ABSTRACT
Objectives
The outcome of elderly acute myeloid leukemia (AML) patients is poor, which was traditionally attributed to patient- and leukemia-related factors. However, studies about the genetic features of these elderly patients have not been integrated and the genetic mechanism of their poor outcome is less known.
Methods
Here, we used next generation sequencing (NGS) to identify the genetic features of elderly AML patients and confirmed the efficacy of chemotherapy based on molecular aberrations. Mutations in 111 genes relevant to hematological malignancy was analysed by virtue of NGS and the genetic differences were compared between elderly (n=52) and young (n=161) AML patients. Furthermore, the outcome of decitabine-based chemotherapy was identified in elderly patients.
Results
Frequencies of adverse genetic alterations, such as RUNX1 and secondary-type mutations (ASXL1, STAG2 and spliceosome), were much higher in elderly patients, while the frequency of WT1 mutations was much lower. Moreover, epigenetic mutations such as DNMT3A, TET2, ASXL1 and IDH2, were also more common in elderly patients. Furthermore, there were 39 elderly patients receiving the decitabine-based chemotherapy, and the results showed that the overall response rate (ORR) and complete remission rate (CR) were 76.9% and 71.8%, respectively. The median overall survival (OS) for those older patients was 12 months, and the 2-year OS probability was 20.5%.
Discussion
Our study provides deep understanding into the molecular mechanisms of the poor outcome of elderly AML patients.
Conclusion
Epigenetic mutations play an important role, and decitabine-based regimen can be used as alternative first-line chemotherapy for elderly patients.
Introduction
AML is characterized by clinical and biological heterogeneity, with the incidence and the prognosis both directly related to advancing age [Citation1–3]. Although clinical outcomes have improved in younger patients over the decades, they remain very poor in older patients. A series of patient- and leukemia-related factors were used to explain their poor outcomes, including poor physical condition, high intensity of chemotherapy, accompanying comorbidities such as severe infections, a higher incidence of unfavorable karyotypes, multidrug resistance protein 1 (MDR1) expression and more [Citation4–7].
In recent years, more and more gene mutations have been gradually discovered in AML, and some mutations have been shown to have prognostic significance, including FLT3, NPM1, CEBPA, KIT, RUNX1, ASXL1, TP53, DNMT3A, TET2, IDH1/2, and WT1 [Citation8–11]. Among them, epigenetic mutations mainly including DNMT3A, ASXL1and TET2 mutations have been identified to be associated with poor prognosis [Citation12–14]. Decitabine, a hypomethylating agent, has been approved for treating AML patients. Most studies have been identified to predict favorable responses [Citation15–20], while others reported their efficacy in AML was controversial [Citation21,Citation22]. However, recent studies were limited to cohorts receiving hypomethylating agents alone or in combination with drugs without anthracyclines. There is a lack of prognostic and predictive biomarkers for elderly AML patients, and whether they will benefit from decitabine plus aclarubicin, cytarabine and G-CSF (DCAG) is unclear.
In this study, we aimed at exploring the genetic features of AML in elderly patients and further identify their clinical outcome with DCAG regimen, which may provide guidance for future therapy.
Methods
Subjects
In total, we analyzed 213 newly diagnosed de novo AML patients which did not include M3 and 79 healthy adults from September 2008 to September 2015, and we took the 2016 WHO classification as reference for AML diagnosis and classification [Citation23]. Among them, there were 52 patients aged 60 years or older, and 161 patients younger than 60 years old. Patients with antecedent hematological diseases, history of myeloid malignancy, or treatment-related AML were excluded. Meanwhile, we collected a total of 79 peripheral blood or saliva samples from healthy adults as the background control. Our study was approved by the Human Ethics Committees of the Chinese PLA General Hospital, and all patients provided written informed consent before they received chemotherapy.
Cytogenetic and mutational analysis
Bone marrow cells were either harvested directly or after 1–3 days of unstimulated culture. Metaphase chromosomes were banded using the trypsin-Giemsa technique and karyotyped according to the International System for Human Cytogenetic Nomenclature. And we used standard methods to extract genomic DNA from BM samples [Citation24].
Analyses of mutations in all 111 known or putative genes relevant to hematological malignancies were performed in all 213 AML and 79 healthy samples. We used 250 K NimbelGen SeqCap EZ Choice kit to trap sequences and enrich libraries according to protocol. Hybrid libraries mixed with probes were run on an Illumina HiSeq 2500 to obtain 100-bp paired-end sequences.
The genomic content of the sequencing panel was 250 K, which containing the entire coding sequence of 111 genes that have mutations associated with AML pathogenesis. Among these, 42 genes originated from the hematopoietic diagnosis and treatment guidelines published by the NCCN (National Comprehensive Cancer Network) and WHO (World Health Organization), and 69 genes were chosen according to recent literature (Supplementary Table S1).
Clinical response
Therapeutic response was defined by the IWG criteria. Patients achieved CR with less than 5% bone marrow (BM) blasts, an absolute neutrophil count ≥1.0×109/L and a platelet count >100 ×109/L. Partial remission (PR) was assigned to those with a decrease of at least 50% (5–25%) of total bone marrow blasts [Citation25]. Those achieving neither CR nor PR were considered patients with non-remission (NR). Relapse was defined as the recurrence of leukemia cells in peripheral blood or the reappearance of >5% blasts in the BM.
Statistical analysis
Frequencies were evaluated using Fisher’s exact test or a chi-square test. Survival analysis was performed by the Kaplan-Meier method. OS was measured from diagnosis time to the date of death from any cause or to the last follow-up. Disease-free survival (DFS) was defined from the date of CR to the time of relapse, death from any cause or the last follow-up. A p-value<0.05 was considered statistically significant.
Results
Patient characteristics
Clinical characteristics of the 213 patients are summarized in . There were 128 males and 85 females. Fifty-two patients were elderly and their median age was 68 years (range 60–88 years). Of the fifty-two older patients, eight patients only received palliative care or even did not receive any chemotherapy owning to their poor performance status, five patients received IA/DA/MA (idarubicin or doxorubicin or mitoxantrone for 3 days and cytarabine for 7 days) chemotherapy, and the other thirty-nine patients received DCAG (decitabine 10 mg/m2 d1-5, aclarubicin 20 mg d1,3,5, cytarabine 10 mg/m2 q12 h d1-5, G-CSF 300μg/day from day 0 to neutrophil recovery) as an induction therapy regimen with a median follow-up time of 27.6 months (interquartile range, 21.4–35.2 m). According to our clinical trial design, if WBC≥10*10^9/L, hydroxyurea or cytarabine is recommended to reduce white blood cells. If WBC<10*10^9/L, G-CSF used from day 0. After achieving CR or PR with two cycles, twenty-seven of these thirty-nine patients continued to accept DCAG as consolidation chemotherapy until relapse, disease progression, unacceptable toxicity, the patients’ attending physicians requested other alternatives, or the patients gave up further treatment. And they received 3 cycles of DCAG consolidation on average (1–6 cycles). Those who did not acquire remission after two cycles of DCAG were offered alternative therapies, or they gave up further treatment.
Table 1. Clinical characteristics of 213 AML patients.
Mutational spectrum and frequencies of genetic alterations between older and younger patients
In the target genomic region, 30,667 SNPs (single-nucleotide polymorphism) and 7355 INDELs (insertion-deletions) which were absent in the 79 healthy background libraries were detected in the 213 AML patients. We ignored mutations that occurred in intronic non-coding regions or the untranslated regions (UTR) of mRNA genes. Several criteria were used to eliminate the influence of germline and harmless mutations on our results (Supplementary Methods). By filtering, approximately 2.79 mutations remained for each sample, and 68 genes of the 111 were identified. Details of those mutated genes are listed in Supplementary Table S2.
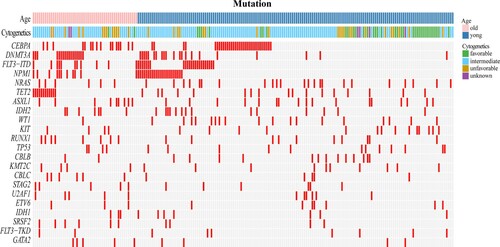
The mutational spectrum of the 213 AML patients was shown in . The most common genetic alteration in the total cohort was in CEBPA (23.0%), followed by mutations in DNMT3A (17.8%), FLT3-ITD (16%), NPM1 (16%), NRAS (13.6%), TET2 (13.6%) and ASXL1 (12.2%). From this figure, we can see that the two groups had some differences in the distribution of mutated genes, karyotypes, and more.
Comparison of cytogenetic and genetic alterations between older and younger patients
Among these patients, 207 patients had chromosome data, including 50 older patients and 157 younger patients (). From the statistical analysis, we can see that the older patients more frequently had unfavorable karyotypes (26.0% vs 13.4%) when compared with younger patients, including complex chromosomal abnormalities (12% vs 3.2%) and −7/del(7q-) (10% vs 2.5%). Older patients also less commonly had favorable karyotypes (4.0% vs 22.9%), such as t (8; 21) (2.0% vs 19.7%) based on MRC classification.
Table 2. Comparison of cytogenetic alterations between older and younger patients.
The distribution of genetic aberrations in the two age groups was also distinct (). In the older group, DNMT3A, CEBPA and FLT3 were the three most commonly mutations (32.7%, 30.80% and 23.1% respectively), followed by mutations in TET2 (23.1%), NPM1 (19.2%), ASXL1 (17.3%), RUNX1 (15.4%) and IDH2 (13.5%). In the younger group, CEBPA, FLT3 and NPM1 were the three most commonly identified mutations (20.5%, 19.3% and 14.9% respectively), followed by mutations in NRAS (14.3%), DNMT3A (13.0%), WT1 (11.2%), TET2 (10.6%) and ASXL1 (10.6%). Overall, compared with younger patients, epigenetic mutations such as DNMT3A, TET2 had much higher frequencies in older patients (32.7% vs 13.0%; 23.1% vs 10.6%, respectively). Likewise, ASXL1 and IDH2 also had higher frequencies than younger patients (17.3% vs 10.6%; 13.5% vs 8.7%, respectively). Furthermore, adverse genetic alterations such as RUNX1 and spliceosome mutations (including SRSF2, SF3B1 and U2AF1) were also more common in older patients than in the younger group (15.4% vs 6.2%; 23.1% vs 7.5%, respectively). The frequency of WT1 mutations was also much lower in older patients (1.9% vs 11.2%) (). The pairwise co-occurrence of mutations in the two age groups was also distinct (). Moreover, secondary-type mutations that predicted poor outcomes [Citation26,Citation27], including ASXL1, STAG2 and spliceosome (SRSF2, SF3B1, U2AF1), were also analyzed in these two age groups. In the 52 elderly AML patients, 17 (32.7%) had secondary-type mutations, while only 28 (17.4%) of the younger AML patients had these mutations (P=0.019).
Figure 2. The Circos plots depicted the relative frequency and pairwise co-occurrence of gene mutations in the older (a) and younger AML patients. (b) The length of the arc corresponds to the frequency of the first gene mutations, and the width of the ribbon corresponds to the percentage of patients who had mutations in the second gene.
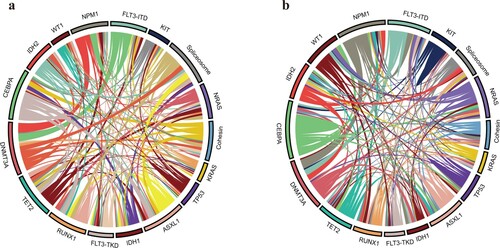
Table 3. Comparison of genetic alterations between older and younger patients.
Treatment outcomes of decitabine-based chemotherapy in older patients
There were 39 older patients receiving DCAG chemotherapy, and 30 patients had an overall response rate (ORR) of 76.9% after the first cycle. Among these patients, 23 (59%) achieved CR, and 7 (17.9%) had PR. After the second cycle, 4 patients with PR acquired CR. And 1 patient with PR acquired CR after the third cycle. Patients with favorable karyotypes all achieved CR (2/2, 100%); 75% (18/24) of patients with intermediate karyotypes and 81.3% (13/16) of patients with normal karyotypes also achieved CR, but 61.5% (8/13) of patients with unfavorable karyotypes achieved CR. In contrast, 7 patients with intermediate karyotypes received standard chemotherapy or palliative treatment, but only 1 achieved CR (14.3%), and the other 6 patients were not in remission (Supplementary Table S3, Figure 4a). The CR rate of those who received standard chemotherapy was 20% (1/5). Nevertheless, among the 27 CR patients receiving DCAG chemotherapy, there was recurrence in 8 cases (29.6%). Their genetic alterations are listed in a. Furthermore, 9 patients receiving DCAG induction therapy achieved NR, and their cytogenetic and molecular characteristics were listed in b. Among these, 3 patients had the mutational co-occurrence of NPM1mut/ FLT3-ITDmut/ DNMT3Amut. As for survival analysis, the 1- and 2-year OS probability was 43.6% and 20.5%, respectively (b).
Figure 3. Genetic and cytogenetic characteristics of older patients receiving DCAG with recurrence (a) and achieving NR (b).
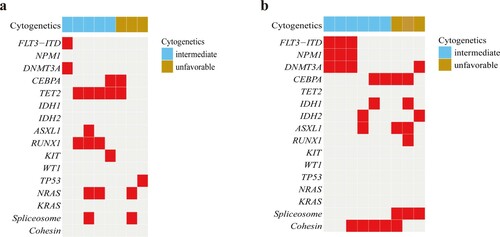
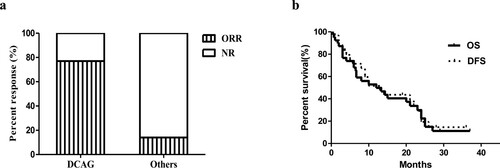
Figure 4. (a) Response rates for two groups of patients. Percent response is noted by each bar graph. ORR is noted by vertical bar, and additional patients with NR are noted by horizontal line. The response rates for patients receiving DCAG and patients receiving standard chemotherapy or palliative treatment were shown respectively. (b) Overall Survival (the solid line) and Disease-Free Survival (the dotted line) in older patients receiving DCAG. Vertical marks reflect the censored data.
Treatment-related toxicity
Overall, the DCAG chemotherapy regimen was well tolerated. The most common adverse event observed was myelosuppression, including thrombocytopenia and neutropenia. Of these, febrile neutropenia occurred in 71.8% (28/39) of patients, but non-hematological toxicities were rare. The median time for granulocyte and platelet recovery was 12 and 16 days respectively in patients achieving CR.
Twenty-nine of 39 elderly patients receiving DCAG chemotherapy died during our study. Of these, nine patients died because of disease relapse (23.1%), seven died from refractory disease (17.9%), three patients gave up treatment (7.7%) after 1 cycle of chemotherapy and all of them died of disease progression, five died of severe infections (12.8%), two died of heart failure (5.1%), one died of gastrointestinal hemorrhage (2.6%) and two died from multiple reasons (5.1%).
Discussion
AML is the most common adult acute leukemia, and its incidence increases with age. In the last two decades, advances in the treatment of AML have greatly improved the prognosis of younger patients. However, there has been little progress for older AML patients [Citation28–30]. Elderly AML patients are poorly tolerant of intensive chemotherapy because of treatment-related toxicities and complications. Approximately 40–55% older patients receiving intensive chemotherapy achieved CR, and the early death rate was up to 19–29% [Citation31,Citation32]. Meanwhile, their 2- and 5-year OS were only 10% and 2%, respectively [Citation33,Citation34]. Despite their significantly worse prognosis, studies on the treatment and outcomes of elderly AML patients showed that the prognosis of intensive treatment was better than those who receive supportive care and palliative chemotherapy or no anti-leukemia treatment [Citation35]. In addition, studies showed that low-dose cytarabine only had a survival benefit for patients with favorable and intermediate risk karyotypes. Moreover, due to the aggressive nature of AML, the mortality of patients with low intensity of chemotherapy can be as high as 26% [Citation36].
The outcome of elderly AML patients is still poor, and an optimal approach has not been established. It thus remains a great challenge to choose treatment options. In addition to the reasons mentioned above, could there be some other factors associated with a poor outcome? We conducted this study to explore the genetic differences between two age groups and in order to further guide clinical decisions.
In our study, we found that unfavorable karyotypes, such as complex chromosomal abnormalities and monosomy 7/7q deletion, were more common in elderly patients, and favorable karyotypes were less common, which agreed with previous studies [Citation37,Citation38]. In addition, we conducted a mutational analysis on 111 genes of clinical value in the two groups by next-generation sequencing. We found that adverse genetic alterations, such as RUNX1 and secondary-type mutations (ASXL1, STAG2 and spliceosome mutations), were more common in older patients. However, the frequency of WT1 mutations was much lower. These further explain the poor prognosis of elderly patients from the molecular mechanism perspective.
Moreover, we found the frequencies of epigenetic mutations such as DNMT3A, TET2, ASXL1 and IDH2 were also higher in older patients. The literature has reported that hypomethylating drugs such as DNMT inhibitors were more effective in AML patients with DNMT3A or TET2 or IDH1/2 mutations. Aberrant DNA methylation is frequently observed in AML, especially among patients with adverse risk features [Citation39,Citation40]. Hypomethylating drugs such as decitabine are pyrimidine nucleoside analogs that can result in the hypomethylation of DNA and restoration of expression of tumor-suppressor genes by inhibiting DNA methyltransferases (DNMTs) [Citation41,Citation42]. They were traditionally used in the treatment of MDS, and recently, decitabine has been identified to have anti-leukemia activity [Citation43,Citation44]. However, the CR rate was low for patients received decitabine alone [Citation45,Citation46]. Recent clinical studies have shown that decitabine can increase sensitivity of the leukemia cells to cytotoxic agents and increase the efficacy of AML induction chemotherapy [Citation47–49]. A low dose of decitabine (15–20 mg/m2/day) has been used as induction therapy for older AML patients and has been shown to have a better remission rate [Citation44,Citation50–52].
Based on these findings, we can sum up our own data regarding the efficacy of decitabine-based chemotherapy in older AML patients. In our study, the ORR of DCAG chemotherapy was 76.9%, and 69.2% of patients achieved CR, which showed obvious advantage when compared with standard chemotherapy or palliative treatment. Most patients in our study who were effective with DCAG chemotherapy regimen contained epigenetic modifier gene mutations like IDH1/IDH2, ASXL1, TET2 or DNMT3A mutations, and this was consistent with our team’s previous research [Citation53]. However, due to the limited sample, there were no statistically genetic subgroups that could be most effective with DCAG regimen for these elderly patients.
As we reported in a previous study [Citation54], those with co-occurrence of NPM1mut/ FLT3-ITDmut/ DNMT3Amut always had adverse outcomes. In our current study, of the 9 patients who achieved NR, 3 had the genotype NPM1mut/ FLT3-ITDmut/ DNMT3Amut. We also found that in patients with this adverse genotype, the efficacy of the DCAG regimen was unsatisfactory. For these patients, we suggest trying a combination of targeted therapy such as an FLT3 inhibitor. There have been studies showing that the addition of sorafenib to chemotherapy improved OS and DFS in older AML patients with FLT3-ITD mutations [Citation55], and when combined with hypomethylating drugs, sorafenib showed promising results [Citation56].
For older AML patients, there was no consensus on treatment options after achieving CR [Citation57,Citation58]. In our study, we found that the DCAG regimen may not be able to clear relapse clones, as 8 out of the 27 patients who achieved CR after DCAG induction therapy relapsed. Among these 8 recurrence patients, 3 had unfavorable-risk karyotypes and 5 had intermediate-risk karyotypes. All of them also had one or more adverse prognostic mutations (that is, FLT3-TID, DNMT3A, TET2, ASX1, RUNX1, KIT, TP53 or Spliceosome mutations). Additional 2–5 cycles of DCAG therapy were given after they achieved CR in our study; therefore, the high relapse rate may suggest more effective maintenance treatment needs to be considered. The median time to relapse for responders from initiation of treatment and finishing treatment was 10 months. 8 patients relapsed during consolidation cycles or after consolidation. For these people, DCAG regimen can be used as bridging therapy.
In conclusion, we suggest genotyping by NGS first to guide individualized target therapy to solve the management dilemma of older AML patients. DCAG is a safe and effective treatment for older AML patients and may be used as a first-line chemotherapy regimen. However, our study was single and retrospective with many limitations. Large scale and prospective clinical trials are needed in the future to confirm this conclusion.
Supplemental Material
Download MS Word (875 KB)Disclosure statement
No potential conflict of interest was reported by the author(s).
Additional information
Funding
References
- Wingo PA, Tong T, Bolden S. Cancer statistics, 1995. CA Cancer J Clin. 1995;45(1):8–30.
- Khwaja A, Bjorkholm M, Gale RE, et al. Acute myeloid leukaemia. Nat Rev Dis Primers. 2016;2:16010. doi:https://doi.org/10.1038/nrdp.2016.10.
- Klepin HD. Elderly acute myeloid leukemia: assessing risk. Curr Hematol Malig Rep. 2015;10(2):118–125. doi:https://doi.org/10.1007/s11899-015-0257-2.
- Eigendorff E, Hochhaus A. Akute leukämien des erwachsenen. Pathologe. 2015;36(5):503–517; quiz 518–9. doi:https://doi.org/10.1007/s00292-015-0087-y.
- Webster JA, Pratz KW. Acute myeloid leukemia in the elderly: therapeutic options and choice. Leuk Lymphoma. 2017: 1–14. doi:https://doi.org/10.1080/10428194.2017.1330956.
- Hassan C, Afshinnekoo E, Li S, et al. Genetic and epigenetic heterogeneity and the impact on cancer relapse. Exp Hematol. 2017. doi:https://doi.org/10.1016/j.exphem.2017.07.002.
- Dombret H, Gardin C. An update of current treatments for adult acute myeloid leukemia. Blood. 2016;127(1):53–61. doi:https://doi.org/10.1182/blood-2015-08-604520.
- Patel JP, Gonen M, Figueroa ME, et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med. 2012;366(12):1079–1089. doi:https://doi.org/10.1056/NEJMoa1112304.
- Hou HA, Lin CC, Chou WC, et al. Integration of cytogenetic and molecular alterations in risk stratification of 318 patients with de novo non-M3 acute myeloid leukemia. Leukemia. 2014;28(1):50–58. doi:https://doi.org/10.1038/leu.2013.236.
- Wakita S, Yamaguchi H, Ueki T, et al. Complex molecular genetic abnormalities involving three or more genetic mutations are important prognostic factors for acute myeloid leukemia. Leukemia. 2016;30(3):545–554. doi:https://doi.org/10.1038/leu.2015.288.
- Ofran Y, Rowe JM. Genetic profiling in acute myeloid leukaemia–where are we and what is its role in patient management. Br J Haematol. 2013;160(3):303–320. doi:https://doi.org/10.1111/bjh.12135.
- Ley TJ, Ding L, Walter MJ, et al. DNMT3A mutations in acute myeloid leukemia. N Engl J Med. 2010;363(25):2424–2433. doi:https://doi.org/10.1056/NEJMoa1005143.
- Liu WJ, Tan XH, Luo XP, et al. Prognostic significance of Tet methylcytosine dioxygenase 2 (TET2) gene mutations in adult patients with acute myeloid leukemia: a meta-analysis. Leuk Lymphoma. 2014;55(12):2691–2698. doi:https://doi.org/10.3109/10428194.2014.893308.
- Schnittger S, Eder C, Jeromin S, et al. ASXL1 exon 12 mutations are frequent in AML with intermediate risk karyotype and are independently associated with an adverse outcome. Leukemia. 2013;27(1):82–91. doi:https://doi.org/10.1038/leu.2012.262.
- Metzeler KH, Walker A, Geyer S, et al. DNMT3A mutations and response to the hypomethylating agent decitabine in acute myeloid leukemia. Leukemia. 2012;26(5):1106–1107. doi:https://doi.org/10.1038/leu.2011.342.
- Bejar R, Lord A, Stevenson K, et al. TET2 mutations predict response to hypomethylating agents in myelodysplastic syndrome patients. Blood. 2014;124(17):2705–2712. doi:https://doi.org/10.1182/blood-2014-06-582809.
- Emadi A, Faramand R, Carter-Cooper B, et al. Presence of isocitrate dehydrogenase mutations may predict clinical response to hypomethylating agents in patients with acute myeloid leukemia. Am J Hematol. 2015;90(5):E77–E79. doi:https://doi.org/10.1002/ajh.23965.
- Itzykson R, Kosmider O, Cluzeau T, et al. Impact of TET2 mutations on response rate to azacitidine in myelodysplastic syndromes and low blast count acute myeloid leukemias. Leukemia. 2011;25(7):1147–1152. doi:https://doi.org/10.1038/leu.2011.71.
- Greenblatt SM, Nimer SD. Chromatin modifiers and the promise of epigenetic therapy in acute leukemia. Leukemia. 2014;28(7):1396–1406. doi:https://doi.org/10.1038/leu.2014.94.
- Lubbert M, Grishina O, Schmoor C, et al. Valproate and retinoic acid in combination with decitabine in elderly nonfit patients with acute myeloid leukemia: results of a multicenter, randomized, 2×2, phase II trial. Am. J Clin Oncol. 2019: JCO1901053. doi:https://doi.org/10.1200/JCO.19.01053.
- Traina F, Visconte V, Elson P, et al. Impact of molecular mutations on treatment response to DNMT inhibitors in myelodysplasia and related neoplasms. Leukemia. 2014;28(1):78–87. doi:https://doi.org/10.1038/leu.2013.269.
- Hiller JK, Schmoor C, Gaidzik VI, et al. Evaluating the impact of genetic and epigenetic aberrations on survival and response in acute myeloid leukemia patients receiving epigenetic therapy. Ann Hematol. 2017;96(4):559–565. doi:https://doi.org/10.1007/s00277-016-2912-7.
- Arber DA, Orazi A, Hasserjian R, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127(20):2391–2405. doi:https://doi.org/10.1182/blood-2016-03-643544.
- Kiyoi H, Naoe T, Nakano Y, et al. Prognostic implication of FLT3 and N-RAS gene mutations in acute myeloid leukemia. Blood. 1999;93(9):3074–3080.
- Döhner H, Estey E, Grimwade D, et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129(4):424–447.
- Lindsley RC, Mar BG, Mazzola E, et al. Acute myeloid leukemia ontogeny is defined by distinct somatic mutations. Blood. 2015;125(9):1367–1376. doi:https://doi.org/10.1182/blood-2014-11-610543.
- Hou HA, Liu CY, Kuo YY, et al. Splicing factor mutations predict poor prognosis in patients with de novo acute myeloid leukemia. Oncotarget. 2016;7(8):9084–9101. doi:https://doi.org/10.18632/oncotarget.7000.
- O'Donnell MR, Tallman MS, Abboud CN, et al. Acute myeloid leukemia, version 3.2017, NCCN clinical practice guidelines in oncology. J Natl Compr Cancer Network. 2017;15(7):926–957. doi:https://doi.org/10.6004/jnccn.2017.0116.
- Krug U, Gale RP, Berdel WE, et al. Therapy of older persons with acute myeloid leukaemia. Leuk Res. 2017;60:1–10. doi:https://doi.org/10.1016/j.leukres.2017.05.020.
- Kiyoi H. Overview: a new era of cancer genome in myeloid malignancies. Oncology. 2015;89(Suppl 1):1–3. doi:https://doi.org/10.1159/000431054.
- Burnett A, Wetzler M, Lowenberg B. Therapeutic advances in acute myeloid leukemia. J Clin Oncol. 2011;29(5):487–494. doi:https://doi.org/10.1200/JCO.2010.30.1820.
- Tallman MS. New strategies for the treatment of acute myeloid leukemia including antibodies and other novel agents. Hematology Am Soc Hematol Educ Program. 2005;2005(1):143–150. doi:https://doi.org/10.1182/asheducation-2005.1.143.
- Rowe JM, Neuberg D, Friedenberg W, et al. A phase 3 study of three induction regimens and of priming with GM-CSF in older adults with acute myeloid leukemia: a trial by the eastern cooperative oncology group. Blood. 2004;103(2):479–485. doi:https://doi.org/10.1182/blood-2003-05-1686.
- Gardin C, Turlure P, Fagot T, et al. Postremission treatment of elderly patients with acute myeloid leukemia in first complete remission after intensive induction chemotherapy: results of the multicenter randomized Acute Leukemia French Association (ALFA) 9803 trial. Blood. 2007;109(12):5129–5135. doi:https://doi.org/10.1182/blood-2007-02-069666.
- Lowenberg B, Zittoun R, Kerkhofs H, et al. On the value of intensive remission-induction chemotherapy in elderly patients of 65+ years with acute myeloid leukemia: a randomized phase III study of the European Organization for Research and Treatment of Cancer Leukemia Group. J Clin Oncol. 1989;7(9):1268–1274. doi:https://doi.org/10.1200/JCO.1989.7.9.1268.
- Burnett AK, Milligan D, Prentice AG, et al. A comparison of low-dose cytarabine and hydroxyurea with or without all-trans retinoic acid for acute myeloid leukemia and high-risk myelodysplastic syndrome in patients not considered fit for intensive treatment. Cancer. 2007;109(6):1114–1124. doi:https://doi.org/10.1002/cncr.22496.
- Dohner H, Estey EH, Amadori S, et al. Diagnosis and management of acute myeloid leukemia in adults: recommendations from an international expert panel, on behalf of the European LeukemiaNet. Blood. 2010;115(3):453–474. doi:https://doi.org/10.1182/blood-2009-07-235358.
- Grimwade D, Hills RK, Moorman AV, et al. Refinement of cytogenetic classification in acute myeloid leukemia: determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood. 2010;116(3):354–365. doi:https://doi.org/10.1182/blood-2009-11-254441.
- Melki JR, Vincent PC, Clark SJ. Concurrent DNA hypermethylation of multiple genes in acute myeloid leukemia. Cancer Res. 1999;59(15):3730–3740.
- Toyota M, Kopecky KJ, Toyota MO, et al. Methylation profiling in acute myeloid leukemia. Blood. 2001;97(9):2823–2829.
- Qin T, Youssef EM, Jelinek J, et al. Effect of cytarabine and decitabine in combination in human leukemic cell lines. Clin Cancer Res. 2007;13(14):4225–4232. doi:https://doi.org/10.1158/1078-0432.CCR-06-2762.
- Cruijsen M, Lubbert M, Wijermans P, et al. Clinical results of hypomethylating agents in AML treatment. J Clin Med. 2015;4(1):1–17. doi:https://doi.org/10.3390/jcm4010001.
- Shen L, Kantarjian H, Guo Y, et al. DNA methylation predicts survival and response to therapy in patients with myelodysplastic syndromes. J Clin Oncol. 2010;28(4):605–613. doi:https://doi.org/10.1200/JCO.2009.23.4781.
- Kantarjian HM, Thomas XG, Dmoszynska A, et al. Multicenter, randomized, open-label, phase III trial of decitabine versus patient choice, with physician advice, of either supportive care or low-dose cytarabine for the treatment of older patients with newly diagnosed acute myeloid leukemia. J Clin Oncol. 2012;30(21):2670–2677. doi:https://doi.org/10.1200/JCO.2011.38.9429.
- Lubbert M, Suciu S, Baila L, et al. Low-dose decitabine versus best supportive care in elderly patients with intermediate- or high-risk myelodysplastic syndrome (MDS) ineligible for intensive chemotherapy: final results of the randomized phase III study of the European Organisation for Research and Treatment of Cancer Leukemia Group and the German MDS Study Group. J Clin Oncol. 2011;29(15):1987–1996. doi:https://doi.org/10.1200/JCO.2010.30.9245.
- Blum W, Garzon R, Klisovic RB, et al. Clinical response and miR-29b predictive significance in older AML patients treated with a 10-day schedule of decitabine. Proc Natl Acad Sci USA. 2010;107(16):7473–7478. doi:https://doi.org/10.1073/pnas.1002650107.
- Bhatla T, Wang J, Morrison DJ, et al. Epigenetic reprogramming reverses the relapse-specific gene expression signature and restores chemosensitivity in childhood B-lymphoblastic leukemia. Blood. 2012;119(22):5201–5210. doi:https://doi.org/10.1182/blood-2012-01-401687.
- Li J, Chen Y, Zhu Y, et al. Efficacy and safety of decitabine in combination with G-CSF, low-dose cytarabine and aclarubicin in newly diagnosed elderly patients with acute myeloid leukemia. Oncotarget. 2015;6(8):6448–6458. doi:https://doi.org/10.18632/oncotarget.3361.
- Scandura JM, Roboz GJ, Moh M, et al. Phase 1 study of epigenetic priming with decitabine prior to standard induction chemotherapy for patients with AML. Blood. 2011;118(6):1472–1480. doi:https://doi.org/10.1182/blood-2010-11-320093.
- Lubbert M, Ruter BH, Claus R, et al. A multicenter phase II trial of decitabine as first-line treatment for older patients with acute myeloid leukemia judged unfit for induction chemotherapy. Haematologica. 2012;97(3):393–401. doi:https://doi.org/10.3324/haematol.2011.048231.
- Quintas-Cardama A, Ravandi F, Liu-Dumlao T, et al. Epigenetic therapy is associated with similar survival compared with intensive chemotherapy in older patients with newly diagnosed acute myeloid leukemia. Blood. 2012;120(24):4840–4845. doi:https://doi.org/10.1182/blood-2012-06-436055.
- Mayer J, Arthur C, Delaunay J, et al. Multivariate and subgroup analyses of a randomized, multinational, phase 3 trial of decitabine vs treatment choice of supportive care or cytarabine in older patients with newly diagnosed acute myeloid leukemia and poor- or intermediate-risk cytogenetics. BMC Cancer. 2014;14:69. doi:https://doi.org/10.1186/1471-2407-14-69.
- Xu Q, Li Y, Jing Y, et al. Epigenetic modifier gene mutations-positive AML patients with intermediate-risk karyotypes benefit from decitabine with CAG regimen. Int J Cancer. 2020;146(5):1457–1467.
- Wang B, Liu Y, Hou G, et al. Mutational spectrum and risk stratification of intermediate-risk acute myeloid leukemia patients based on next-generation sequencing. Oncotarget. 2016;7(22):32065–32078. doi:https://doi.org/10.18632/oncotarget.7028.
- Thomas X, Le Jeune C. Treatment of elderly patients with acute myeloid leukemia. Curr Treat Options Oncol. 2017;18(1):2. doi:https://doi.org/10.1007/s11864-017-0445-5.
- Chang E, Ganguly S, Rajkhowa T, et al. The combination of FLT3 and DNA methyltransferase inhibition is synergistically cytotoxic to FLT3/ITD acute myeloid leukemia cells. Leukemia. 2016;30(5):1025–1032. doi:https://doi.org/10.1038/leu.2015.346.
- Yanada M, Naoe T. Acute myeloid leukemia in older adults. Int J Hematol. 2012;96(2):186–193. doi:https://doi.org/10.1007/s12185-012-1137-3.
- Shallis RM, Boddu PC, Bewersdorf JP, et al. The golden age for patients in their golden years: The progressive upheaval of age and the treatment of newly-diagnosed acute myeloid leukemia. Blood Rev. 2019: 100639. doi:https://doi.org/10.1016/j.blre.2019.100639.