ABSTRACT
Phylloquinone (vitamin K1) is a thylakoid-embedded electron carrier essential for photosynthesis. Paradoxically, we found that phylloquinone biosynthesis is retained in the nonphotosynthetic holoparasite Phelipanche aegyptiaca (Egyptian broomrape). The phylloquinone pathway genes are preferentially expressed during development of the invasive organ, the haustorium, and exhibit strong coexpression with redox-active proteins known to be involved in parasitism. Unlike in photoautotrophic taxa, the late pathway genes of the holoparasite lack the chloroplast-targeting sequence and their proteins are targeted to the plasma membrane instead. Plasma membrane phylloquinone may enable Phelipanche to sense changes in the redox environment during host interactions. The N-truncated isoforms are conserved in several other Orobanchaceae root holoparasites, and interestingly, in a number of closely related photoautotrophic species as well. This suggests an ancient origin of distinct phylloquinone pathways predating the evolution of parasitic plants in the Orobanchaceae. These findings represent exciting opportunities to probe plasma membrane phylloquinone function and diversification in parasitic and nonparasitic plant responses to external redox chemistry in the rhizosphere.
Holoparasitic plants have evolved a minimalist lifestyle and depend entirely on their hosts to survive and complete their life cycle. This evolutionary shift is marked by greatly reduced seed reserve size, by loss of photosynthetic machinery, and by possession of a specialized invasive organ called haustorium.Citation1,Citation2 The multicellular haustoria have functions ranging from host penetration to host vascular connection for nutrient and water acquisition.Citation3 While great strides have been made in understanding the sensing and perception of host-derived signals by parasitic seeds for germination and haustorium initiation, the redox machinery underpinning these processes remains poorly studied. We recently described a novel phylloquinone (PhQ, vitamin K1) biosynthetic pathway in Phelipanche aegyptiaca (Egyptian broomrape) and proposed its participation in plasma membrane (PM) redox signaling and haustorium development.Citation4
The discovery of the full complement of PhQ biosynthetic genes in a nonphotosynthetic holoparasite may seem counterintuitive at first, given the well-known role of PhQ as a redox cofactor in photosystem I electron transport and oxidative folding of photosystem II subunits.Citation5 However, our finding of an alternative, PM-targeted PhQ pathway in broomrapesCitation4 is consistent with the long suggested involvement of PhQ in the PM electron transport chain of heterotrophic plant tissues.Citation6–10 A PM-localized electron transport chain is also central to xenognosis of parasitic plants, the signaling organogenesis process that couples the vegetative-to-parasitic phase transition with host-derived chemical cues.Citation11 According to the electrochemical framework for xenognosis, host-derived haustorium-inducing factors (HIFs) are perceived by PM-bound electron carriers to initiate redox signaling essential for haustorium formation.Citation11 In Striga asiatica, haustorium signaling only commences within a narrow range of redox potentials and shares similar characteristics with organellar redox circuits such as photosynthesis and oxidative phosphorylation.Citation11,Citation12 This led Keyes and colleagues Citation11 to speculate on organellar involvement in the redox relay of haustorium signaling. Our finding that holoparasitic P. aegyptiaca repurposed PhQ from defunct photosynthetic electron transport for PM redox systems represents a parsimonious explanation of the model.Citation4
Gene network analysis revealed strong coexpression of P. aegyptiaca PhQ genes with components of the PM electron transport chain, including NAD(P)H quinone reductase (PaQR2) and apoplastic peroxidases (PaPRX1 and PaPRX2) known to be associated with parasitism. The apoplastic PRXs have established roles in reactive oxygen species (ROS) generation for haustorium development.Citation13,Citation14 PaPRXs are also among the most abundantly expressed genes in early stages of broomrape development, especially during haustorium penetration to host roots.Citation4,Citation15 QR2 expression is highly induced by the common HIF 2,6-dimethoxy1,4-benzoquinones (DMBQ) in several hemiparasites, including Triphysaria versicolor, S. asiatica, S. hermonthica, and Phtheirospermum japonicum,Citation16–18 and its silencing by RNAi reduced haustorium formation in Phtheirospermum japonicum.Citation16 Broomrapes do not respond to DMBQ,Citation19 but PaQR2 was also strongly induced upon exposure to host roots during haustorium initiation.Citation4 QR2 is evolutionarily conserved, with PM associationCitation20,Citation21 and menadione (vitamin K3) reduction activities that can be competitively inhibited by dicumarol, a vitamin K antagonist.Citation22 The data suggest that QR2 may have roles both as a penultimate enzyme in PhQ biosynthesis, and in PM redox cycling and ROS detoxification.Citation4
The haustorium induction chemistry of DMBQ-insensitive broomrapes is poorly understood.Citation23 Cytokinins have been shown to induce haustorium formation, and cytokinin perception was positively correlated with parasite aggressiveness in P. ramose.Citation24 Two cytokinin marker genes PrCKX2 and PrCKX4 encoding cytokinin oxidases were specifically induced upon exposure to cytokinins or host root exudates during haustorium initiation.Citation24 We identified a single ortholog PaCKX2 in the PhQ-centered gene network for P. aegyptiaca.Citation4 PaCKX2 exhibited strong coexpression with four of the seven PhQ pathway genes, whereas PhQ gene coexpression was much weaker for CKX2/4 orthologs in DMBQ-responsive hemiparasites S. hermonthica (three of nine PhQ genes) and T. versicolor (one of ten PhQ genes). The patterns are consistent with what we observed for parasitism genes and redox genes involved in transmembrane electron transport.Citation4 PaCKX2 thus provides another line of support for the coupling of PhQ biosynthesis, haustorium signaling, and PM redox systems in P. aegyptiaca.
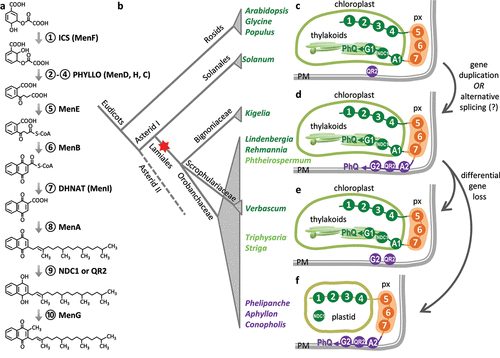
The PhQ biosynthetic pathway is compartmentalized in different organelles (). The early (steps 1–4) and intermediate (steps 5–7) pathway steps occur in chloroplasts and peroxisomes, respectively, with the late steps determining the ultimate targeting of PhQ. The chloroplast envelope, plastoglobule, and thylakoid membrane localization of the late pathway enzymes MenA1, NDC1 and MenG1, respectively,Citation25–27 completes the canonical PhQ biosynthesis in chloroplasts in support of its photosynthetic role (). The alternative pathway configuration of P. aegyptiaca PhQ biosynthesis is mediated by PM enzymes, PaMenA2, PaQR2, and PaMenG2Citation4 (). PaMenA2 and PaMenG2 have shorter N-termini lacking the chloroplast transit signal when compared to their plastidial counterparts. A search of the One Thousand Plant Transcriptomes (1KP) databaseCitation28 revealed two other Orobanchaceae holoparasites, Aphyllon (syn. Orobanche) fasciculata and Conopholis americana which harbor the alternative MenA2 and MenG2 but not the canonical MenA1 and MenG1 (). We also identified distinct genes encoding putative plastidial (MenA1/G1) and PM (MenA2/G2) isoforms in the draft genome assembly of hemiparasite Phtheirospermum japonicumCitation29 and in the 1KP transcriptomes of nonparasitic Lindenbergia philippensis, Rehmannia glutinosa, and Kigelia africana().Citation4 Additional parasitic and nonparasitic taxa – all belonging to Lamiales – contain both MenG1 and MenG2 (T. versicolor, S. hermonthica, and Verbascum thapsus) (, e). These data are consistent with the notion of a PM PhQ pathway that predates the origin of the parasitic plants (). Gene duplication of MenA and MenG in the common ancestor of Lamiales likely gave rise to the two distinct isoforms, followed by differential gene losses in photosynthetic and nonphotosynthetic lineagesCitation4 ().
Figure 1. Evolution of phylloquinone biosynthetic pathways
The taxonomically restricted occurrence of MenA2 and MenG2 raises questions about the biosynthetic origin of PhQ previously associated with PM electron transport in heterotrophic tissues of non-Lamiales.Citation6,Citation9,Citation30,Citation31 We found evidence of alternative splicing for both AtMenA and AtMenG in Arabidopsis thaliana,Citation32 with several isoforms predicted to lack the plastid-targeting signal. We suggest that alternative splicing can afford flexible PhQ biosynthesis for PM redox functions in species without MenA2 and MenG2. One such example can be inferred from recent studies with a group of PM-localized leucine-rich-repeat receptor kinases (Hydrogen Peroxide-induced Ca2+ Increases [HPCA] or Cannot Respond to DMBQ [CARD]) that function as sensors for ROS as well as DMBQ to induce Ca2+ influx in roots.Citation33,Citation34 This ROS/quinone signaling pathway is involved in pathogen defense of A. thaliana and haustorium formation of Phtheirospermum japonicum, and depends on redox activation of CARD via disulfide bond formation between two cysteine residues in the extracellular sensing domain.Citation33,Citation34 AtCARD1 exhibits higher substrate affinities for menadione and 1,4-naphthoquinone than DMBQ, and its mutation can be rescued by Phtheirospermum japonicum and S. asiatica CARD orthologs.Citation33 The work supports a conserved ROS/quinone signaling mechanism predating the origin of parasitism. We suggest that PM-localized PhQ may function as a redox cofactor in disulfide bond formation of CARDs, like plastidial PhQ involvement in disulfide bond formation of photosystem II subunits.Citation4
Besides pathogen defense, PhQ may participate in PM-mediated ROS sensing, signaling and processing in response to environmental stimuli, such as ozone, heavy metals, and water and temperature stresses.Citation35 Supporting evidence can indeed be gleaned from the PhQ-centered gene network modeling of parasitic plants.Citation4 In eudicot roots, ferric reductase oxidases (FRO) of the flavocytochrome b family mediate electron transfer across the PM associated with iron and copper reduction for uptake from soils.Citation36,Citation37 The PhQ-coexpressed PaFRO1 is orthologous to A. thaliana AtFRO4/AtFRO5 tandem duplicates that are essential for Cu uptake in root tips.Citation36 The A. thaliana QR2 (AtFQR1) exhibits root-biased expression and is highly responsive to auxin and allelochemicals.Citation38,Citation39 These data point to root PM redox systems associated with biotic and abiotic interactions as potential avenues for research into PM PhQ function in nonparasitic plants.
In summary, adaptation of an extant PhQ pathway for PM redox signaling associated with haustorium development adds to documented evolutionary innovation of parasitic plants. Our work Citation4offers a repertoire of candidate genes for hypothesis-driven research to ascertain their roles in transmembrane redox processes associated with parasitism. A better understanding of haustorium signaling is of significance to both basic and applied research for effective control of noxious weeds that cause devastating crop losses around the world. Conservation of the alternative PhQ biosynthetic pathway in nonparasitic angiosperms should motivate future research into the PM function of PhQ in photoautotrophic species.
Acknowledgments
THE authors thank James Westwood and colleagues for parasitic plant transcriptome resources funded by National Science Foundation (OS-1238057), other collaborators who contributed to the original study, and Scott Harding for helpful discussion.
Disclosure statement
No potential conflicts of interest were disclosed.
Additional information
Funding
References
- Joel DM, Bar H, Mayer AM, Plakhine D, Ziadne H, Westwood JH, Welbaum GE. Seed ultrastructure and water absorption pathway of the root-parasitic plant Phelipanche aegyptiaca (Orobanchaceae). Ann Bot. 2012;109(1):1–4. doi:https://doi.org/10.1093/aob/mcr261.
- Westwood JH, Yoder JI, Timko MP, dePamphilis CW. The evolution of parasitism in plants. Trends Plant Sci. 2010;15(4):227–235. doi:https://doi.org/10.1016/j.tplants.2010.01.004.
- Yoshida S, Cui S, Ichihashi Y, Shirasu K. The haustorium, a specialized invasive organ in parasitic plants. Annu Rev Plant Biol. 2016;67(1):643–667. doi:https://doi.org/10.1146/annurev-arplant-043015-111702.
- Gu X, Chen I-G, Harding SA, Nyamdari B, Ortega MA, Clermont K, Westwood JH, Tsai C-J. Plasma membrane phylloquinone biosynthesis in nonphotosynthetic parasitic plants. Plant Physiol. 2021;185(4):1443–1456. doi:https://doi.org/10.1093/plphys/kiab031.
- Basset GJ, Latimer S, Fatihi A, Soubeyrand E, Block A. Phylloquinone (Vitamin K1): occurrence, biosynthesis and functions. Mini Rev Med Chem. 2017;17(12):1028–1038. doi:https://doi.org/10.2174/1389557516666160623082714.
- Biniek C, Heyno E, Kruk J, Sparla F, Trost P, Krieger-Liszkay A. Role of the NAD(P)H quinone oxidoreductase NQR and the cytochrome b AIR12 in controlling superoxide generation at the plasma membrane. Planta. 2017;245(4):807–817. doi:https://doi.org/10.1007/s00425-016-2643-y.
- Döring O, Lüthje S, Böttger M. Modification of the activity of the plasma membrane redox system of zea mays L. Roots by Vitamin K3 and dicumarol. J Exp Bot. 1992;43(2):175–181. doi:https://doi.org/10.1093/jxb/43.2.175.
- Lochner K, Döring O, Böttger M. Phylloquinone, what can we learn from plants? BioFactors. 2003;18(1–4):73–78. doi:https://doi.org/10.1002/biof.5520180209.
- Lüthje S, Van Gestelen P, Córdoba-Pedregosa MC, González-Reyes JA, Asard H, Villalba JM, Böttger M. Quinones in plant plasma membranes — a missing link? Protoplasma. 1998;205(1–4):43–51. doi:https://doi.org/10.1007/BF01279292.
- Schopfer P, Heyno E, Drepper F, Krieger-Liszkay A. Naphthoquinone-dependent generation of superoxide radicals by quinone reductase isolated from the plasma membrane of soybean. Plant Physiol. 2008;147(2):864–878. doi:https://doi.org/10.1104/pp.108.118745.
- Keyes JW, O’Malley CR, Kim D, Lynn GD. Signaling organogenesis in parasitic angiosperms: xenognosin generation, perception, and response. J Plant Growth Regul. 2000;19(2):217–231. doi:https://doi.org/10.1007/s003440000024.
- Smith CE, Ruttledge T, Zeng Z, O’Malley RC, Lynn DG (1996). A mechanism for inducing plant development: the genesis of a specific inhibitor. Proceedings of the National Academy of Sciences 93, 6986–6991. doi:https://doi.org/10.1073/pnas.93.14.6986
- González-Verdejo CI, Barandiaran X, Moreno MT, Cubero JI, Di Pietro A. A peroxidase gene expressed during early developmental stages of the parasitic plant Orobanche ramosa. J Exp Bot. 2006;57(1):185–192. doi:https://doi.org/10.1093/jxb/erj024.
- Kim D, Kocz R, Boone L, John Keyes W, Lynn DG. On becoming a parasite: evaluating the role of wall oxidases in parasitic plant development. Chem Biol. 1998;5(2):103–117. doi:https://doi.org/10.1016/S1074-5521(98)90144-2.
- Yang Z, Wafula EK, Honaas LA, Zhang H, Das M, Fernandez-Aparicio M, Huang K, Bandaranayake PCG, Wu B, Der JP, et al. Comparative transcriptome analyses reveal core parasitism genes and suggest gene duplication and repurposing as sources of structural novelty. Mol Biol Evol. 2015;32(3):767–790. doi:https://doi.org/10.1093/molbev/msu343.
- Ishida JK, Yoshida S, Shirasu K. Quinone oxidoreductase 2 is involved in haustorium development of the parasitic plant Phtheirospermum japonicum. Plant Signal Behav. 2017;12(7):e1319029–e1319029. doi:https://doi.org/10.1080/15592324.2017.1319029.
- Liang L, Liu Y, Jariwala J, Lynn DG, Palmer AG. Detection and adaptation in parasitic angiosperm host selection. Am J Plant Sci. 2016;7(8):1275–1290. doi:https://doi.org/10.4236/ajps.2016.78123.
- Matvienko M, Wojtowicz A, Wrobel R, Jamison D, Goldwasser Y, Yoder JI. Quinone oxidoreductase message levels are differentially regulated in parasitic and non-parasitic plants exposed to allelopathic quinones. The Plant Journal. 2001;25(4):375–387. doi:https://doi.org/10.1046/j.1365-313x.2001.00971.x.
- Westwood JH. Characterization of the Orobanche –arabidopsis system for studying parasite–host interactions. Weed Sci. 2000;48(6):742–748. doi:https://doi.org/10.1614/0043-1745(2000)048[0742:COTOAS]2.0.CO;2.
- Marmagne A, Rouet M-A, Ferro M, Rolland N, Alcon C, Joyard J, Garin J, Barbier-Brygoo H, Ephritikhine G. Identification of new intrinsic proteins in Arabidopsis plasma membrane proteome. Mol Cell Proteomi. 2004;3(7):675–691. doi:https://doi.org/10.1074/mcp.M400001-MCP200.
- Natera SHA, Ford KL, Cassin AM, Patterson JH, Newbigin EJ, Bacic A. Analysis of the Oryza sativa plasma membrane proteome using combined protein and peptide fractionation approaches in conjunction with mass spectrometry. J Proteome Res. 2008;7(3):1159–1187. doi:https://doi.org/10.1021/pr070255c.
- Wrobel RL, Matvienko M, Yoder JI. Heterologous expression and biochemical characterization of an NAD(P)H: quinoneoxidoreductase from the hemiparasitic plant Triphysaria versicolor. Plant Physiol Biochem. 2002;40(3):265–272. doi:https://doi.org/10.1016/S0981-9428(02)01361-X.
- Fernández-Aparicio M, Reboud X, Gibot-Leclerc S. Broomrape weeds underground mechanisms of parasitism and associated strategies for their control: a review . Frontiers Plant Sci. 2016;7:135. doi:https://doi.org/10.3389/fpls.2016.00135
- Goyet V, Billard E, Pouvreau J-B, Lechat -M-M, Pelletier S, Bahut M, Monteau F, Spíchal L, Delavault P, Montiel G, et al. Haustorium initiation in the obligate parasitic plant Phelipanche ramosa involves a host-exudated cytokinin signal. J Exp Bot. 2017;68(20):5539–5552. doi:https://doi.org/10.1093/jxb/erx359.
- Eugeni Piller L, Besagni C, Ksas B, Rumeau D, Bréhélin C, Glauser G, Kessler F, Havaux M (2011). Chloroplast lipid droplet type II NAD(P)H quinone oxidoreductase is essential for prenylquinone metabolism and vitamin K1 accumulation. Proceedings of the National Academy of Sciences 108, 14354–14359. doi:https://doi.org/10.1073/pnas.1104790108
- Kaiping S, Soll J, Schultz G. Site of methylation of 2-phytyl-1,4-naphthoquinol in phylloquinone (vitamin K1) synthesis in spinach chloroplasts. Phytochemistry. 1984;23(1):89–91. doi:https://doi.org/10.1016/0031-9422(84)83083-6.
- Schultz G, Soll J, Ellerbrock BH. Site of prenylation reaction in synthesis of phylloquinone (Vitamin K1) by spinach chloroplasts. Europ J Biochem. 1981;117(2):329–332. doi:https://doi.org/10.1111/j.1432-1033.1981.tb06341.x.
- Leebens-Mack JH, Barker MS, Carpenter EJ, Deyholos MK, Gitzendanner MA, Graham SW, Grosse I, Li Z, Melkonian M, Mirarab S, et al., One Thousand Plant Transcriptomes, I. One thousand plant transcriptomes and the phylogenomics of green plants. Nature. 2019;574:679–685. doi:https://doi.org/10.1038/s41586-019-1693-2
- Cui S, Kubota T, Nishiyama T, Ishida JK, Shigenobu S, Shibata TF, Toyoda A, Hasebe M, Shirasu K, Yoshida S. Ethylene signaling mediates host invasion by parasitic plants. Sci Adv. 2020;6(44):eabc2385. doi:https://doi.org/10.1126/sciadv.abc2385.
- Barr R, Pan RS, Crane FL, Brightman AO, Morré DJ. Destruction of vitamin K1 of cultured carrot cells by ultraviolet radiation and its effect on plasma membrane electron transport reactions. Biochem Int. 1992;27:449–456.
- Bridge A, Barr R, Morré DJ. The plasma membrane NADH oxidase of soybean has vitamin K1 hydroquinone oxidase activity. Biochimica Et Biophysica Acta (BBA) - Biomembranes. 2000;1463(2):448–458. doi:https://doi.org/10.1016/S0005-2736(99)00239-4.
- Li S, Yamada M, Han X, Ohler U, Benfey PN. High-resolution expression map of the Arabidopsis root reveals alternative splicing and lincRNA regulation. Dev Cell. 2016;39(4):508–522. doi:https://doi.org/10.1016/j.devcel.2016.10.012.
- Laohavisit A, Wakatake T, Ishihama N, Mulvey H, Takizawa K, Suzuki T, Shirasu K. Quinone perception in plants via leucine-rich-repeat receptor-like kinases. Nature. 2020; 587:92-97. doi:https://doi.org/10.1038/s41586-020-2655-4.
- Wu F, Chi Y, Jiang Z, Xu Y, Xie L, Huang F, Wan D, Ni J, Yuan F, Wu X, et al. Hydrogen peroxide sensor HPCA1 is an LRR receptor kinase in Arabidopsis. Nature. 2020;578(7796):577–581. doi:https://doi.org/10.1038/s41586-020-2032-3.
- Podgórska A, Burian M, Szal B. Extra-cellular but extra-ordinarily important for cells: apoplastic reactive oxygen species metabolism. Front Plant Sci. 2017;8. doi:https://doi.org/10.3389/fpls.2017.01353.
- Bernal M, Casero D, Singh V, Wilson GT, Grande A, Yang H, Dodani SC, Pellegrini M, Huijser P, Connolly EL, et al. Transcriptome sequencing identifies SPL7 -regulated copper acquisition genes FRO4/FRO5 and the copper dependence of iron homeostasis in arabidopsis. Plant Cell. 2012;24(2):738–761. doi:https://doi.org/10.1105/tpc.111.090431.
- Robinson NJ, Procter CM, Connolly EL, Guerinot ML. A ferric-chelate reductase for iron uptake from soils. Nature. 1999;397(6721):694. doi:https://doi.org/10.1038/17800.
- Baerson SR, Sánchez-Moreiras A, Pedrol-Bonjoch N, Schulz M, Kagan IA, Agarwal AK, Reigosa MJ, Duke SO. Detoxification and transcriptome response in Arabidopsis seedlings exposed to the allelochemical benzoxazolin-2(3H)-one. J Biol Chem. 2005;280(23):21867–21881. doi:https://doi.org/10.1074/jbc.M500694200.
- Laskowski MJ, Dreher KA, Gehring MA, Abel S, Gensler AL, Sussex IM. FQR1, a novel primary auxin-response gene, encodes a flavin mononucleotide-binding quinone reductase. Plant Physiol. 2002;128(2):578–590. doi:https://doi.org/10.1104/pp.010581.
