ABSTRACT
Cholinesterase is consisting of acetylcholinesterase (AChE) and pseudocholinesterase in vertebrates and invertebrates. AChE gene has been identified in several plant species, while pseudocholinesterase gene has not yet been found in any plant species. In this study, we report that the AChE gene paralog encodes propionylcholinesterase (PChE), a pseudocholinesterase in rice. PChE was found to be located adjacent to AChE (Os07g0586200) on rice chromosome 7 and designated as Os07g0586100. Phylogenetic tree analysis showed a close relationship between rice AChE and PChE. PChE-overexpressing rice had higher hydrolytic activity toward propionylthiocholine than acetylthiocholine and showed extremely low activity against butyrylthiocholine. Therefore, the PChE gene product was characterized as a propionylcholinesterase, a pseudocholinesterase. The rice PChE displayed lower sensitivity to the cholinesterase inhibitor, neostigmine bromide, than electric eel, maize, and rice AChEs. The recombinant PChE functions as a 171 kDa homotetramer. PChE was expressed during the later developmental stage, and it was found be localized in the extracellular spaces of the rice leaf tissue. These results suggest that the rice plant possesses PChE, which functions in the extracellular spaces at a later developmental stage. To the best of our knowledge, this study provides the first direct evidence and molecular characterization of PChE in plants.
Introduction
The ACh-mediated nerve system comprising three components – acetylcholine (ACh), ACh-receptor (AChR), and acetylcholinesterase (AChE) – is a well-known signal transmission system in animals.Citation1–4 AChE (E.C. 3.1.1.7) is a key enzyme that stops neurotransmission by catalyzing the hydrolysis of the neurotransmitter ACh in the synaptic cleft.Citation1–4 Vertebrates and invertebrates also contain an evolutionarily-related pseudocholinesterase (E.C. 3.1.1.8) comprising butyrylcholinesterase (BChE) and propionylcholinesterase (PChE).Citation5 These pseudocholinesterases preferentially cleave their specific substrates, viz., butyrylcholine (BCh) and propionylcholine (PCh), rather than ACh.Citation6 The function of BChE is unclear in animals; however, it is believed to coregulate the activity of the neurotransmitter AChCitation7 and play a role in neuronal development.Citation8 PChE has not been well studied, and there are few reports on the function of PChE. Several reports have suggested that PChE can play a role in inflammatory processesCitation9 and in the nervous and neuromuscular systems.Citation10 PChE in sea urchin embryos and larvae from sperm respond to stress against the toxicity of engineered nanoparticles.Citation11
The enzymatic activity hydrolyzing ACh has been widely detected in plants. In the recent studies, ACh-hydrolyzing activity has been detected in pollen grains; this enzymatic activity is related to self-incompatibility regulation, pollen germination, pollen tube growth, and stigma penetration.Citation12,Citation13 However, the amino acid sequence of the plant protein with ACh-hydrolyzing activity and the nucleotide sequence of its encoding gene have been unknown until recently.
In our efforts to study the gene encoding a protein having ACh-hydrolyzing activity in plants, we have recently identified AChE genes in maize (Zea mays L.), rice (O. sativa L. ssp. Japonica cv. Nipponbare), siratro (Macroptilium atropurpureum Urb.), and glasswort (Salicornia europaea L.).Citation14–17 We reported that plant AChE is a positive regulator of heat stress and shoot gravitropism based on a genetic study using transgenic plants altering the expression of plant AChE genes.Citation17–19 In contrast, direct evidence of the molecules and genes responsible for both BChE and PChE in plants have not yet been identified, although there are few reports on BChE and PChE activities in the seeds of Alliumaltaicum and roots of Solanum tuberosum.Citation20,Citation21 Therefore, in this study, we aimed to identify a pseudocholinesterase gene in plants and characterize its gene product.
This study is the first to identify and characterize the enzymatic and molecular properties of PChE in rice, providing new insights into plant neurobiology.
Materials and methods
Plant specimens
The rice seeds (Oryza sativa L. cv. Nipponbare) were grown on glass beads in a growth chamber at 28°C with a photoperiod of 16 h light/day. For immunoblotting analysis and cholinesterase activity assay, the rice plants were harvested after 14 days of sowing. For RT-PCR analysis, the rice seedlings were harvested after 4, 6, 8, 10, 15, and 20 days of sowing. For immunohistochemistry studies, the rice plants were grown for 1 month.
Immunoblotting analysis
Rice PChE protein was extracted from 2 g of rice leaves, according to the protocol described in previous studies.Citation14 Thereafter, the extracted samples were subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE, 12.5% gel) and blotted onto a nitrocellulose membrane. Subsequently, the nitrocellulose membrane was stained with Ponceau S staining solution [0.1% (w/v) Ponceau S in 1% (v/v) acetic acid] as a loading control and washed with distilled water; then, a maize AChE polyclonal antibody was added as a probe. This antibody was generated by Hokkaido System Science Co., Ltd. (Hokkaido, Japan) against a recombinant maize AChE protein expressed in E.coli. As described in previous studies, a 1:20 dilution of the maize AChE polyclonal antibody was used as the primary antibody, and a 1:16000 dilution of anti-rabbit IgG-horseradish peroxidase conjugate (Sigma–Aldrich, St. Louis, MO, U.S.A.) was added as the secondary antibody.Citation18 To obtain the secondary antibody control, a nitrocellulose membrane was stained only with the secondary antibody without addition of the maize AChE polyclonal antibody after the blotting process.
Cholinesterase activity assay
Cholinesterase (ChE) activity was measured with three substrates, viz., ASCh, PSCh, and BSCh, using Ellman’s reagent (5,5-dithiobis-2-nitrobenzoic acid, DTNB), as described previously.Citation14
Cloning of the rice PChE gene and construction of a vector for gene overexpression in rice
The PChE gene (accession no. AK071404) was identified based on BLAST searches using the rice AChE amino acid sequence (accession no. AK073754) as the query sequence. Total RNA was extracted and purified from 0.1 g of 5-day-old rice seedling shoots, and cDNA was obtained from total RNA using the SuperScript II first-strand synthesis system (Invitrogen, Carlsbad, CA, U.S.A.) with oligo (dt) primer. Rice PChE was amplified using PCR with specific primers (Supplementary Table 1) with the first-strand cDNA as the template, as described previously.Citation17 The amplified PChE gene was cloned into the pT7Blue blunt vector (pT7Blue-rice PChE) in the antisense orientation. The gene and plant binary vector, p2K-1+ was then digested with KpnI and SpeI. The PChE was introduced downstream of the maize ubiquitin1 promoter (pUbi) in the p2K-1+ vector. This construct was named pUbi::rice PChE. Agrobacterium tumefaciens strain EHA 101-transformed pUbi::rice PChE was used to generate transgenic rice plants by Agrobacterium-mediated transformation of rice calli.Citation22 Regenerated transgenic rice was grown in a closed growth system (Koitotron type KG: Koito Ind. Ltd. Co., Tokyo, Japan) at 28°C under a photoperiod of 16 h light/day.
Construction of binary vectors for gene downregulation
The inverted repeat region was obtained by joining the 5́- and 3́-gene-specific regions of PChE.
Initially, the 5́-gene-specific region of PChE was amplified using the specific primers (Supplementary Table 1) and subcloned into the pT7Blue blunt vector (Novagen, Madison, WI, U.S.A.). The 3́-gene-specific regions of rice PChE were then amplified and ligated into the above-mentioned vector digested with SmaI. Next, the chimeric gene-specific region was amplified using the 5́-UTR-F and 3́-UTR-R primers (Supplementary Table 1), inserted into the pENTR/D-TOPO cloning vector (Invitrogen, Carlsbad, CA, U.S.A.) to generate an entry vector, and then introduced into the RNA silencing vector, pANDA vector, using an LR clonase reaction between the entry vector and the pANDA vector.Citation23,Citation24 This RNAi plasmid was designated as PChE-KD. Agrobacterium tumefaciens strain EHA 101-transformed PChE-KD was used to generate the transgenic rice plants, as described above.
RT-PCR analysis
The transcription level of PChE was analyzed by semiquantitative RT-PCR. First-strand cDNA was synthesized according to the method described by Yamamoto et al.Citation17 One microliter of the synthesized first-strand cDNA was added into the PCR mixture with different sets of primers (Supplementary Table 1) and Blend Taq Polymerase (TOYOBO, Osaka, Japan). The rice ubiquitin gene (UBQ; accession no. D12629) was used as an internal control, as described in previous studies.Citation23,Citation24 PChE and UBQ were amplified with 35 and 25 cycles, respectively. After PCR reaction, agarose gel electrophoresis (1.5% gel) was carried out to separate the PCR products.
The transcription levels of PChE in both PChE-overexpressing rice and RNAi rice were analyzed using semiquantitative RT-PCR. Total RNA was extracted from 4-day-old transgenic rice seedlings (T2 generation) according to the method described by Yamamoto et al.Citation17 The synthesis of first-strand cDNA, PCR, and agarose gel electrophoresis of PCR products were performed following the same procedures described above.
Enzymatic characterization of rice PChE
To determine the substrate selectivity of rice PChE, three kinds of substrates (ASCh, PSCh, and BSCh) were used at concentrations ranging from 0.05 to 100 mM. The inhibitor, neostigmine bromide, was tested at an overdose range of 1–30 μM. The total protein extract from the aerial parts of 14-day-old PChE-overexpressing rice plants (pUbi::PChE No. 18) was used as the sample for this purpose.
Immunohistochemistry
Leaf tissues of wild-type or PChE-overexpressing rice were fixed for 16 hours at 4°C using 5% formaldehyde, 5% acetic acid, and 70% ethanol (FAA). The fixed leaves were dehydrated and embedded in paraffin (Paraplast Plus, Sigma–Aldrich, St. Louis, MO, U.S.A.). Sections of 6 µm thickness were cut using a rotary microtome; thereafter, these sections were deparaffinized and rehydrated. Subsequently, they were labeled with anti-maize AChE (diluted 1:50 with 0.5% BSA in PBST), followed by Alexa Fluor 488-conjugated anti-rabbit IgG (diluted 1:300 with 0.5% BSA in PBST) (Molecular Probes, Eugene, OR, USA), as described previously.Citation18 To obtain the secondary antibody control, the sections were stained using only Alexa Fluor 488-conjugated anti-rabbit IgG without addition of the primary antibody. Images were captured using an Olympus BX51 fluorescence microscope (Olympus, Tokyo, Japan).
Estimation of molecular mass of active form of AChE and PChE from transgenic rice
The crude enzyme solution, including rice AChE or PChE, was extracted from the leaves of AChE-overexpressing rice (pUbi::AChE No. 4) generated in a previous studyCitation17 and PChE-overexpressing rice (pUbi::PChE No. 18) generated in the present study. Two grams of these leaves were ground to a fine powder in liquid nitrogen with a mortar and pestle. Fifteen milliliters of extraction buffer (10 mM potassium phosphate buffer, pH 7.0, 10 mM EDTA) was added to the powdered sample and the mixture was homogenated. The homogenate was centrifuged at 15,000 × g at 4°C for 30 min, and the supernatant was collected as the sample for gel filtration chromatography.
Next, 500 μL of each crude enzyme solution was injected into a Superdex 200 Increase 10/300 GL column (GE Healthcare, U.S.A.) equilibrated with 50 mM potassium phosphate buffer (pH 7.0) containing 150 mM NaCl. The elution volumes of the enzymes in their native forms were detected by measuring the enzyme activity using the Ellman’s reagent as described above. A gel filtration calibration kit (GE Healthcare, U.S.A.) was used to estimate the molecular mass of rice AChE and PChE according to the manufacturer’s instructions.
Sequence analysis
The amino acid sequence of the rice PChE protein was analyzed for signal peptides using the SignalP5.0 (http://www.cbs.dtu.dk/services/SignalP-5.0/) and an in-silico search using the protein family database Pfam (http://www.sanger.ac.uk/Software/Pfam). For phylogenetic tree analysis, the rice PChE homologues (accession numbers XM_016837348, XM_003554018, AK451334, and AK366643) were obtained using a tBLASTn search (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Phylogenetic trees were calculated from the distances provided by ClustalW analysis using the neighbor-joining method, as described by Yamamoto et al.Citation15 MEGA X software (https://www.megasoftware.net/) was used for the construction of the phylogenetic tree.
Results
Cloning and sequence analysis of the rice PChE gene
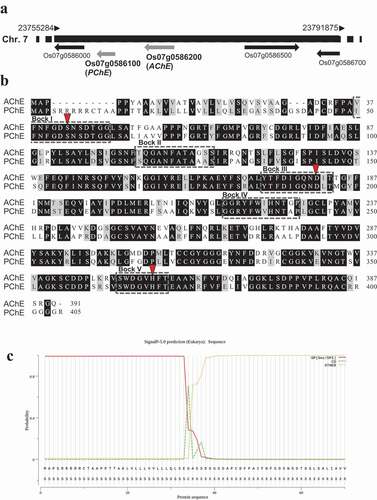
BLAST searches using the rice AChE amino acid sequence (accession number AK073754) as the query sequence revealed a paralogous gene, PChE, that shared high similarity (81% and 69% at the nucleotide and amino acid levels, respectively) with rice AChE. The rice PChE gene (designated Os07g0586100) was located adjacent to the rice AChE gene (designated Os07g0586200) on chromosome 7 (). Os07g0586100 was 1440 nucleotides long and encoded 405 amino acids. The in-silico search using Pfam revealed that the putative PChE protein belonged to the putative GDSL lipase family and contained five blocks of conserved amino acids distributed from residues 50–382, similar to rice AChE (). The 34 amino acid residues in the N-terminal of the PChE protein were predicted to be a signal peptide that routes the protein to a secretory pathway in a manner similar to that observed for rice AChE ().
Figure 1. Sequence analysis of rice AChE and PChE. (a) Chromosomal locations of rice AChE and PChE. Rice AChE and PChE are located on chromosome 7 and designated as Os07g0586200 and Os07g0586100, respectively. (b) Alignment of the amino acid sequences of rice AChE and PChE. The black box shows conserved amino acids. The gray box indicates nonidentical but functionally conserved amino acids. The conserved sequences in the GDSL enzyme family are indicated by boxes (dashed lines, Blocks I–VCitation25–27). Arrowheads indicate the amino acid residues related to the putative catalytic triad in the GDSL family enzymes. The GenBank accession numbers used in the analysis are as follows: rice AChE: AK073754; rice PChE: AK071404. (c) Prediction of signal peptide cleavage site of rice PChE using SignalP5.0. The three kinds of possibilities are reported on the plot, namely, Sec/SPI (secretion signal peptide), CS (cleavage site), and OTHER (probability that the sequence does not have any kind of signal peptide)
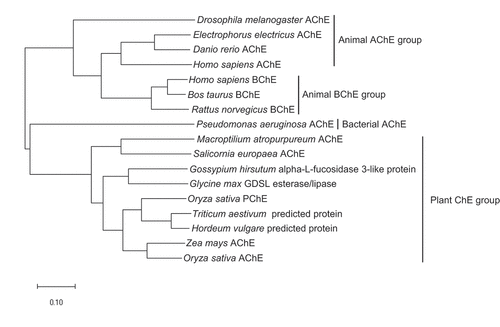
The phylogenetic tree based on the amino acid sequences of cholinesterases isolated from various organisms showed clear divergence for four groups, namely, animal AChE, animal BChE, bacterial AChE, and plant ChE. In addition, the phylogenetic analysis revealed that the amino acid sequence of rice PChE clustered in the same monophyletic group as two predicted proteins of Triticum aestivum and Hordeum vulgare. Moreover, the rice PChE clustered in the same paraphyletic group as the maize and rice AChE proteins ().
Figure 2. Phylogenetic tree derived from the analysis of amino acid sequences of the cholinesterase genes in various organisms. The scale bar indicates the number of amino acid substitutions. GenBank accession numbers used in the analysis are as follows: Drosophila melanogaster AChE, X05893; Electrophorus electricus AChE, AF030422; Danio rerio AChE, NM_131846; Homo sapiens AChE, M55040; Homo sapiens BChE, NM_000055; Bos taurus BChE, NM_001076906; Rattus norvegicus BChE, AF244349; Pseudomonas aeruginosa AChE, KF981438; Macroptilium atropurpureum AChE, AB294246; Salicornia europaea AChE, AB489863; Gossypium hirsutum alpha-L-fucosidase 3-like protein, XM_016837348; Glycine max GDSL esterase/lipase, XM_003554018; Oryza sativa PChE, AK071404; Triticum aestivum predicted protein, AK451334; Hordeum vulgare predicted protein, AK366643; Zea mays AChE, AB093208; Oryza sativa AChE, AK073754
ChE activity in transgenic rice plants
AChE activity assay using ASCh as the substrate was performed in PChE-overexpressing and PChE-suppressing rice to clarify whether the PChE gene product had choline ester-hydrolyzing activity.
The PChE gene was introduced to the maize ubiquitin1 promoter pUbi to overexpress the target gene and was then transformed into the rice. ChE activity assays were performed against two independent hygromycin-resistant plants. shows that pUbi::PChE No. 9 and No. 18 (T2 generation) had 5.3-fold and 6.8-fold higher ChE activity than wild-type plants, respectively. In RT-PCR analysis, PChE-overexpressing rice showed an increase in gene expression levels compared to wild-type plants, thereby supporting the fact that transgenic rice (pUbi::PChE No. 9 and No. 18) have a higher ChE activity than wild-type plants (). However, no phenotypic changes were observed between wild-type and PChE-overexpressing plants under normal growth conditions ().
Figure 3. Generation of transgenic rice with altered expression of PChE gene (a) ChE activity in two different transformants overexpressing PChE. The values are presented as means ± SE of three replicates. The results were analyzed using Student’s t-test; asterisk indicates p values <.01, that is, these are considered significantly different from the wild-type. (b) RT-PCR analysis of PChE mRNA expression in transgenic (pUbi::PChE No. 9 and No. 18) and wild-type rice. The left side of each panel designates the primers used; AChE: acetylcholinesterase gene; PChE: propionylcholinesterase gene; UBQ: ubiquitin-encoding gene as an internal control. (c) Typical phenotypes of 14-day-old transgenic rice plants (T2 generation) overexpressing the rice PChE. (d) ChE activity of transgenic (PChE-KD No. 9 and No. 11) and wild-type rice plants measured using Ellman’s reagent with ASCh as the substrate as described in the Materials and methods section. The values are presented as means ± SE of three replicates. The results were analyzed using Student’s t-test; asterisk indicates p values <.01, that is, these are considered significantly different from the wild-type. (e) RT-PCR analysis of PChE mRNA expression in transgenic (PChE-KD No. 9 and No. 11) and wild-type rice. The left side of each panel designates the primers used; AChE: acetylcholinesterase; PChE: propionylcholinesterase gene; UBQ: ubiquitin-encoding gene as an internal control. (f) Typical phenotypes of 14-day-old transgenic rice plants (T2 generation) with suppressed expression of the rice PChE.
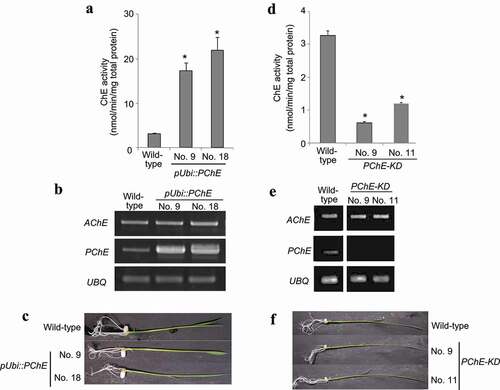
Next, the transgenic rice in which the expression of PChE gene was suppressed by RNAi was generated. Two independent hygromycin-resistant transgenic lines (PChE-KD No. 9 and No. 11) were selected for the ChE activity and RT-PCR assays. RNAi expression reduced ChE activity and its gene expression levels, compared with those in wild-type plants (, e). There was no phenotypic difference between 14-day-old wild-type and PChE-suppressing rice plants (). These results indicate that the PChE gene encodes a choline ester-hydrolyzing enzyme.
Enzymatic properties of rice PChE
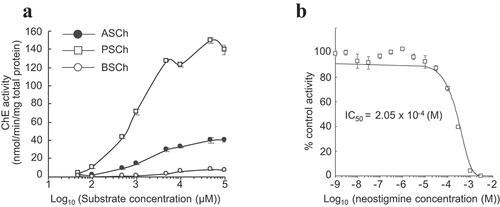
pUbi::PChE No. 18, which had the highest PChE expression, was selected to investigate the substrate selectivity of PChE. The enzymatic activity was analyzed using three kinds of substrates: ASCh, PSCh, and BSCh. PChE showed higher enzymatic activity against PSCh than against ASCh. The activity of PChE increased with increasing concentrations of PSCh and ASCh. However, it exhibited almost no hydrolytic activity against BSCh (). In addition, PChE activity against PSCh was inhibited by neostigmine bromide, with an apparent IC50 value of 2.05 × 10−4 M for PChE (, Supplementary Table 2). Nonetheless, the IC50 value of rice PChE to neostigmine bromide was approximately 4-fold higher than that of electric eel AChE, similar to that of siratro AChE (Supplementary Table 2).
Molecular properties of rice PChE
The molecular mass of monomeric PChE protein in the crude enzyme solution obtained from the leaves of PChE-overexpressing rice (pUbi::PChE No.9 and No. 18) was determined on a immunoblotting analysis labeled with the maize AChE antibody. When the samples obtained from the leaves of PChE-overexpressing rice were subjected to immunoblotting analysis, the main band appeared at approximately 45 kDa. There was no specific signal from the maize AChE antibody in the samples obtained from the leaves of wild-type plants, which were used as a control (). This result suggests that the antibody cannot detect endogenous AChE and PChE proteins in wild-type plants due to low affinity to both ChE proteins. Furthermore, the same observation was obtained in case of the secondary antibody control Supplementary (). Therefore, after immunoblotting analysis of the samples obtained from the leaves of PChE-overexpressing rice plants, the band that appears approximately at 45 kDa would be the PChE band. In addition, truncated forms of the enzyme were detected at values similar to that of rice AChE described in the study by Yamamoto et alCitation17 ().
Figure 5. Molecular characterization of rice PChE (a) Immunoblotting analysis of total protein isolated from transgenic (pUbi::PChE No. 9 and No. 18) and wild-type rice. The upper panel exhibits immunoblotting analysis with anti-maize AChE as probe. The lower panel shows loading control stained with Ponceau S to confirm equal loading of samples. (b) Estimation of the molecular mass of the active form of rice PChE using gel filtration chromatography. X-axis represents the ratio of void volume and elution volume (Ve/Vo) and Y-axis represents the log molecular mass (kDa). The closed circle shows molecular mass of the active form of rice PChE. The open squares show a series of marker proteins: ferritin (440 kDa); aldorose (158 kDa); conalbumin (75 kDa); ovalbumin (43 kDa); carbonic anhydrase (29 kDa); Ribonuclease A (13.7 kDa); Aprotinin (6.5 kDa). (c) Elution profile of active form-AChE (line, left Y-axis) and PChE (dotted line, right Y-axis) on a Superdex 200 Increase 10/300 GL column. X-axis represents the elution volume (mL). Y-axis shows ChE activity as the absorbance at 412 nm
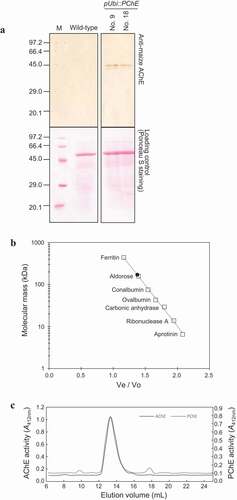
The native molecular mass of rice PChE was determined by the size exclusion chromatography method by using crude enzyme solution extracted from pUbi::PChE No. 18, and it was estimated to be approximately 171 kDa (). In addition, the native molecular mass of rice AChE was determined by using the above method. The elution volume of active peak of both rice AChE and PChE were identical in the size exclusion chromatography (). These results indicated that the active forms of both rice AChE and PChE exist as an approximately 171 kDa homotetramer.
Expression profiles and subcellular localization of rice PChE
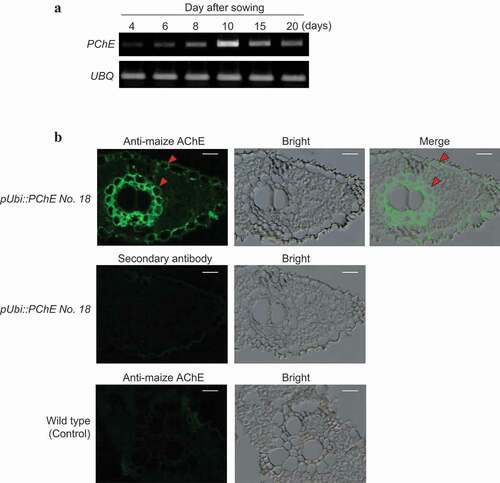
The transcription level of PChE was low during early development (4–8 days after sowing); however, it increased gradually and peaked at approximately 10 days after sowing (). PChE protein in the leaf tissues of the transgenic rice plant pUbi::PChE No. 18 was detected using maize AChE antibody to clarify the subcellular localization of PChE. The PChE protein was mainly localized in the extracellular spaces of the leaves of rice plants (). Thus, rice PChE functions in the extracellular spaces during the later developmental stages.
Figure 6. Expression profile and subcellular localization of PChE (a) Expression profile of PChE was investigated by semiquantitative RT-PCR analysis using total RNA extracted 4, 6, 8, 10, 15, and 20 days after sowing. The left side of each panel designates the primers used: PChE (propionylcholinesterase gene); UBQ (ubiquitin-encoding gene) as an internal control (Supplementary Table 1). (b) Subcellular localization of PChE was performed by immunofluorescence methods. The left side of each panel designates the plant specimen. The cross sections (6 μm thickness) of leaves of PChE-overexpressing rice (T2 generation) were labeled using maize AChE antibody as described in the “Materials and methods section” (upper panels). For the secondary antibody control, the sections were stained using only Alexa Fluor 488-conjugated anti-rabbit IgG without addition of the primary antibody (a center left panel). The sections obtained from wild-type plants were used as controls (lower panels). The arrowheads indicate the points of PChE localization. Each scale bar represents 10 μm. The left, middle, and right panels show fluorescent, bright field, and merged images, respectively
Discussion
In this study, we aimed to identify a propionylcholinesterase gene in rice plants and characterize the enzymatic activity and the molecular properties of its gene product. PChE (Os07g0586100) had a high similarity with the rice AChE (Os07g0586200) as observed in the in-silico search. We speculated that Os07g0586100 is closely related to rice AChE and therefore, Os07g0586100 could be a pseudocholinesterase gene in rice plants. Furthermore, Os07g0586100 was located adjacent to AChE (Os07g0586200) on rice chromosome 7. This close physical association suggests that AChE and PChE might compose a gene cluster – a set of two or more genes that encode the same or similar products and share common biological functions. There are other reports of related plant genes residing at the same location, e.g., the rice momilactone gene cluster.Citation28 The phylogenetic tree, which shows the evolutionary relationships among various ChE, indicated that rice PChE and AChE belong to a paraphyletic group, signifying that there are close evolutionary relationships between rice PChE and AChE.
The PChE gene was overexpressed and suppressed in rice to clarify whether PChE proteins have choline ester-hydrolyzing activity. The PChE-overexpressing plants exhibited higher ChE activity than the wild-type plants, while transgenic rice with suppressed PChE expression had significantly reduced activity against ASCh. Moreover, rice PChE hydrolyzed PSCh at a greater rate than ASCh. In addition, rice PChE showed extremely low activity against BSCh, similar to that of siratro, maize, and electric eel AChEs.Citation15 These results strongly suggest that rice PChE is a propionylcholinesterase. This is the first report of the PChE gene and proteins in plants, although PChE proteins have been previously identified from several animal species, namely Murex trunculus,Citation29 Hirudo medicinalis,Citation30 and Murex brandaris.Citation6
The peak of PChE activity in the crude extract from transgenic plants appeared as a single active peak, with an apparent molecular mass of 171 kDa observed upon analytical gel filtration. Immunoblotting analysis revealed that the molecular mass of monomeric PChE in the transgenic rice plants was approximately 45 kDa. The molecular mass of the active form and the monomeric rice AChE was similar to that of rice PChE.Citation17 These results suggest that the active form of rice PChE and AChE exist as a 171 kDa homotetramer comprising approximately 45 kDa monomers. In animals, the active AChE has been found as tetramers that assemble around a collagen tail and a proline-rich peptide to anchor to the cell surface or the extracellular matrix. BChE also exists primarily as a soluble tetramer.Citation31 Therefore, both rice AChE and PChE function in the same quaternary protein structure, similar to that of animal cholinesterase, although it is unclear whether a collagen tail and a proline-rich peptide are assembled in this case.
During rice seedling growth, PChE expression gradually increased in the later stages of early development (10–20 days after sowing). We had previously reported that the transcription level of rice AChE was high from 4 to 10 days after sowing;Citation17 AChE, but not PChE, was primarily expressed early, and the expression of the two genes was reversed in the later developmental stages, indicating sequential regulation. Thus, the function of AChE may be important during early developmental stages (4–10 days after sowing), while PChE function may be critical in the later stages (10–20 days after sowing). However, there were no phenotypic differences between 14-day-old wild-type rice plants and transgenic rice plants with either overexpressed or suppressed PChE. These results suggest that PChE cannot be related to the growth of rice plants during early development.
Based on the results of immunohistochemistry, we speculated rice PChE was localized in the extracellular spaces of rice leaf tissues, where the PChE gene was found to be overexpressed. Furthermore, SignalP program presumed that the rice PChE is targeted to the secretory pathway. Maize and rice AChE proteins were detected, and their respective genes were found to be overexpressed in the extracellular spaces of the leaves of transgenic rice.Citation17,Citation32 Therefore, rice PChE functions in the extracellular spaces of cells, similar to other plant AChEs.
Propionylcholine (PCh) has also been found in the leaves of five species of flowering plants, which have ACh and ACh-hydrolyzing activities.Citation33 Indeed, propionylcholine in animals is known as a choline-derivative precursor of ACh that predominantly acts on muscarinic receptors in the colonic surface epithelium.Citation34,Citation35 Therefore, we advocate a hypothesis that PCh acts on certain receptors, such as AChR, and that PChE regulates the activity of PCh by degrading it in plants. In the future, we propose to identify the gene encoding a PCh-binding protein and elucidate the signal transduction pathways that signal via PChE.
Supplemental Material
Download Zip (245.6 KB)Acknowledgments
We would like to thank Dr. K. Shimamoto and Dr. D. Miki (Nara Institute of Science and Technology, Japan) for providing p2K-1+ and pANDA vectors, respectively.
Disclosure statement
No potential conflict of interest was reported by the author(s).
Supplementary material
Supplemental data for this article can be accessed on the publisher’s website
Additional information
Funding
References
- Soreq H, Acetylcholinesterase SS. new roles for an old actor. Nat Rev Neurosci. 2001;2:1–9. doi:https://doi.org/10.1038/35067589.
- Hill M. The neuromuscular junction disorders. J Neurol Neurosurg Psychiatry. 2003;74:ii32–ii37. doi:https://doi.org/10.1136/jnnp.74.suppl_2.ii32.
- Lai K-O, Ip NY. Central synapse and neuromuscular junction: same players, different roles. Trends Genet. 2003;19(7):395–402. doi:https://doi.org/10.1016/s0168-9525(03)00147-1.
- Ferraro E, Molinari F, Berghella L. Molecular control of neuromuscular junction development. J Cachexia Sarcopenia Muscle. 2012;3(1):13–23. doi:https://doi.org/10.1007/s13539-011-0041-7.
- Falugi C, Aluigi MG. Early appearance and possible functions of non-neuromuscular cholinesterase activities. Front Mol Neurosci. 2012;5:54. doi:https://doi.org/10.3389/fnmol.2012.00054.
- Talesa V, Contenti S, Mangiabene C, Pascolini R, Rosi G, Principato GB. Propionylcholinesterase from Murex brandaris: comparison with other invertebrate cholinesterases. Comparative Biochemistry and Physiology Part C: Comparative Pharmacology. 1990;96(1):39–43. doi:https://doi.org/10.1016/0742-8413(90)90041-7.
- Darvesh S, Hopkins DA, Geula C. Neurobiology of butyrylcholinesterase. Nat Rev Neurosci. 2003;4(2):131–138. doi:https://doi.org/10.1038/nrn1035.
- Geula C, Nagykery N. Butyrylcholinesterase activity in the rat forebrain and upper brainstem: postnatal development and adult distribution. Exp Neurol. 2007;204(2):640–657. doi:https://doi.org/10.1093/pcp/pch048.
- Kekwick RGO. Serum-cholinesterase activity in health and in liver disease. Biochem J. 1960;76(3):420–424. doi:https://doi.org/10.1016/j.plipres.2004.09.002.
- Talesa V, Principato GB, Giovannini E, Di Giovanni MV, Rosi G. Dimeric forms of cholinesterase in Sipunculus nudus. Eur J Biochem. 1993;215:267–275. doi:https://doi.org/10.1111/j.1432-1033.1993.tb18031.x.
- Mesarič T, Sepčić K, Drobne D, Makovec D, Faimali M, Morgana S, Falugi C, Gambardella C. Sperm exposure to carbon-based nanomaterials causes abnormalities in early development of purple sea urchin (Paracentrotus lividus). Aquat Toxicol. 2015;163:158–166. doi:https://doi.org/10.1016/j.aquatox.2015.04.012.
- Tezuka T, Akita I, Yoshino N, Suzuki Y. Regulation of self-incompatibility by acetylcholine and cAMP in Lilium longiflorum. J Plant Physiol. 2007;164:878–885. doi:https://doi.org/10.1016/j.jplph.2006.05.013.
- Rejón JD, Zienkiewicz A, Rodríguez-García MI, Castro AJ. Profiling and functional classification of esterases in olive (Olea europaea) pollen during germination. Ann Bot. 2012;110:1035–1045. doi:https://doi.org/10.1093/aob/mcs174.
- Sagane Y, Nakagawa T, Yamamoto K, Michikawa S, Oguri S, Momonoki YS. Molecular characterization of maize acetylcholinesterase: a novel enzyme family in the plant kingdom. Plant Physiol. 2005;138:1359–1371. doi:https://doi.org/10.1104/pp.105.062927.
- Yamamoto K, Oguri S, Momonoki YS. Characterization of trimeric acetylcholinesterase from a legume plant, Macroptilium atropurpureum Urb. Planta. 2008;227(4):809–822. doi:https://doi.org/10.1007/s00425-007-0658-0.
- Yamamoto K, Oguri S, Chiba S, Momonoki YS. Molecular cloning of acetylcholinesterase gene from Salicornia europaea L. Plant Signal Behav. 2009;4:361–366. doi:https://doi.org/10.4161/psb.4.5.8360.
- Yamamoto K, Shida S, Honda Y, Shono M, Miyake H, Oguri S, Sakamoto H, Momonoki YS. Overexpression of acetylcholinesterase gene in rice results in enhancement of shoot gravitropism. Biochem Biophys Res Commun. 2015;465:488–493. doi:https://doi.org/10.1016/j.bbrc.2015.08.044.
- Yamamoto K, Sakamoto H, Momonoki YS. Maize acetylcholinesterase is a positive regulator of heat tolerance in plants. J Plant Physiol. 2011;168:1987–1992. doi:https://doi.org/10.1016/j.jplph.2011.06.001.
- Yamamoto K, Sakamoto H, Momonoki YS. Altered expression of acetylcholinesterase gene in rice results in enhancement or suppression of shoot gravitropism. Plant Signal Behav. 2016;11:e1163464. doi:https://doi.org/10.1080/15592324.2016.1163464.
- Hadačová V, Vacková K, Klozová E, Kutáček M, Pitterová K. Cholinesterase activity in some species of theAllium genus. Biol Plant. 1983;25(3):209–215. doi:https://doi.org/10.1007/bf02902105.
- Roshchina VV. Neurotransmitters in plant life. Science Publishers Inc., Enfield, USA; 2001;99–150.
- Hiei Y, Ohta S, Komari T, Kumashiro T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 1994;6:271–282. doi:https://doi.org/10.1046/j.1365-313x.1994.6020271.x.
- Miki D, Simple SK. RNAi vectors for stable and transient suppression of gene function in rice. Plant Cell Physiol. 2004;45:490–495. doi:https://doi.org/10.1093/pcp/pch048.
- Miki D, Itoh R, Shimamoto K. RNA silencing of single and multiple members in a gene family of rice. Plant Physiol. 2005;138:1903–1913. doi:https://doi.org/10.1104/pp.105.063933.
- Upton C, Buckley JT. A new family of lipolytic enzymes? Trends Biochem Sci. 1995;20:178–179. doi:https://doi.org/10.1016/s0968-0004(00)89002-7.
- Brick DJ, Brumlik MJ, Buckley JT, Cao J-X, Davies PC, Misra S, Tranbarger TJ, Upton C. A new family of lipolytic plant enzymes with members in rice, arabidopsis and maize. FEBS Lett. 1995;377:475–480. doi:https://doi.org/10.1016/0014-5793(95)01405-5.
- Akoh CC, Lee G-C, Liaw Y-C, Huang T-H, Shaw J-F. GDSL family of serine esterases/lipases. Prog Lipid Res. 2004;43(6):534–552. doi:https://doi.org/10.1016/j.plipres.2004.09.002.
- Shimura K, Okada A, Okada K, Jikumaru Y, Ko K-W, Toyomasu T, Sassa T, Hasegawa M, Kodama O, Shibuya N, et al. Identification of a biosynthetic gene cluster in rice for momilactones. J Biol Chem. 2007;282(47):34013–34018. doi:https://doi.org/10.1074/jbc.m703344200.
- Grigor’eva GM, Rakic LM. Acetylcholinesterase in the brain of the marine gastropod Murex trunculus L. (Prosobranchia). Isolation and properties. Comparative Biochemistry and Physiology Part C: Comparative Pharmacology. 1979;64(2):199–206. doi:https://doi.org/10.1016/0306-4492(79)90049-2.
- Principato GB, Ambrosini MV, Liotti FS, Giovannini E. Propionylcholinesterase in Hirudo medicinalis: purification, partial characterization and comparative study with a mammalian acetylcholinesterase. Comparative Biochemistry and Physiology Part C: Comparative Pharmacology. 1981;70(2):209–213. doi:https://doi.org/10.1016/0306-4492(81)90053-8.
- Leung MR, Zeev-Ben-Mordehai T. Cryo-electron microscopy of cholinesterases, present and future. J Neurochem. 2020;00:1–8. doi:https://doi.org/10.1111/jnc.15245.
- Yamamoto K, Momonoki YS. Subcellular localization of overexpressed maize AChE gene in rice plant. Plant Signal Behav. 2008;3(8):576–577. doi:https://doi.org/10.1007/s00425-007-0658-0.
- Miura GA, Shih T-M. Identification of proprionylcholine in higher plants. Physiol Plant. 1984;62(3):341–343. doi:https://doi.org/10.1111/j.1399-3054.1984.tb04583.x.
- Wang Q, Gao P, Wang X, Duan Y. The early diagnosis and monitoring of squamous cell carcinoma via saliva metabolomics. Sci Rep. 2008;4:6802. doi:https://doi.org/10.1007/s00425-007-0658-0.
- Moreno S, Gerbig S, Schulz S, Spengler B, Diener M, Bader S. Epithelial propionyl- and butyrylcholine as novel regulators of colonic ion transport. Br J Pharmacol. 2016;173(18):2766–2779. doi:https://doi.org/10.4161/psb.4.5.8360.