ABSTRACT
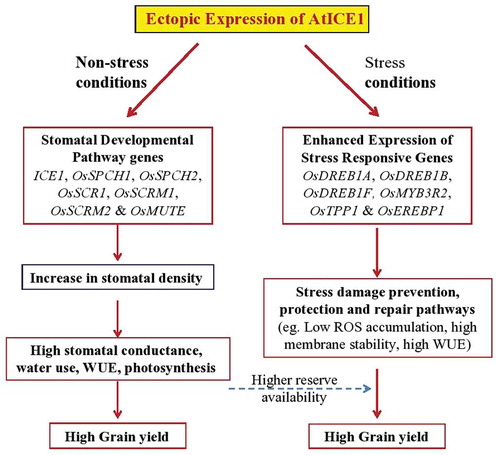
ICE1 (Inducer of CBF Expression 1), a MYC-type bHLH transcription factor, is a regulator of cold tolerance in Arabidopsis. Indica rice, which occupies the major rice cultivated area, is highly sensitive to cold stress. Hence in this study, Arabidopsis ICE1 (AtICE1) was overexpressed in indica rice to analyze its role in reproductive stage cold and other abiotic stress tolerance to indica rice. AtICE1 was overexpressed by using stress inducible AtRD29A promoter in mega rice cv. MTU1010. Under cold stress conditions, AtICE1 overexpression lines showed lower accumulation of MDA and H2O2, higher membrane stability, and thus higher seedling survival rate than the WT plants. Expression levels of OsDREB1A, OsMYB3R2, and OsTPP1 were significantly higher in transgenics as compared with WT under cold stress conditions. AtICE1 transgenic rice plants produced 44–60% higher grain yield as compared with WT plants under control conditions in three independent experiments. Of the three AtICE1 overexpression lines, two lines produced significantly higher grain yield as compared with WT plants after recovery from cold, salt and drought stresses. AtICE1 overexpression lines showed significantly higher stomatal density and conductance under non-stress conditions. qRT-PCR analysis showed that expression levels of stomatal pathway genes viz., OsSPCH1, OsSPCH2, OsSCR1, OsSCRM1, OsSCRM2 and OsMUTE were significantly higher in AtICE1 transgenics as compared with WT plants. The components of water use viz., stomatal conductance, photosynthesis, and instantaneous WUE were higher in transgenics as compared with WT plants. The results showed that AtICE1 confers multiple stress tolerance to indica rice, and the role of ICE1 in stress tolerance and stomatal development is conserved across species.
Introduction
Abiotic stresses such as drought, soil salinity and cold stresses adversely affect plant productivity and quality. Adaptability to the environmental stresses is controlled by complex cascades of molecular networks. In higher plants, the CBF/DREB1 (C-repeat Binding Factor/Dehydration Responsive Element Binding factor 1) transcriptional regulatory cascade is essential for the induction of a set of stress responsive genes for cold acclimation and freezing tolerance.Citation1–3 The DREB1/CBF family of TFs bind to the DRE/CRT cis elements and activate the expression of downstream stress-responsive genes.Citation4,Citation5 Inducer of CBF Expression 1 (ICE1), a bHLH MYC type transcription factor, binds to the MYC recognition cis-elements (CANNTG) in the promoter of CBF3 gene and activates cold induced transcription of CBF3.Citation6–17 The ICE1- CBF pathway positively regulates the expression of several cold responsive genes, and thus regulates cold acclimation and freezing tolerance in Arabidopsis.Citation6,Citation18,Citation19 Posttranslational regulation of ICE1 is essential for CBF expression and cold tolerance. The ICE1 protein is subjected to 26S proteosome mediated degradation under prolonged cold stress by High expression of Osmotically responsive gene 1 (HOS1), an E3 ubiquitin protein ligase.Citation20 Under cold stress, MPK3/6 catalyzed phosphorylation of ICE1 at Ser94, Thr366 and Ser403 promotes the degradation of ICE1.Citation13,Citation14 Further, BIN2 (BRASSINOSTEROID-INSENSITIVE2) mediated phosphorylation at Ser94 of ICE1 has also been shown to promote ICE1 degradation.Citation17 The ICE1 protein stability is positively regulated by sumoylation of ICE1 by SAP and Miz (SIZ1), an E3 SUMO protein ligase,Citation21 and Open Stomata 1 (OST1, SnRK2.6) mediated phosphorylation of S278 of ICE1 under cold stress.Citation10 Both SIZ1 mediated sumoylation and OST1 mediated phosphorylation suppress ICE1 degradation under cold stress and promote cold tolerance. Besides, transcriptional activity of ICE1 is negatively regulated by its interaction with transcription factors MYB15,Citation22 jasmonate ZIM-domain 1/4 (JAZ1 and JAZ4) proteins,Citation9 MYC67, and MYC70.Citation15 In addition to cold tolerance, ICE1 also regulates stomatal development,Citation23 suppression of flowering under cold stress,Citation12 primary seed dormancy and endosperm reserve mobilization,Citation16 male fertility, and ABA signaling during germination of seeds.Citation24 ICE1–CBF cold stress signaling pathway is conserved across diverse plant species.Citation25 From several plant species ICE1 homologs have been cloned and when the ICE1 is overexpressed in the same species from which it was cloned or expressed in other species, it conferred enhanced expression of CBF homologs and cold, freezing and/or multiple abiotic stress tolerance (Table S1).
Rice is highly sensitive to drought, salt and cold stresses.Citation26 In general japonica subspecies, predominantly cultivated in temperate zone is more cold tolerant than the indica subspecies which cultivated in tropics and subtropics of Philippines, India, Pakistan, Java, Sri Lanka, Indonesia, central and southern China, and in some African countries.Citation27,Citation28 Identification and use of genes which can confer multiple stress tolerance is important to enhance rice productivity. Hence we explored the potential of AtICE1 in conferring multiple abiotic stress tolerance in indica rice cultivar. Earlier AtICE1 and its homologs from different plant species have been overexpressed mainly in japonica cultivars and found to enhance cold or drought tolerance (Table S1). AtICE1 overexpression from constitutive 35S promoter conferred seedling stage chilling tolerance in japonica rice variety Kenjiandao Citation29 and rice ICE1 overexpression under constitutive ubiquitin promoter enhance cold tolerance in japonica rice cultivar Nipponbare.Citation30 Similarly, overexpression of ICE1 from Chinese cabbage, radish and woad from constitutive Ubiquitin promoter in japonica rice cultivars have been shown to enhance seedling stage cold tolerance.Citation31–33
In previous studies, overexpression of ICE1 and its homologs have been shown to confer multiple stress tolerance in Arabidopsis,Citation34–36 Chrysanthemum,Citation37 and tobacco.Citation38–40 Thus, previous studies focused on cold or drought tolerance of seedling/vegetative stage of japonica cultivars constitutively overexpressing ICE1 from constitutive promoter. This study was conducted to analyze the potential of AtICE1 expressed from stress inducible AtRD29A promoter in conferring cold, drought and salt stress tolerance to indica rice cultivar.
Materials and methods
Construction of AtICE1 vector and rice transformation
Full-length CDS (1485bp) of Arabidopsis ICE1 (At3g26744) was amplified with gene specific primers using LA Taq DNA Polymerase (DSS Takara, Japan) from the AtICE1 super promoter clone,Citation6 and cloned in modified pCAMBIA1300 under stress inducible promoter AtRD29A (Figure S1). Binary construct was confirmed by colony PCR, restriction digestion with BamHI and KpnI (NEB, USA) followed by sequencing of plasmid clone. Confirmed clone was transformed in Agrobacterium strain EHA105. Rice transformation was performed in indica rice variety cv. MTU1010 via Agrobacterium-mediated cocultivation method.Citation41
Molecular confirmation of transgenic rice plants
DNA isolation was performed by CTAB method using leaf samples from putative transgenic plants (T2).Citation42 PCR confirmation of transgenic plants was done with AtICE1, HPTII, AtRD29A-F, and NOS-R primers using Brazilian Taq DNA Polymerase (Invitrogen, USA). PCR conditions were 94°C for 3 min for denaturation and enzyme activation, followed by 30 cycles of 94°C for 30 s, 56°C for 30 s, 68°C for 1 min 30 s and final extension for 10 min at 72°C on a PCR machine (Biorad, USA). The details of primers used in this study are available in Table S1. Southern blot analysis was performed using 10 µg of DNA from different transgenic overexpression lines and WT plant. DNA samples were restriction digested with KpnI for overnight. Restricted products were separated in 0.8% agarose gel at 40 volt for 5 h and capillary transferred in nylon membrane following standard protocol. HPTII probe was prepared using DIG DNA Labeling and Detection kit (Roche, USA) following manufacturer’s protocol.
qRT-PCR analysis of AtICE1 transgenic rice
AtICE1 overexpressing rice seedlings (25 days) were treated with cold (4°C), 20% PEG6000 (−0.49 MPa) and 200 mM NaCl (−1.01 MPa; 20 dS/m) in Yoshida plant nutrient medium for 8 h. At the end of treatment, leaf tissues were harvested and frozen in liquid nitrogen. Total RNA was isolated using RNA isolation kit (Ambion, Invitrogen, USA). Purified 2 µg of total RNA was used for cDNA preparation using reverse transcriptase enzyme (Superscript III RT, Invitrogen, USA). qRT-PCR reaction was performed by using qRT-PCR kit (Kapa Sybr® Fast, USA). PCR conditions were 95°C for 30 s, followed by 40 cycles of 95°C for 5 s and 60°C for 40 s on Real Time PCR machine (ABI, USA). Expression of AtICE1 (At3g26744), OsDREB1A (AF300970), OsDREB1B (AF300972), OsDREB1F (AY785897), OsMYB3R2 (AK066520), OsTPP1 (AB120515), OsEREBP1 (AF193803) and OsICE1 (AK109915) were analyzed. Ubiquitin5 gene was used as reference.Citation43 Relative fold change in gene expression was calculated by using comparative 2−ΔΔCT method. The list of primers used in this study is available in Table S2.
Abiotic stress tolerance of AtICE1 transgenic rice Cold tolerance at seedling stage
Cold stress tolerance in AtICE1 transgenic (T2) and WT plants were evaluated at seedlings stage. Seedlings were grown in soilrite in small pots at 30°C under 16/8 h light/dark photoperiod in culture room with replicates. 25 days old seedlings were shifted in cold chamber for cold treatment at 15°C for 48 h (acclimation) followed by 6°C for 48 h. At the end of 48 h, membrane stability was measured following the protocol of Blum and Ebercon (1981).Citation44 Total chlorophyll content was measured following the method of Hiscox and Israelstam, (1979).Citation45 MDA is indicator of lipid peroxidation due to oxidative stress and it was measured as method given by Heath and Packer (1968).Citation46 H2O2 was determined following the methods described earlier,Citation47 with three biological replicates. Recovery was done at 32°C for 1 week and survival rate was recorded.
Cold stress tolerance at booting stage
Transgenic lines (T2) were germinated on MS medium supplemented with hygromycin (50 mg/L). AtICE1 overexpressing transgenic rice and WT seedlings were transplanted in pots, with at least three replicates, and grown in transgenic greenhouse at 30 + 2°C and 60–70% RH. At booting stage, one set of AtICE1 and WT plants were shifted to cold chamber for cold acclimation at 15°C for 2 days followed by 8°C for 3 days with 16/8 h light/dark photoperiod in cold chamber. At the end of the stress, membrane stability was measured by electrolyte leakage following the protocol of Blum and Ebercon (1981).Citation44 Chlorophyll fluorescence was monitored in fully expanded leaves under control and cold stress by using fluorescence meter (FluorPen FP110, USA). Recovery was done at 30 ± 2°C in greenhouse and grown till maturity, and yield data were recorded. Harvest index was calculated as follows: HI(%) = (Grain yield/Biomass) x 100.
Salt tolerance
AtICE1 overexpressing transgenic lines (T2) and WT plants grown in the same pot, with at least three replicates, and grown in transgenic greenhouse at 30 ± 2°C and 60–70% RH. At booting stage, one set of plants were subjected to salt stress by irrigating with NaCl (200 mM) solution till EC of soil reached ~>8 dS. Electrical conductivity of soil was measured from control and salt stress pots with three biological repetitions. Soil sample was collected and saturated with ddH2O and the conductivity was measured by portable conductivity meter (Eutech instruments, USA). Plants were subjected to salt stress for 15 days till plants develop the senescence symptoms. At the end of the stress, membrane stability was measured following the protocol of Blum and Ebercon, (1981).Citation44 Total chlorophyll content was measured following the method of Hiscox and Israelstam, (1979).Citation45 Recovery was done by pouring excess amount of water in pots to leach out the salts by drainage. Recovered plants were grown till maturity under irrigated condition, and yield data were recorded.
Water use and drought tolerance at panicle initiation stage
Quantification of water use by AtICE1 transgenics (T3) and WT plants under control and drought stress was done at vegetative (panicle initiation) stage under control and drought stress, with three biological repetitions. Single plant was planted per pot and grown in soil pots. At panicle initiation stage (~70-day-old plants) drought stress was imposed by withholding irrigation for 10 days, while control set of plants were irrigated to maintain soil moisture saturation. Soil surface was covered with polythene film to avoid direct evaporation from soil surface. Fresh weights of pots were recorded daily. The water use under control, the saturated pot weight was recorded initially and daily irrigation was done to maintain the saturation weight. The water use was quantified from the amount of water added everyday in control pots to maintain the soil moisture saturation. The soil moisture depletion in drought imposed pots was quantified by recording the fresh weight of pots throughout the drought period. Soil Metric Potential (SMP) was measured in pots by using tensiometer (Empl, India) and soil moisture content (SMC) was measured by gravimetric method,Citation48 under drought stress. Photosynthesis and stomatal conductance was measured using portable IRGA (LiCOR6400, USA) when the soil matric potential was about −50 kPa in drought stressed pots. Drought stressed plants were recovered by rewatering at the end of stress period and grown till maturity to record yield data.
Stomatal density and gene expression analysis
Stomatal density was measured in WT and overexpressing AtICE1 transgenic lines from 70-day-old control plants. Abaxial surface of flag leaf was imprinted using glue and fixed in glass slides for stomatal density measurement from method.Citation49 Stomatal images were captured in light microscope (EVOS®FL, USA) at 40x resolutions to count number of stomata. Developing leaves before expansion were collected to study differential expression of stomatal genes. Newly emerging leaf was used for RNA extraction and expression analysis of genes for stomatal development. Expression of stomatal developmental genes OsMUTE (Os05g51820), OsFAMA (Os05g50900), OsSPCH1 (Os06g33450), OsSPCH2 (Os02g15760), OsSCRM1 (Os11g32100), OsSCRM2 (Os01g71310) and OsSCR1 (Os11g03110) were analyzed. Ubiquitin5 gene was used as reference. Primers sequences used in this study are given in Table S2.
Statistical analysis
The analysis of variance for test of significance for gene expression data was done by using R studio software and alphabets denotes the variation (P < .05). One factor analysis of variation for physiological and biochemical data was analyzed with statistical software package (OPSTAT) at 5% level of significance.
Results
Development of AtICE1 transgenic rice
Putative transgenic lines obtained from genetic transformation of rice cv. MTU1010 were confirmed by PCR with primers for AtICE1 and HTPII genes. Southern blot analysis in T2 generation confirmed the integration of transgene in transgenic lines Ox-1, Ox-2 and Ox-3 (Figure S2).
Expression analysis of stress responsive genes
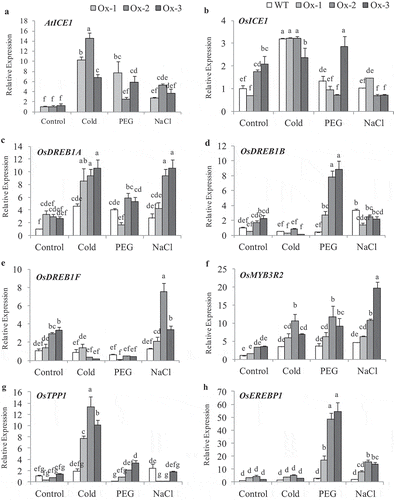
Expression analysis of transgene AtICE1 and stress responsive genes were analyzed in transgenic rice plants (T2) subjected to 20% PEG (−0.49 MPa), 200 mM NaCl (−1.01 MPa; 20 dS m-1) and cold (4°C) treatments. AtICE1 expression was upregulated under osmotic, cold and salt stresses over control in transgenic plants, as AtICE1 was expressed from stress inducible AtRD29A promoter in transgenic plants ()). Native ICE1 (OsICE1) gene was significantly upregulated under cold stress in both WT and AtICE1 transgenics as compared with control, while under osmotic and salt stress only in one of the transgenic line the expression was higher than the WT plants ()). Cold stress upregulated the expression of ICE1 target genes OsDREB1A, OsMYB3R2, and OsTPP1 in AtICE1 overexpression lines as compare to WT ()). Osmotic stress significantly upregulated the expression of OsDREB1B, OsMYB3R2, OsTPP1, and OsEREBP1 in AtICE1 transgenic lines ()). In salt stress, OsDREB1A, OsDREB1F, OsMYB3R2, and OsEREBP1 expression were higher in transgenic lines as compared with WT plants ().
Figure 1. Expression analysis of stress responsive genes in shoot tissues from AtICE1 overexpression transgenic lines. Total RNA was isolated from 25 days old seedlings were treated with PEG6000 (20%), NaCl (200 mM) and Cold (4°C) for 8 h was used for gene expression analysis. (a) AtICE1 expression in transgenic lines. (b) Expression of native OsICE1 in WT and Transgenic plants. (c-h) Expression of ICE1 target genes and stress responsive genes. The series legends are same for B-H. The experiment were performed with three biological replicates (n = 3). Error bar indicates ±SE. Statistical differences are shown on the bars by labeling significantly different groups with different letters (P < .05)
AtICE1 confers multiple stress tolerance to rice
AtICE1 improves cold stress tolerance
AtICE1 overexpressing transgenic rice (T2) seedlings exhibited significantly better tolerance to cold stress as compared to WT. Low temperature stress at 15°C and 6°C for 48 h lead to symptoms such as leaf wilting and dryness which were less in transgenic rice as compared to WT ()). Membrane stability was significantly higher in AtICE1 (69–72%) transgenics as compare to WT (47%) seedlings under cold stress ()). Accumulation of MDA and H2O2 content were significantly lower in AtICE1 transgenic rice lines as compare to WT under cold stress ()). Upon recovery from cold stress AtICE1 transgenic rice lines showed higher survival rate (87–91%) as compared to WT (57%) seedlings ()). These results showed that AtICE1 overexpression confers cold tolerance to indica rice cultivar. At reproductive stage also, transgenic rice lines overexpressing AtICE1 showed higher levels of tolerance to cold stress (8°C) as compared to WT ()). Analysis of the maximum quantum yield of the Photosystem II (PSII) in dark-adapted leaves showed that significantly higher PSII yield in AtICE1 plants as compared to WT plants under cold stress ()). Membrane stability of AtICE1 overexpression lines were significantly higher compared to WT under salt stress ()). After recover, all the three transgenic lines produced significantly higher (58–214%)) grain yield over WT plants ()).
Figure 2. Cold stress tolerance of AtICE1 overexpressing transgenic rice (T2) lines. (a) Photograph of rice seedling before cold stress (control), at the end of cold stress 6°C for 48 h and one week after recovery. (b–d) Graph shows Membrane stability, MDA, and H2O2 content under cold stress. (e) Survival rate of seedlings at the end of recovery period. Graph shown the mean value from three biological replicates (n = 3). Error bar indicates ±SE. *means P < .05
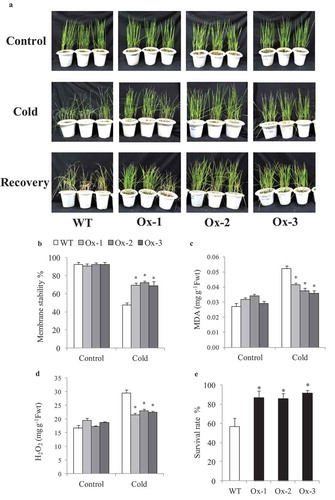
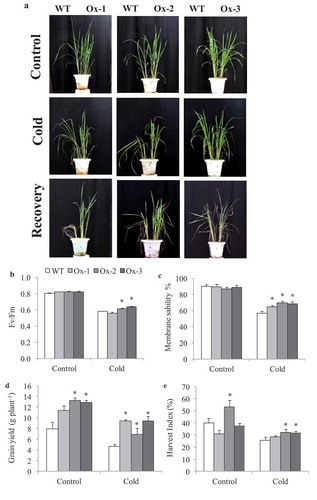
Figure 3. Evaluation for cold stress tolerance in overexpressing AtICE1 overexpression transgenic rice (T3 generation) at reproductive stage. (a) Response under cold stress for 15°C for 48 hrs and 8°C for 72 hrs and recovered at 32°C in greenhouse. (b,c) Quantum yield (Fv/FM), membrane stability were recorded under cold stress. (d,e) Grain yield and harvest index at maturity after cold stress recovery in AtICE1 overexpression lines and WT plants. The experiment was performed with three biological replicates (n = 3). Error bar indicates ±SE. *means P < .05
AtICE1 improves salt tolerance
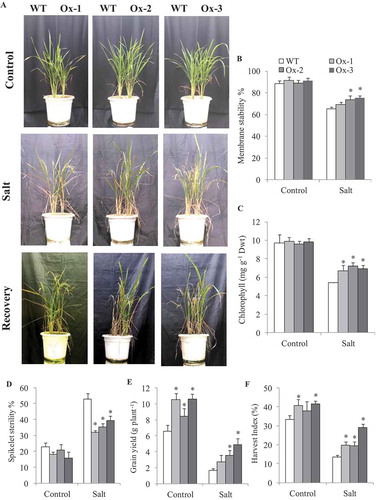
WT and AtICE1 overexpression lines (T2) grown in the same pot were subjected to salt stress (7.7–9.3 dS m−1) at booting stage (), Figure S3a). Membrane stability of AtICE1 overexpression lines Ox-2 and Ox-3 were significantly higher as compared to WT under salt stress ()). Total chlorophyll content was significantly higher in AtICE1 as compared to WT plant ()). Spikelet sterility was significantly lower in transgenic lines as compared to WT under salt ()). Grain yield of all three transgenic lines were significantly higher under control conditions. Under salt stress AtICE1 Ox-2 and Ox-3 lines produced significantly higher grain yield than WT plants ()). However, the differences in biomass, tiller number and panicle number between WT and transgenic lines were nonsignificant except for Ox-3 which produced significantly higher tiller number and panicle number under control conditions (Figure S3(b-d)).
Figure 4. Evaluation of overexpressing AtICE1 transgenic rice (T2) under salt stress at reproductive stage. (a) Photograph showing phenotype of plant before and after stress and recovery in AtICE1 transgenic and WT plants. (b,c) Comparative analysis of membrane stability, chlorophyll content under stress. (d–f) Spikelet sterility, grain yield and harvest index after maturity. The experiment was performed with three biological replicates (n = 3). Error bar indicates ±SE. *means P < .05
AtICE1 improves water use and drought tolerance
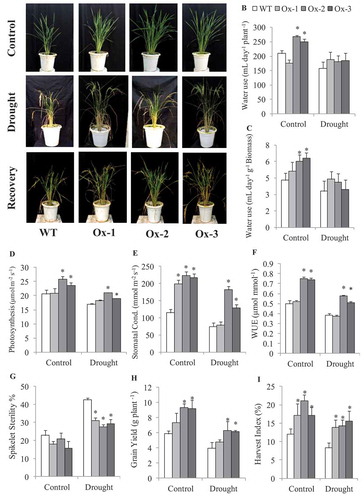
Analysis of water use in AtICE1 overexpressing and WT plants was studied under control and drought at panicle initiation stage ()). AtICE1 Ox-2 and Ox-3 lines utilized about 250 mL per day as compared to WT which used about 210 mL per day. At the end of 10 days drought stress, the soil moisture content was 5.2% in the pots where WT was grown, while in the pots of transgenic lines, it varied from 5.05 to 6.05%, which was statistically on par with WT pot (Figure S4(a)). The mean water use under drought stress was 180.5–189 mL for AtICE1 transgenic lines as compared with 158 mL for WT plants ()). On per unit biomass basis, the whole plant transpiration was 4.3 mL day−1 g−1 biomass in WT, while that of AtICE1 transgenic lines varied from 5.13 to 6.31 mL day−1 g−1 biomass under non-stress conditions. Under drought stress conditions, the mean water use over the stress period was 3.31 mL day−1 g−1 biomass in WT, while that of AtICE1 transgenic lines varied from 3.46 to 4.41 mL day−1 g−1 biomass ()). In consistent with higher whole plant water use, stomatal conductance was significantly higher in AtICE1 transgenics except that of Ox-1 under drought stress as compared with WT plants ()). Rate of photosynthesis and instantaneous WUE were also significantly higher in Ox-2 and Ox-3 lines as compared to WT plants under control and drought stress (~ −50 kPa) conditions ()).
Figure 5. Evaluation of AtICE1 transgenic rice (T3) for Water use at vegetative stage. (a) Photographs were captured at control, drought and recovery after 21 days. (b,c) Water used per day by AtICE1 transgenic lines and WT plants under control and drought stress. (d-f) Rate of photosynthesis and stomatal conductance and WUE under control and drought stress (−50 kPa). (g,h) Grain yield and Harvest index at maturity under control and drought. The experiment was performed with three biological replicates (n = 3). Error bar indicates ±SE. One way Anova was done for test of significance with online software (opstat).*means P < .05
After the end of one cycle of drought stress, the plants were recovered and grown to maturity. Analysis of plant biomass and grain yield was done at maturity to compare AtICE1 overexpressing transgenic and WT plants after one cycle of drought stress. Spikelet sterility was significantly lower in AtICE1 transgenics lines ()). Grain yield was significantly higher in Ox-2 and Ox-3 as compare to WT under control and drought stress ()). The biomass was not significantly different between WT and transgenic plants under non-stress and stress conditions (Figure S4(b)). Tillers per plant were significantly higher in transgenic lines as compared with WT plants except that of Ox-3 under drought stress (Figure S4(c)). Number of panicles per plant was significantly higher in transgenic lines as compared with WT plants under control conditions, while under drought stress, only Ox-1 had significantly higher panicle number (Figure S4(d)).
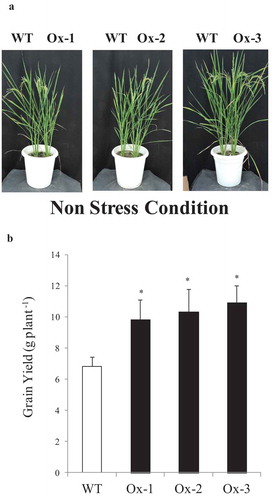
In all three independent experiments described above AtICE1 transgenic lines produced higher grain yield under control conditions. AtICE1 transgenic rice showed better growth and produced relatively higher no. of tillers and panicles compare to WT plant (Figures S3(c,d) and S4(c,d)). Spikelet sterility was found less in AtICE1 transgenic rice compare to WT plants under non stress condition in green house. Significant contribution of these yield traits in AtICE1 transgenic rice helps in better seed setting and produce higher grain yield compare to WT plants. The mean of all three experiment showed that AtICE1 overexpression rice lines produced 44–66% higher grain yield over WT plants ()).
Figure 6. Overexpression of AtICE1 improved yield under nonstress conditions. (a) Response of AtICE1 transgenic and WT plants under non stress condition. (b) Yield data from control plants of three independent experiments (cold, salt, and drought) conducted at different time of the year were pooled analyzed. Each experiment was performed with six biological replicates (n = 18). Error bar indicates ±SE. *means P < .05. *means P < .05
AtICE1 enhances stomatal density
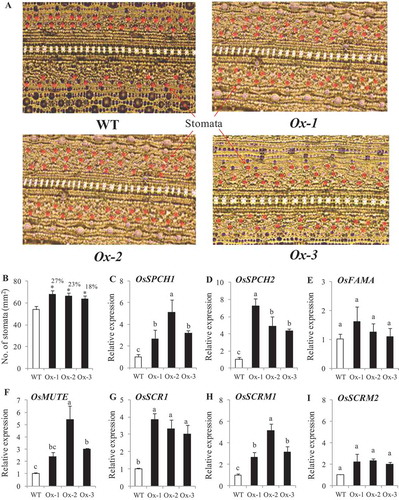
No. of stomata per unit area were higher in transgenics plants as compared with WT plants under nonstress conditions ()). Stomatal density of Ox-1, Ox-2 and Ox-3 lines were 27%, 23%, and 18% higher respectively as compared to WT plants ()). Expression analysis of genes for stomatal development viz, SPCH1, SPCH2, FAMA, MUTE, SCR1, SCRM1, and SCRM2 showed that all the genes except FAMA and SCRM2 were significantly higher in transgenic lines as compared with WT plants ()).
Figure 7. Stomatal density in AtICE1 overexpressing transgenic rice (T2) under control condition. (a) Image of abaxial leaf surface from fully expanded leaves under microscope (40x). (b) Stomatal density under control condition. (c-i) qRT-PCR expression analysis of genes for stomatal development. RNA was extracted from newly emerging leaves under control condition. The experiment was performed with three biological replicates (n = 3). Error bar indicates ±SE. *means P < .05. Column with different letters shows significant differences * P < .05
Discussion
The present study showed that stress inducible overexpression of AtICE1 gene confers cold, salt and drought stress tolerance of indica rice. Cold, osmotic stress, and NaCl treatment has significantly upregulated ICE1 downstream target genes in rice. Previous studies have shown that AtICE1 overexpression confers cold tolerance to Arabidopsis,Citation6 rice,Citation29 and Cucumis sativas,Citation50 drought tolerance to Arabidopsis,Citation34 and tolerance to salt and osmotic stress in tobacco,Citation39 at seedling stage. The ability of AtICE1 to confer cold, salt, and drought tolerance at reproductive stage in crop plants was not examined earlier. However, overexpression of ICE1 homologs from tomato, grape, C. sinensis, Brassica campestris, and Chrysanthemum conferred tolerance cold, salt and drought stresses in tobacco, Arabidopsis or Chrysanthemum,Citation35,Citation37,Citation38,Citation40 at vegetative stage. Thus, our study showed that AtICE1 overexpression from stress inducible promoters imparts multiple stress tolerance to rice.
Analysis of ICE1 downstream target genes revealed that expression levels of some target genes were significantly higher under stress conditions in AtICE1 transgenic lines as compared with WT plants. Expression levels of OsDREB1A, OsDREB1B, OsDREB1F, OsMYB3R2, OsTPP1, and OsEREBP1 genes were higher in AtICE1 overexpression lines than in WT in one or other stress conditions. Previous studies have shown that overexpression of OsDREB1A,Citation51 OsDREB1B,Citation52 OsDREB1F,Citation53 OsMYB3R2,Citation54 OsTPP1,Citation30,Citation55 and OsEREBP1,Citation56 enhanced tolerance of transgenic plants to biotic and abiotic stresses at seedling stage. In recent study OsICE1 transgenic rice induces OsTPP1 expression and increases trehalose level which improved the chilling tolerance in rice.Citation30 In this study AtICE1 transgenic lines produced significantly less H2O2 and MDA, and thus better membrane stability as compared to WT under cold stress. Thus as shown previously, besides the regulation of cold tolerance through CBFs, ICE1 may confer cold tolerance through enhanced antioxidative enzymes and polyamines (Table S1). Among the 34 studies on overexpression of AtICE1 or its homolog (Table S1)Citation6,Citation29-40,Citation50,Citation57-76, only in one study on reproductive stage drought tolerance and yield was examined (Table S1).Citation58 Similar to our results, Chander et al (2018),Citation58 found that OsICE1 overexpression confer higher grain yield over WT plants under drought stress in japonica rice cv. Nipponbare.
AtICE1 overexpression from stress-inducible RD29A promoter significantly increased stomatal density, stomatal conductance, photosynthesis, instantaneous WUE and whole plant water use (), which might have contributed to 44–60% higher grain yield of transgenics over WT plants under control conditions (). Lower spikelet sterility under drought and salt stress significantly improves the grain yield in AtICE1 transgenic rice lines. AtRD29A promoter was used for stress inducible expression of AtICE1 gene. AtRD29A promoter was found to induce transgene expression even under control growth conditions of rice (~32°C), and the expression levels were further enhanced under stress condition.Citation41 In this study also AtICE1 expression was found under non stress condition which was enhanced further under stress conditions in AtICE1 overexpression lines ()). The expression of OsICE1 gene was also significantly higher AtICE1 transgenics Ox-2 and Ox-3 than that of WT plants. This base level expression of AtRD29A driven AtICE1, and OsICE1 in transgenic rice lines might have contributed to enhanced stomatal density, and thus stomatal conductance, water use, photosynthesis and yield under non-stress conditions.
ICE1 play positive role in multiple stages of stomatal development and by forming heterodimers with SPCH, MUTE, and FAMA.Citation23 Interaction between AtICE1 and OsSPCH, OsMUTE, and OsFAMA might play role in stomatal density and behavior. The SCR1 impacts on formation of heterodimers by AtICE1 during establishment of the stomatal cell lineage and increases stomatal density and decreases size. ICE1/SCREAM (SCRM) and SCRM2, which directly interact with SPCH, MUTE, and FAMA and specify the sequential actions of SPCH, MUTE, and FAMA, indicating that SCRM and SCRM2 together determined successive initiation, proliferation and terminal differentiation of stomatal cell lineages.Citation23 In the present study, we analyzed the stomatal density and relative gene expression of AtICE1 transgenic rice compare to WT. Leaf microscopic study confers the higher stomatal density in AtICE1 overexpressing transgenic rice. In the developing leaves, in addition to AtICE1, significantly higher expression of OsSPCH1, OsSPCH2, OsMUTE1, OsSCR1, OsSCRM1, and OsSCRM2 were observed in AtICE1 overexpressing transgenic rice as compared to WT plants (). Thus, enhanced expression of ICE1 and other stomatal developmental pathway genes might have contributed to the enhanced stomatal density in AtICE1 transgenic rice lines.
In conclusion, we showed that overexpression of AtICE1 gene significantly enhanced the cold, drought and salt stress tolerance at reproductive stage. In addition, the multiple stress tolerance, overexpression of AtICE1 gene from AtRD29A promoter, which shows some basal activity in rice even under non-stress conditions, significantly enhanced the grain yield. Often, overexpression of genes for stress tolerance shows some adverse effect on yield under nonstress condition. However, ICE1 is useful the enhance grain yield under nonstress conditions as well as stress conditions in rice. Plant can respond to drought through dehydration avoidance (stomatal closure, water mining, etc.) and cellular tolerance to dehydration. The drought tolerance of ICE1 appears to be through cellular tolerance to dehydration mechanisms, i.e., through enhanced availability of photosynthates, reduced ROS levels and membrane stability. Enhanced yield under nonstress conditions could be attributed to higher stomatal density, stomatal conductance, photosynthesis, and thus grain yield. Enhanced stress tolerance could be attributed to the enhanced expression of stress responsive genes, and thus stress tolerance (). Thus our study showed that AtICE1 overexpression is a potential strategy to improve abiotic stress tolerance and grain yield in indica rice.
Author contributions
RKV did the gene cloning and gene expression analysis, RKV and VVSK did transgenic development. RKV managed the stress experiments and plant material. RKV and SKY did physiological and biochemical analysis. MVR and TSK helped in project preparation and data analysis. VC designed the project and supervised the project. VC and RKV designed the experiment, analysed the data and wrote the manuscript. All authors read and approved the manuscript.
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
Supplemental Material
Download MS Word (25 KB)Supplemental Material
Download MS Power Point (337.7 KB)Supplementary material
Supplemental data for this article can be accessed on the publisher’s website.
Additional information
Funding
References
- Thomashow MF. Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annu Rev Plant Physiol Plant Mol Biol. 1999;50:1–13. doi:https://doi.org/10.1146/annurev.arplant.50.1.571.
- Chinnusamy V, Zhu J, Zhu JK. Cold stress regulation of gene expression in plants. Trends Plant Sci. 2007;12(10):444–451. doi:https://doi.org/10.1016/j.tplants.2007.07.002.
- Zhu JK. Abiotic stress signaling and responses in plants. Cell. 2016;167(2):313–324. doi:https://doi.org/10.1016/j.cell.2016.08.029.
- Stockinger EJ, Gilmour SJ, Thomashow MF. Arabidopsis thaliana CBF1 encodes an AP2 domain-containing transcriptional activator that binds to the C-repeat/DRE, a cis-acting DNA regulatory element that stimulates transcription in response to low temperature and water deficit. Proc Natl Acad Sci USA. 1997;94(3):1035–1040. doi:https://doi.org/10.1073/pnas.94.3.1035.
- Liu Q, Kasuga M, Sakuma Y, Abe H, Miura S, Yamaguchi-Shinozaki K, Shinozaki K. Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell. 1998;10(8):1391–1406. doi:https://doi.org/10.1105/tpc.10.8.1391.
- Chinnusamy V, Ohta M, Kanrar S, Lee BH, Hong X, Agarwal M, Zhu JK. ICE1: a regulator of cold-induced transcription and freezing tolerance in Arabidopsis. Genes Dev. 2003;17(8):1043–1054. doi:https://doi.org/10.1101/gad.1077503.
- Zuo ZF, Kang HG, Park MY, Jeong H, Sun HJ, Yang DH, Lee YE, Song PS, Lee HY. Overexpression of ICE1, a regulator of cold-induced transcriptome, confers cold tolerance to transgenic Zoysia japonica. J Plant Biol. 2019;62:137. doi:https://doi.org/10.1016/j.plantsci.2019.110254.
- Zhu Y, Yang H, Mang HG, Hua J. Induction of BAP1 by a moderate decrease in temperature is mediated by ICE1 in Arabidopsis. Plant Physiol. 2011;155:580–588. doi:https://doi.org/10.1104/pp.110.169466.
- Hu Y, Jiang L, Wang F, Yu D. Jasmonate regulates the inducer of CBF expression-C-repeat binding factor/DRE binding factor1 cascade and freezing tolerance in Arabidopsis. Plant Cell. 2013;25(8):2907–2924. doi:https://doi.org/10.1105/tpc.113.112631.
- Ding Y, Li H, Zhang X, Xie Q, Gong Z, Yang S. OST1 kinase modulates freezing tolerance by enhancing ICE1 stability in Arabidopsis. Dev Cell. 2015b;32(3):278–289. doi:https://doi.org/10.1016/j.devcel.2014.12.023.
- Kim YS, Lee M, Lee JH, Lee HJ, Park CM. The unified ICE-CBF pathway provides a transcriptional feedback control of freezing tolerance during cold acclimation in Arabidopsis. Plant Mol Biol. 2015;89(1):187–201. doi:https://doi.org/10.1007/s11103-015-0365-3.
- Lee JH, Jung JH, Park CM. INDUCER OF CBF EXPRESSION 1 integrates cold signals into FLOWERING LOCUS C-mediated flowering pathways in Arabidopsis. Plant J. 2015;84(1):29–40. doi:https://doi.org/10.1111/tpj.12956.
- Li H, Ding Y, Shi Y, Zhang X, Zhang S, Gong Z, Yang S. MPK3- and MPK6-mediated ICE1 phosphorylation negatively regulates ICE1 stability and freezing tolerance in Arabidopsis. Dev Cell. 2017;43(5):630–642. doi:https://doi.org/10.1016/j.devcel.2017.09.025.
- Zhao C, Wang P, Si T, Hsu CC, Wang L, Zayed O, Yu Z, Zhu Y, Dong J, Tao WA, et al. MAP kinase cascades regulate the cold response by modulating ICE1 protein stability. Dev Cell. 2017;43(5):618–629. doi:https://doi.org/10.1016/j.devcel.2017.09.024.
- Ohta M, Sato A, Renhu N, Yamamoto T, Oka N, Zhu JK, Tada Y, Suzaki T, Miura K. MYC-type transcription factors, MYC67 and MYC70, interact with ICE1 and negatively regulate cold tolerance in Arabidopsis. Sci Rep. 2018;8(1):11622. doi:https://doi.org/10.1038/s41598-018-29722-x.
- MacGregor DR, Zhang N, Iwasaki M, Chen M, Dave A, Lopez-Molina L, Penfield S. ICE1 and ZOU determine the depth of primary seed dormancy in Arabidopsis independently of their role in endosperm development. Plant J. 2019;98(2):277–290. doi:https://doi.org/10.1111/tpj.14211.
- Ye K, Li H, Ding Y, Shi Y, Song CP, Gong Z, Yang S. BRASSINOSTEROID-INSENSITIVE2 negatively regulates the stability of transcription factor ice1 in response to cold stress in Arabidopsis. Plant Cell. 2019;31(11):2682–2696. doi:https://doi.org/10.1105/tpc.19.00058.
- Chinnusamy V, Zhu J, Zhu JK. Gene regulation during cold acclimation in plants. Physiol Plant. 2006;126(1):52–61. doi:https://doi.org/10.1111/j.1399-3054.2006.00596.x.
- Lee BH, Henderson DA, Zhu JK. The Arabidopsis cold responsive transcriptome and its regulation by ICE1. Plant Cell. 2005;17(11):3155–3175. doi:https://doi.org/10.1105/tpc.105.035568.
- Dong CH, Agarwal M, Zhang Y, Xie Q, Zhu JK. The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degradation of ICE1. Proc Natl Acad Sci USA. 2006;103(21):8281–8286. doi:https://doi.org/10.1073/pnas.0602874103.
- Miura K, Jin JB, Lee J, Yoo CY, Stirm V, Miura T, Ashworth EN, Bressan RA, Yun DJ, Hasegawa PM. SIZ1-mediated sumoylation of ICE1 controls CBF3/DREB1A expression and freezing tolerance in Arabidopsis. Plant Cell. 2007;2007(19):1403–1414. doi:https://doi.org/10.1105/tpc.106.048397.
- Agarwal M, Hao Y, Kapoor A, Dong CH, Fujii H, Zheng X, Zhu JK. type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J Biol Chem. 2006;281(49):37636–37645. doi:https://doi.org/10.1074/jbc.M605895200.
- Kanaoka MM, Pillitteri LJ, Fujii H, Yoshida Y, Bogenschutz NL, Takabayashi J, Zhu JK, Torii KU. SCREAM/ICE1 and SCREAM2 specify three cell-state transitional steps leading to Arabidopsis stomatal differentiation. Plant Cell. 2008;20(7):1775–1785. doi:https://doi.org/10.1105/tpc.108.060848.
- Hu Y, Han X, Yang M, Zhang M, Pan J, Yu D. The transcription factor INDUCER OF CBF EXPRESSION1 interacts with ABSCISIC ACID INSENSITIVE5 and DELLA proteins to fine-tune abscisic acid signaling during seed germination in Arabidopsis. Plant Cell. 2019;31(7):1520–1538. doi:https://doi.org/10.1105/tpc.18.00825.
- Chinnusamy V, Zhu JK, Sunkar R. Gene regulation during cold stress acclimation in plants. Methods Mol Biol. 2010;639:39–55. doi:https://doi.org/10.1007/978-1-60761-702-0_3.
- Balasubramanian V, Sie M, Hijmans RJ, Otsuka K. Increasing rice production in Sub-Saharan Africa: challenges and opportunities. Adv Agro. 2007;94:55–133. doi:https://doi.org/10.1080/1343943X.2019.1617638.
- Lv Y, Guo Z, Li X, Ye H, Xiong L. New insights into the genetic basis of natural chilling and cold shock tolerance in rice by genome-wide association analysis. Plant Cell Environ. 2016;39(3):556–570. doi:https://doi.org/10.1111/pce.12635.
- Shakiba E, Edwards JD, Jodari F, Duke SE, Baldo AM, Korniliev P, McCouch SR, Eizenga GC. Genetic architecture of cold tolerance in rice (Oryza sativa) determined through high resolution genome-wide analysis. PLoS One. 2017;12:e0172133. doi:https://doi.org/10.1371/journal.pone.0172133.
- Xiang DJ, Hu XY, Zhang Y, Yin KD. Over-expression of ICE1 gene in transgenic rice improves cold tolerance. Rice Sci. 2008;15(3):173–178. doi:https://doi.org/10.1016/S1672-6308(08)60039-6.
- Zhang Z, Li J, Li F, Liu H, Yang W, Chong K, Xu Y. Phosphorylates OsbHLH002/OsICE1 and inhibits its ubiquitination to activate OsTPP1 and enhances rice chilling tolerance. Dev Cell. 2017;43:731–743. doi:https://doi.org/10.1016/j.devcel.2017.11.016.
- Xiang D, Man L, Yin K, Song Q, Wang L, Zhao M, Xu Z. Overexpression of a ItICE1 gene from Isatis tinctoria enhances cold tolerance in rice. Mol Breed. 2013;32:617–628. doi:https://doi.org/10.1007/s11032-013-9894-0.
- Xiang D, Chai Y, Man L, Sun Y, Zhang T, Wei C, Xie Z, Li H, Zhang W, Liu D, et al. Overexpression of a heading Chinese cabbage ICE1 gene confers freezing tolerance in transgenic rice. Plant Cell Tiss Organ Cult. 2017;128:43–54. doi:https://doi.org/10.1007/s11240-016-1080-8.
- Man L, Xiang D, Wang L, Zhang W, Wang X, Qi G. Stress responsive gene RsICE1 from Raphanus sativus increases cold tolerance in rice. Protoplasma. 2017;254(2):945–956. doi:https://doi.org/10.1007/s00709-016-1004-9.
- Fuhui H, Liu Z, Xie H, Zhu J, Zhang J, Kraus J. Increased drought tolerance through the suppression of ESKMO1 gene and overexpression of CBF-related genes in Arabidopsis. PLoS ONE. 2014;9(9):e106509. doi:https://doi.org/10.1371/journal.pone.0106509.
- Li J, Wanga L, Zhua W, Wanga N, Xin H, Li S. Characterization of two VvICE1 genes isolated from ‘Muscat Hamburg’grapevine and their effect on the tolerance to abiotic stresses. Sci Hort. 2014;165:266–273. doi: https://doi.org/10.1016/j.scienta.2013.11.002.
- Ding ZT, Li C, Shi H, Wang H, Wang Y. Pattern of CsICE1 expression under cold or drought treatment and functional verification through analysis of transgenic Arabidopsis. Genet Mol Res. 2015b;14(3):11259–11270. doi:https://doi.org/10.4238/2015.September.22.20.
- Chen L, Chen Y, Jiang J, Chen S, Chen F, Guan Z, Fang W. The constitutive expression of Chrysanthemum dichrum ICE1 in Chrysanthemum grandiflorum improves the level of low temperature, salinity and drought tolerance. Plant Cell Rep. 2012a;31(9):1747–1758. doi:https://doi.org/10.1007/s00299-012-1288-y.
- Feng HL, Ma NN, Meng X, Zhang S, Wang JR, Chai S, Meng QW. A novel tomato MYC-type ICE1-like transcription factor, SlICE1a, confers cold, osmotic and salt tolerance in transgenic tobacco. Plant Physiol Biochem. 2013;73:309–320. doi:https://doi.org/10.1016/j.plaphy.2013.09.014.
- Nagaveni B, Rama N, Jeyaraman R, Chinnusamy V, Karaba NN. Ectopic expression of AtICE1 and OsICE1 transcription factor delays stress induced senescence and improves tolerance to abiotic stresses in tobacco. J Plant Biochem Biotechnol. 2016;25:285–293. doi:https://doi.org/10.1007/s13562-015-0340-8.
- Zhang T, Mo J, Zhou K, Chang Y, Liu Z. Overexpression of Brassica campestris BcICE1 gene increases abiotic stress tolerance in tobacco. Plant Physiol Biochem. 2018;132:515–523. doi:https://doi.org/10.1016/j.plaphy.2018.09.039.
- Verma RK, Santosh KVV, Yadav SK, Pushkar S, Rao MV, Chinnusamy V. Overexpression of ABA receptor PYL10 gene confers drought and cold tolerance to indica rice. Front Plant Sci. 2019;10:1488. doi:https://doi.org/10.3389/fpls.2019.01488.
- Doyle JJ, Doyle JL. Isolation of plant DNA from fresh tissue. Focus. 1990;12:13–15. doi:https://doi.org/10.2225/vol10-issue3-fulltext-5.
- Jain N, Vergish S, Khurana JP. Validation of house-keeping genes for normalization of gene expression data during diurnal/circadian studies in rice by RT-qPCR. Sci Rep. 2018;8:3203. doi:https://doi.org/10.1038/s41598-018-21374-1.
- Blum A, Ebercon A. Cell membrane stability as a measure of drought and heat tolerance in wheat. Crop Sci. 1981;21:43–47. doi:https://doi.org/10.2135/cropsci1981.0011183X002100010013x.
- Hiscox JD, Israelstam GF. A method for the extraction of chlorophyll from leaf tissue without maceration. Canad J Bot. 1979;57(12):1332–1334. doi:https://doi.org/10.1139/b79-163.
- Heath RL, Parker L. Photoperoxidation in isolated chloroplasts: I. Kinetics and stiochiometry of fatty acid peroxidation. Arch Biochem Biophy. 1968;125(1):189–198. doi:https://doi.org/10.1016/0003-9861(68)90654-1.
- Alexieva V, Sergiev I, Mapelli S, Karanov E. The effect of drought and ultraviolet radiation on growth and stress markers in pea and wheat. Plant Cell Environ. 2001;24(12):1337–1344. doi:https://doi.org/10.1046/j.1365-3040.2001.00778.x.
- Reynolds SG. The gravimetry method of soil moisture determination. J Hydrol. 1970;11(3):258–273. doi:https://doi.org/10.1016/0022-1694(70)90066-1.
- Kusumi K. Measuring Stomatal Density in Rice. Bio-protocol. 2013;3:e753. doi:https://doi.org/10.21769/BioProtoc.753.
- Liu L, Duan L, Zhang J, Zhang Z, Mi G, Ren H. Cucumber (Cucumis sativus L.) over-expressing cold-induced transcriptome regulator ICE1 exhibits changed morphological characters and enhances chilling tolerance. Sci Hort. 2010;124(1):29–33. doi: https://doi.org/10.1016/j.scienta.2009.11.018.
- Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt-, and cold-responsive gene expression. Plant J. 2003;33(4):751–763. doi:https://doi.org/10.1046/j.1365-313x.2003.01661.x.
- Ito Y, Katsura K, Maruyama K, Taji T, Kobayashi M, Seki M. Functional analysis of rice DREB1/CBF-type transcription factors involved in cold-responsive gene expression in transgenic rice. Plant Cell Physiol. 2006;47(1):141–153. doi:https://doi.org/10.1093/pcp/pci230.
- Wang Q, Guan Y, Wu Y, Chen H, Chen F, Chu C. Overexpression of a rice OsDREB1F gene increases salt, drought, and low temperature tolerance in both Arabidopsis and rice. Plant Mol Biol. 2008;67(6):589–602. doi:https://doi.org/10.1007/s11103-008-9340-6.
- Ma Q, Dai X, Xu Y, Guo J, Liu Y, Chen N, Xiao J, Zhang D, Xu Z, Zhang X, et al. Enhanced tolerance to chilling stress in OsMYB3R-2 transgenic rice is mediated by alteration in cell cycle and ectopic expression of stress genes. Plant Physiol. 2009;150(1):244–256. doi:https://doi.org/10.1104/pp.108.133454.
- Ge LF, Chao DY, Shi M, Zhu MZ, Gao JP, Lin HX. Overexpression of the trehalose-6-phosphate phosphatase gene OsTPP1 confers stress tolerance in rice and results in the activation of stress responsive genes. Planta. 2008;228(1):191–201. doi:https://doi.org/10.1007/s00425-008-0729-x.
- Jisha V, Dampanaboina L, Vadassery J, Mithofer A, Kappara S, Ramanan R. Overexpression of an AP2/ERF type transcription factor OsEREBP1 confers biotic and abiotic stress tolerance in rice. PLoS ONE. 2015;10(6):e0127831. doi:https://doi.org/10.1371/journal.pone.0127831.
- Badawi M, Reddy YV, Agharbaoui Z, Tominaga Y, Danyluk J, Sarhan F, Houde M. Structure and functional analysis of wheat ICE (Inducer of CBF Expression) genes. Plant Cell Physiol. 2008;49(8):1237–1249. doi:https://doi.org/10.1093/pcp/pcn100.
- Chander S, Almeida DM, Serra TS, Jardim-Messeder D, Barrosa PM, Lourenço TF, Figueiredo DD, Margis-Pinheiro M, Costa JM, Oliveira MM, et al. OsICE1 transcription factor improves photosynthetic performance and reduces grain losses in rice plants subjected to drought. Environ Exp Bot. 2018;150:88–98. doi:https://doi.org/10.1016/j.envexpbot.2018.02.004.
- Miura K, Shiba H, Ohta M, Kang SW, Sato A, Yuasa T, Iwaya-Inoue M, Kamada H, Ezura H. SlICE1 encoding a MYC-type transcription factor controls cold tolerance in tomato, Solanum lycopersicum. Plant Biotechnol. 2012;29(3):253–260. doi:https://doi.org/10.5511/plantbiotechnology.12.0303a.
- Feng XM, Zhao Q, Zhao LL, Qiao Y, Xie XB, Li HF, Yao YX, You CX, Hao YJ. The cold-induced basic helix-loop-helix transcription factor gene MdCIbHLH1 encodes an ICE-like protein in apple. BMC Plant Biol. 2012;12:22. doi:https://doi.org/10.1186/1471-2229-12-22.
- Chen Y, Jiang J, Song A, Chen S, Shan H, Luo H, Gu C, Sun J, Zhu L, Fang W, et al. Ambient temperature enhanced freezing tolerance of Chrysanthemum dichrum CdICE1Arabidopsis via miR398. BMC Biol. 2013b;11:121. doi:https://doi.org/10.1186/1741-7007-11-121.
- Dong C, Zhang Z, Ren JP, Qin Y, Huang JF, Wang Y, Cai BH, Wang BL, Tao JM. Stress-responsive gene ICE1 from Vitis amurensis increases cold tolerance in tobacco. Plant Physiol Biochem. 2013;71:212–217. doi:https://doi.org/10.1016/j.plaphy.2013.07.012.
- Xu W, Jiao Y, Li R, Zhang N, Xiao D, Ding X. Chinese wild-growing Vitis amurensis ICE1 and ICE2 encode MYC-type bHLH transcription activators that regulate cold tolerance in Arabidopsis. PLoS ONE. 2014;9:e102303. doi:https://doi.org/10.1371/journal.pone.0102303.
- Lin Y, Zheng H, Zhang Q, Liu C, Zhang Z. Functional profiling of EcaICE1 transcription factor gene from Eucalyptus camaldulens is involved in cold response in tobacco plants. J Plant Biochem Biotechnol. 2014a;23(2):141–150. doi:https://doi.org/10.1007/s13562-013-0192-z.
- Peng PH, Lin CH, Tsai HW, Lin TY. Cold response in phalaenopsis aphrodite and characterization of PaCBF1 and PaICE1. Plant Cell Physiol. 2014;55(9):1623–1635. doi:https://doi.org/10.1093/pcp/pcu093.
- Yu XH, Juan JX, Gao ZL, Zhang Y, Li WY, Jiang XM. Cloning and transformation of INDUCER of CBF EXPRESSION1 (ICE1) in tomato. Genet Mol Res. 2015;14(4):13131–13143. doi:https://doi.org/10.4238/2015.October.26.9.
- Huang X, Li K, Jin C, Zhang S. ICE1 of Pyrus ussuriensis functions in cold tolerance by enhancing PuDREBa transcriptional levels through interacting with PuHHP1. Sci Rep. 2015a;5:17620. doi:https://doi.org/10.1038/srep1762.
- Huang X, Zhang Q, Zhu D, Fu X, Wang M, Zhang Q, Moriguchi T, Liu J. ICE1 of Poncirus trifoliata functions in cold tolerance by modulating polyamine levels through interacting with arginine decarboxylase. J Exp Bot. 2015b;66(11):3259–3274. doi:https://doi.org/10.1093/jxb/erv138.
- Deng C, Ye H, Fan M, Pu T, Yan J. The rice transcription factors OsICE confer enhanced cold tolerance in transgenic Arabidopsis. Plant Signal Behav. 2017;12(5):e1316442. doi:https://doi.org/10.1080/15592324.2017.1316442.
- Lu X, Yang L, Yu M, Lai J, Wang C, McNeil D, Zhou M, Yang C. A novel Zea mays ssp. mexicana L. MYC-type ICE-like transcription factor gene ZmmICE1, enhances freezing tolerance in transgenic Arabidopsis thaliana. Plant Physiol Biochem. 2017;113:78–88. doi:https://doi.org/10.1016/j.plaphy.2017.02.002.
- Yuan HM, Sheng Y, Chen WJ, Lu YQ, Tang X, Yang MO, Huang X. Overexpression of Hevea brasiliensis HbICE1 enhances cold tolerance in Arabidopsis. Front Plant Sci. 2017;8:1462. doi:https://doi.org/10.3389/fpls.2017.01462.
- Zuo ZF, Kang HG, Park MY, Jeong H, Sun HJ, Yang DH, Lee Y.E., Song, P.S., Lee, H.Y. Overexpression of ICE1, a regulator 643 of cold-induced transcriptome, confers cold tolerance to transgenic Zoysia japonica. J. Plant Biol. 2019;62:137. doi:https://doi.org/10.1007/s12374 -018-0330-1.
- Zuo ZF, Kang HG, Hong QC, Park MY, Sun HJ, Kim J, Song PS, Lee HY. Anovel basic helix-loop-helix transcription factor, ZjICE2 from Zoysia japonica confers abiotic stress tolerance to transgenic plants via activating the DREB/CBF regulon and enhancing ROS scavenging. Plant Mol Biol. 2020;102:447–462. doi:https://doi.org/10.1007/s11103-019-00957-0.
- Zhou L, He YJ, Li J, Li LZ, Liu Y, Chen HY. An eggplant SmICE1a gene encoding MYC‐type ICE1‐like transcription factor enhances freezing tolerance in transgenic Arabidopsis thaliana. Plant Biol. 2020;22:3450–3458. doi:https://doi.org/10.1111/plb.13095.
- Zhu X, Zhao X, Ren T, Ma Y, Wang Y, Fang W. CsICE1 Functions in cold tolerance by regulating polyamine levels may through interacting with arginine decarboxylase in the tea tree. Agriculture. 2020;10(6):201. doi:https://doi.org/10.3390/agriculture10060201.
- Luo P, Li Z, Chen W, Xing W, Yang J, Cui Y. Overexpression of RmICE1, a bHLH transcription factor from Rosa multiflora, enhances cold tolerance via modulating ROS levels and activating the expression of stress-responsive genes. Environ Exp Bot. 2020;178:104160. doi:https://doi.org/10.1016/j.envexpbot.2020.104160.