human placenta methylome. Proc Natl Acad Sci U S A. [Internet]. 2013;110(15):6037–6042. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23530188
- Chiu RWK, Chim SSC, Wong IHN, et al. Hypermethylation of RASSF1A in human and rhesus placentas. Am J Pathol. [Internet]. 2007;170(7271):941–950. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17322379
- Nelissen ECM, van Montfoort APA, Dumoulin JCM, et al. Epigenetics and the placenta. Hum Reprod Update. [Internet]. 2009;17(7271):397–417. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20959349
- Januar V, Desoye G, Novakovic B, et al. Epigenetic regulation of human placental function and pregnancy outcome: considerations for causal inference. Am J Obstet Gynecol. 2015;213(4):S182–96.
- Bhattacharya A, Freedman AN, Avula V, et al. Placental genomics mediates genetic associations with complex health traits and disease. Nat Commun. [Internet]. 2022;13(1):706. Available from: http://www.ncbi.nlm.nih.gov/pubmed/35121757
- Loke YJ, Galati JC, Morley R, et al. Association of maternal and nutrient supply line factors with DNA methylation at the imprinted IGF2/H19 locus in multiple tissues of newborn twins. Epigenetics. 2013;8:1069–1079.
- Paquette AG, Houseman EA, Green BB, et al. Regions of variable DNA methylation in human placenta associated with newborn neurobehavior. Epigenetics. [Internet]. 2016;11(8):603–613. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27366929
- Wilson SL, Robinson WP. Utility of DNA methylation to assess placental health. Placenta. [Internet]. 2018;64(Suppl 1):S23–8. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29273273
- Khankin EV, Royle C, Karumanchi SA. Placental vasculature in health and disease. Semin Thromb Hemost. [Internet]. 2010;36(3):309–320. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20490981
- Lyon MF. Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature. [Internet]. 1961;190(4773):372–373. Available from: http://www.ncbi.nlm.nih.gov/pubmed/13764598
- Duncan CG, Grimm SA, Morgan DL, et al. Dosage compensation and DNA methylation landscape of the X chromosome in mouse liver. Sci Rep [Internet]. 2018;8:10138. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29973619.
- Alexander J, Teague AM, Chen J, et al. Offspring sex impacts DNA methylation and gene expression in placentae from women with diabetes during pregnancy. PLoS One. [Internet]. 2018;13(2):e0190698. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29470513
- Liu L, Zhang X, Rong C, et al. Distinct DNA methylomes of human placentas between pre-eclampsia and gestational diabetes mellitus. Cell Physiol Biochem. [Internet]. 2014;34: 1877–1889. Available from http://www.ncbi.nlm.nih.gov/pubmed/25503509
- Patti ME, Butte AJ, Crunkhorn S, et al. Coordinated reduction of genes of oxidative metabolism in humans with insulin resistance and diabetes: potential role of PGC1 and NRF1. Proc Natl Acad Sci U S A. [Internet]. 2003;100(14):8466–8471. Available from: http://www.ncbi.nlm.nih.gov/pubmed/12832613
- Barres R, Osler ME, Yan J, et al. Non-CpG methylation of the PGC-1alpha promoter through DNMT3B controls mitochondrial density. Cell Metab. [Internet]. 2009;10(3):189–198. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19723495
- Barrès R, Yan J, Egan B, et al. Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab. 2012;15(3):405–411.
- Wang L, Fan H, Zhou L, et al. Altered expression of PGC-1 α and PDX1 and their methylation status are associated with fetal glucose metabolism in gestational diabetes mellitus. Biochem Biophys Res Commun. [Internet]. 2018;501(1):300–306. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29730292
- Côté S, Gagné-Ouellet V, Guay S-P, et al. PPARGC1α gene DNA methylation variations in human placenta mediate the link between maternal hyperglycemia and leptin levels in newborns. Clin Epigenetics. [Internet]. 2016;8(1):72. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27340502
- Jiang S, Teague AM, Tryggestad JB, et al. Role of metformin in epigenetic regulation of placental mitochondrial biogenesis in maternal diabetes. Sci Rep. [Internet]. 2020;10(1):8314. Available from: http://www.ncbi.nlm.nih.gov/pubmed/32433500
- Xie X, Gao H, Zeng W, et al. Placental DNA methylation of peroxisome-proliferator-activated receptor-γ co-activator-1α promoter is associated with maternal gestational glucose level. Clin Sci (Lond). [Internet]. 2015;129(4):385–394. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25875376
- Lesseur C, Armstrong DA, Paquette AG, et al. Maternal obesity and gestational diabetes are associated with placental leptin DNA methylation. Am J Obstet Gynecol. [Internet]. 2014;211(6):654.e1–9. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24954653
- El HN, Pliushch G, Schneider E, et al. Metabolic programming of MEST DNA methylation by intrauterine exposure to gestational diabetes mellitus. Diabetes. [Internet]. 2013;62: 1320–1328. Available from http://www.ncbi.nlm.nih.gov/pubmed/23209187
- Gagné-Ouellet V, Breton E, Thibeault K, et al. Mediation analysis supports a causal relationship between maternal hyperglycemia and placental DNA methylation variations at the leptin gene locus and cord blood leptin levels. Int J Mol Sci. [Internet]. 2020;21(1):329. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31947745
- Haghiac M, Basu S, Presley L, et al. Patterns of adiponectin expression in term pregnancy: impact of obesity. J Clin Endocrinol Metab. [Internet]. 2014;99(9):3427–3434. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24796925
- Nogues P, Dos Santos E, Jammes H, et al. Maternal obesity influences expression and DNA methylation of the adiponectin and leptin systems in human third-trimester placenta. Clin Epigenetics. [Internet]. 2019;11(1):20. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30732639
- Bouchard L, Hivert MF, Guay SP, et al. Placental adiponectin gene DNA methylation levels are associated with mothers’ blood glucose concentration. Diabetes. [Internet]. 2012;61(5):1272–1280. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22396200
- Hjort L, Vryer R, Grunnet LG, et al. Telomere length is reduced in 9- to 16-year-old girls exposed to gestational diabetes in utero. Diabetologia. 2018;61(4):870–880.
- Shrestha D, Workalemahu T, Tekola-Ayele F. Maternal dyslipidemia during early pregnancy and epigenetic ageing of the placenta. Epigenetics. [Internet]. 2019;14(10):1030–1039. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31179827
- Workalemahu T, Shrestha D, Tajuddin SM, et al. Maternal cardiometabolic factors and genetic ancestry influence epigenetic aging of the placenta. J Dev Orig Health Dis. [Internet]. 2021;12(1):34–41. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31948495
- Moore GE, Ishida M, Demetriou C, et al. The role and interaction of imprinted genes in human fetal growth. Philos Trans R Soc Lond B Biol Sci. [Internet]. 2015;370(1663):20140074. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25602077
- Fowden AL, Coan PM, Angiolini E, et al. Imprinted genes and the epigenetic regulation of placental phenotype. Prog Biophys Mol Biol. [Internet]. 2011;106(1):281–288. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21108957
- Bressan FF, De Bem THC, Perecin F, et al. Unearthing the roles of imprinted genes in the placenta. Placenta. [Internet]. 2009;30(10):823–834. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19679348
- Coan PM, Burton GJ, Ferguson-Smith AC. Imprinted genes in the placenta--a review. Placenta. [Internet]. 2005;26: A:S10–20. Available from http://www.ncbi.nlm.nih.gov/pubmed/15837057
- Allach El Khattabi L, Backer S, Pinard A, et al. A genome-wide search for new imprinted genes in the human placenta identifies DSCAM as the first imprinted gene on chromosome 21. Eur J Hum Genet. [Internet]. 2019;27(1):49–60. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30206355
- Pilvar D, Reiman M, Pilvar A, et al. Parent-of-origin-specific allelic expression in the human placenta is limited to established imprinted loci and it is stably maintained across pregnancy. Clin Epigenetics. [Internet]. 2019;11(1):94. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31242935
- Noguer-Dance M, Abu-Amero S, Al-Khtib M, et al. The primate-specific microRNA gene cluster (C19MC) is imprinted in the placenta. Hum Mol Genet. [Internet]. 2010;19(18):3566–3582. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20610438
- Wagschal A, Feil R. Genomic imprinting in the placenta. Cytogenet Genome Res. [Internet]. 2006;113(1–4):90–98. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16575167
- Hivert M-F, Cardenas A, Allard C, et al. Interplay of placental DNA methylation and maternal insulin sensitivity in pregnancy. Diabetes. [Internet]. 2020;69: 484–492. Available from http://www.ncbi.nlm.nih.gov/pubmed/31882564
- El Hajj N, Pliushch G, Schneider E, et al. Metabolic programming of MEST DNA methylation by intrauterine exposure to gestational diabetes mellitus. Diabetes. 2013;62(4):1320–1328.
- Mayer W, Hemberger M, Frank HG, et al. Expression of the imprinted genes MEST/Mest in human and murine placenta suggests a role in angiogenesis. Dev Dyn. [Internet]. 2000;217(1):1–10. Available from: http://www.ncbi.nlm.nih.gov/pubmed/10679925
- Hiden U, Maier A, Bilban M, et al. Insulin control of placental gene expression shifts from mother to foetus over the course of pregnancy. Diabetologia. [Internet]. 2006;49(1):123–131. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16344925
- Houde -A-A, Ruchat S-M, Allard C, et al. LRP1B, BRD2 and CACNA1D: new candidate genes in fetal metabolic programming of newborns exposed to maternal hyperglycemia. Epigenomics. [Internet]. 2015;7(7):1111–1122. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26586120
- Desoye G, Wells JCK. Pregnancies in diabetes and obesity: the capacity-load model of placental adaptation. Diabetes. [Internet]. 2021;70(4):823–830. Available from: http://www.ncbi.nlm.nih.gov/pubmed/33741605
- Desoye G. The human placenta in diabetes and obesity: friend or foe? the 2017 norbert freinkel award lecture. Diabetes Care. [Internet]. 2018;41(7):1362–1369. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29934479
- Konwar C, Del Gobbo G, Yuan V, et al. Considerations when processing and interpreting genomics data of the placenta. Placenta. [Internet]. 2019;84:57–62. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30642669
- Non AL, Binder AM, Barault L, et al. DNA methylation of stress-related genes and LINE-1 repetitive elements across the healthy human placenta. Placenta. [Internet]. 2012;33(3):183–187. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22222044
- Hogg K, Price EM, Robinson WP. Improved reporting of DNA methylation data derived from studies of the human placenta. Epigenetics. [Internet]. 2014;9(3):333–337. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24394602
- Houseman EA, Kile ML, Christiani DC, et al. Reference-free deconvolution of DNA methylation data and mediation by cell composition effects. BMC Bioinformatics. [Internet]. 2016;17(1):259. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27358049
- Yuan V, Hui D, Yin Y, et al. Cell-specific characterization of the placental methylome. BMC Genomics. [Internet]. 2021;22(1):6. Available from: http://www.ncbi.nlm.nih.gov/pubmed/33407091
- Majali-Martinez A, Weiss-Fuchs U, Miedl H, et al. Type 1 diabetes mellitus and the first trimester placenta: hyperglycemia-induced effects on trophoblast proliferation, cell cycle regulators, and invasion. Int J Mol Sci. [Internet]. 2021;22(20):10989. Available from: http://www.ncbi.nlm.nih.gov/pubmed/34681648
- Lassance L, Haghiac M, Leahy P, et al. Identification of early transcriptome signatures in placenta exposed to insulin and obesity. Am J Obstet Gynecol. [Internet]. 2015;212(5):647.e1–11. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25731694
- Novakovic B, Gordon L, Robinson WP, et al. Glucose as a fetal nutrient: dynamic regulation of several glucose transporter genes by DNA methylation in the human placenta across gestation. J Nutr Biochem. [Internet]. 2013;24(1):282–288. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22901689
- Novakovic B, Yuen RK, Gordon L, et al. Evidence for widespread changes in promoter methylation profile in human placenta in response to increasing gestational age and environmental/stochastic factors. BMC Genomics. [Internet]. 2011;12: 529. Available from http://www.ncbi.nlm.nih.gov/pubmed/22032438
- Desoye G, Cervar-Zivkovic M. Diabetes mellitus, obesity, and the placenta. Obstet Gynecol Clin North Am. [Internet]. 2020;47(1):65–79. Available from: http://www.ncbi.nlm.nih.gov/pubmed/32008672
- Del Vecchio G, Li Q, Li W, et al. Cell-free DNA methylation and transcriptomic signature prediction of pregnancies with adverse outcomes. Epigenetics. [Internet]. 2021;16(6):642–661. Available from: http://www.ncbi.nlm.nih.gov/pubmed/33045922
- Green BB, Houseman EA, Johnson KC, et al. Hydroxymethylation is uniquely distributed within term placenta, and is associated with gene expression. FASEB J. [Internet]. 2016;30: 2874–2884. Available from http://www.ncbi.nlm.nih.gov/pubmed/27118675
- Hernandez Mora JR, Sanchez-Delgado M, Petazzi P, et al. Profiling of oxBS-450K 5-hydroxymethylcytosine in human placenta and brain reveals enrichment at imprinted loci. Epigenetics. [Internet]. 2018;13(2):182–191. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28678681
- Nestor CE, Ottaviano R, Reddington J, et al. Tissue type is a major modifier of the 5-hydroxymethylcytosine content of human genes. Genome Res. [Internet]. 2012;22(3):467–477. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22106369
- Stern C, Schwarz S, Moser G, et al. Placental endocrine activity: adaptation and disruption of maternal glucose metabolism in pregnancy and the influence of fetal sex. Int J Mol Sci. [Internet]. 2021;22(23):12722. Available from: http://www.ncbi.nlm.nih.gov/pubmed/34884524
- Higgins M, Felle P, Mooney EE, et al. Stereology of the placenta in type 1 and type 2 diabetes. Placenta. [Internet]. 2011;32(8):564–569. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21621839
- Lu S, Wang J, Kakongoma N, et al. DNA methylation and expression profiles of placenta and umbilical cord blood reveal the characteristics of gestational diabetes mellitus patients and offspring. Clin Epigenetics. [Internet]. 2022;14(1):69. Available from: http://www.ncbi.nlm.nih.gov/pubmed/35606885
- Horvatiček M, Perić M, Bečeheli I, et al. Maternal metabolic state and fetal sex and genotype modulate methylation of the serotonin receptor type 2A gene (HTR2A) in the Human Placenta. Biomedicines. [Internet]. 2022;10(2):467. Available from: http://www.ncbi.nlm.nih.gov/pubmed/35203678
- Sletner L, Moen AEF, Yajnik CS, et al. Maternal glucose and LDL-cholesterol levels are related to placental leptin gene methylation, and, together with nutritional factors, largely explain a higher methylation level among ethnic South Asians. Front Endocrinol (Lausanne). [Internet]. 2021;12: 809916. Availablefrom http://www.ncbi.nlm.nih.gov/pubmed/35002980
- Franzago M, Fraticelli F, Marchioni M, et al. Fat mass and obesity-associated (FTO) gene epigenetic modifications in gestational diabetes: new insights and possible pathophysiological connections. Acta Diabetol. [Internet]. 2021;58(8):997–1007. Available from: http://www.ncbi.nlm.nih.gov/pubmed/33743080
- Chen C, Jiang Y, Yan T, et al. Placental maternally expressed gene 3 differentially methylated region methylation profile is associated with maternal glucose concentration and newborn birthweight. J Diabetes Investig. [Internet]. 2021;12(6):1074–1082. Available from: http://www.ncbi.nlm.nih.gov/pubmed/33090678
- Awamleh Z, Butcher DT, Hanley A, et al. Exposure to Gestational Diabetes Mellitus (GDM) alters DNA methylation in placenta and fetal cord blood. Diabetes Res Clin Pract. [Internet]. 2021;174:108690. Available from: http://www.ncbi.nlm.nih.gov/pubmed/33549677
- Wang Y, Wang T, Huo Y, et al. Placenta expression of vitamin D and related genes in pregnant women with gestational diabetes mellitus. J Steroid Biochem Mol Biol. [Internet]. 2020;204:105754. Available from: http://www.ncbi.nlm.nih.gov/pubmed/32946925
- Zhao B-H, Jiang Y, Zhu H, et al. Placental delta-like 1 gene DNA methylation levels are related to mothers’ blood glucose concentration. J Diabetes Res. [Internet]. 2019:9521510. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31886292
- Steyn A, Crowther NJ, Norris SA, et al. Epigenetic modification of the pentose phosphate pathway and the IGF-axis in women with gestational diabetes mellitus. Epigenomics. [Internet]. 2019;11(12):1371–1385. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31583916
- Zhang Y, Zhang T, Chen Y. Comprehensive analysis of gene expression profiles and DNA methylome reveals Oas1, Ppie, Polr2g as pathogenic target genes of gestational diabetes mellitus.Sci Rep [Internet]. 2018;8(1):16244.
- Gagné-Ouellet V, Houde -A-A, Guay S-P, et al. Placental lipoprotein lipase DNA methylation alterations are associated with gestational diabetes and body composition at 5 years of age. Epigenetics. [Internet]. 2017;12(8):616–625. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28486003
- Blazevic S, Horvaticek M, Kesic M, et al. Epigenetic adaptation of the placental serotonin transporter gene (SLC6A4) to gestational diabetes mellitus. PLoS One. [Internet]. 2017;12(6):e0179934. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28650965
- Reichetzeder C, Dwi Putra SE, Pfab T, et al. Increased global placental DNA methylation levels are associated with gestational diabetes. Clin Epigenetics. [Internet]. 2016;8(1):82. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27462376
- Rong C, Cui X, Chen J, et al. DNA methylation profiles in placenta and its association with gestational diabetes mellitus. Exp Clin Endocrinol Diabetes. [Internet]. 2015;123(5):282–288. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25962407
- Binder AM, LaRocca J, Lesseur C, et al. Epigenome-wide and transcriptome-wide analyses reveal gestational diabetes is associated with alterations in the human leukocyte antigen complex. Clin Epigenetics. [Internet]. 2015;7(1):79. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26244062
- Finer S, Mathews C, Lowe R, et al. Maternal gestational diabetes is associated with genome-wide DNA methylation variation in placenta and cord blood of exposed offspring. Hum Mol Genet. [Internet]. 2015;24(11):3021–3029. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25634562
- Petropoulos S, Guillemin C, Ergaz Z, et al. Gestational diabetes alters offspring dna methylation profiles in human and rat: identification of key pathways involved in endocrine system disorders insulin signaling, diabetes signaling, and ILK signaling. Endocrinology. [Internet]. 2015;156: 2222–2238. Available from http://www.ncbi.nlm.nih.gov/pubmed/25514087
- Houde AA, St-Pierre J, Hivert MF, et al.,Placental lipoprotein lipase DNA methylation levels are associated with gestational diabetes mellitus and maternal and cord blood lipid profiles.J Dev Orig Heal Dis [Internet] 2014; 5:132–141. Available from.http://www.ncbi.nlm.nih.gov/pubmed/24847699.
- Ruchat SM, Houde AA, Voisin G, et al. Gestational diabetes mellitus epigenetically affects genes predominantly involved in metabolic diseases. Epigenetics. [Internet]. 2013;8(9):935–943. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23975224
- Reese SE, Zhao S, Wu MC, et al. DNA methylation score as a biomarker in newborns for sustained maternal smoking during pregnancy. Environ Health Perspect. [Internet]. 2017;125(4):760–766. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27323799
- Knabl J, Hiden U, Hüttenbrenner R, et al. GDM alters expression of placental estrogen receptor α in a cell type and gender-specific manner. Reprod Sci. [Internet]. 2015;22(12):1488–1495. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25947892
- Cvitic S, Novakovic B, Gordon L, et al. Human fetoplacental arterial and venous endothelial cells are differentially programmed by gestational diabetes mellitus, resulting in cell-specific barrier function changes. Diabetologia. [Internet]. 2018;61:2398–2411. Available from: http://www.ncbi.nlm.nih.gov/pubmed/30091044
- Díaz-Pérez FI, Hiden U, Gauster M, et al. Post-transcriptional down regulation of ICAM-1 in feto-placental endothelium in GDM. Cell Adh Migr. [Internet]. 2016;10(1–2):18–27. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26761204
- Shimanuki Y, Mitomi H, Fukumura Y, et al. Alteration of Delta-like ligand 1 and Notch 1 receptor in various placental disorders with special reference to early onset preeclampsia. Hum Pathol. [Internet]. 2015;46(8):1129–1137. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26014475
- Hoch D, Bachbauer M, Pöchlauer C, et al. Maternal obesity alters placental cell cycle regulators in the first trimester of human pregnancy: new insights for BRCA1. Int J Mol Sci. [Internet]. 2020;21(2):468. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31940810
- Shen W-B, Ni J, Yao R, et al. Maternal obesity increases DNA methylation and decreases RNA methylation in the human placenta. Reprod Toxicol. [Internet]. 2022;107:90–96. Available from: http://www.ncbi.nlm.nih.gov/pubmed/34890771
- Ghildayal N, Fore R, Lutz SM, et al.Early-pregnancy maternal body mass index is associated with common DNA methylation markers in cord blood and placenta: a paired-tissue epigenome-wide association study.Epigenetics. [Internet] 2021;1–11. Available fromhttp://www.ncbi.nlm.nih.gov/pubmed/34384032
- Shrestha D, Ouidir M, Workalemahu T, et al. Placental DNA methylation changes associated with maternal prepregnancy BMI and gestational weight gain. Int J Obes (Lond). [Internet]. 2020;44(6):1406–1416. Available from: http://www.ncbi.nlm.nih.gov/pubmed/32071425
- Prats-Puig A, Xargay-Torrent S, Carreras-Badosa G, et al. Methylation of the C19MC microRNA locus in the placenta: association with maternal and chilhood body size. Int J Obes (Lond). [Internet]. 2020;44(1):13–22. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31554916
- Mitsuya K, Parker AN, Liu L, et al. Alterations in the placental methylome with maternal obesity and evidence for metabolic regulation. PLoS One. [Internet]. 2017;12(10):e0186115. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29045485
- Nomura Y, Lambertini L, Rialdi A, et al. Global methylation in the placenta and umbilical cord blood from pregnancies with maternal gestational diabetes, preeclampsia, and obesity. Reprod Sci. [Internet]. 2014;21(1):131–137. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23765376
- Ouidir M, Zeng X, Workalemahu T, et al. Early pregnancy dyslipidemia is associated with placental DNA methylation at loci relevant for cardiometabolic diseases. Epigenomics. [Internet]. 2020;12(11):921–934. Available from: http://www.ncbi.nlm.nih.gov/pubmed/32677467
- Guay S-P, Houde -A-A, Breton E, et al. DNA methylation at LRP1 gene locus mediates the association between maternal total cholesterol changes in pregnancy and cord blood leptin levels. J Dev Orig Health Dis. [Internet]. 2020;11(4):369–378. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31753053
- Yan W, Zhang Y, Wang L, et al. Maternal dietary glycaemic change during gestation influences insulin-related gene methylation in the placental tissue: a genome-wide methylation analysis. Genes Nutr. [Internet]. 2019;14(1):17. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31086609
- Cardenas A, Gagné-Ouellet V, Allard C, et al. Placental DNA methylation adaptation to maternal glycemic response in pregnancy. Diabetes. [Internet]. 2018;67(8):1673–1683. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29752424
- Desgagné V, Hivert M-F, St-Pierre J, et al. Epigenetic dysregulation of the IGF system in placenta of newborns exposed to maternal impaired glucose tolerance. Epigenomics. [Internet]. 2014;6(2):193–207. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24811788
- Houde -A-A, Guay S-P, Desgagné V, et al. Adaptations of placental and cord blood ABCA1 DNA methylation profile to maternal metabolic status. Epigenetics . [Internet]. 2013;8(12):1289–1302. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24113149
- Wallace C, Smyth DJ, Maisuria-Armer M, et al. The imprinted DLK1-MEG3 gene region on chromosome 14q32.2 alters susceptibility to type 1 diabetes. Nat Genet. [Internet]. 2010;42(1):68–71. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19966805
- Kagami M, Matsuoka K, Nagai T, et al. Paternal uniparental disomy 14 and related disorders: placental gene expression analyses and histological examinations. Epigenetics. [Internet]. 2012;7(10):1142–1150. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22917972
- Bliek J, Alders M, Maas SM, et al. Lessons from BWS twins: complex maternal and paternal hypomethylation and a common source of haematopoietic stem cells. Eur J Hum Genet. [Internet]. 2009;17(12):1625–1634. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19513094
- Gou C, Liu X, Shi X, et al. Placental expressions of CDKN1C and KCNQ1OT1 in monozygotic twins with selective intrauterine growthrestriction. Twin Res Hum Genet. [Internet]. 2017;20(5):389–394. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28803575
- McMinn J, Wei M, Schupf N, et al. Unbalanced placental expression of imprinted genes in human intrauterine growth restriction. Placenta. [Internet]. 2006;27(6–7):540–549. Available from: http://www.ncbi.nlm.nih.gov/pubmed/16125225
- Deyssenroth MA, Marsit CJ, Chen J, et al. In-depth characterization of the placental imprintome reveals novel differentially methylated regions across birth weight categories. Epigenetics. [Internet]. 2020;15(1–2):47–60. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31403346
- Vasconcelos S, Ramalho C, Marques CJ, et al. Altered expression of epigenetic regulators and imprinted genes in human placenta and fetal tissues from second trimester spontaneous pregnancy losses. Epigenetics. [Internet]. 2019;14(12):1234–1244. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31221015
- Dent CL, Isles AR. Brain-expressed imprinted genes and adult behaviour: the example of Nesp and Grb10. Mamm Genome. [Internet]. 2014;25(1–2):87–93. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23974804
- Haggarty P, Hoad G, Horgan GW, et al. DNA methyltransferase candidate polymorphisms, imprinting methylation, and birth outcome.PLoS One. 2013;8(7):e68896.
- Rahat B, Mahajan A, Bagga R, et al. Epigenetic modifications at DMRs of placental genes are subjected to variations in normal gestation, pathological conditions and folate supplementation. Sci Rep. [Internet]. 2017;7(1):40774. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28098215