1. Introduction
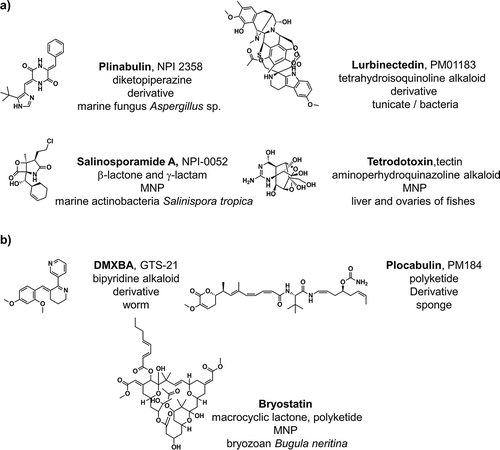
A statistical analysis of the number of new U.S. Food and Drug Administration (FDA) approvals in the period between 1969 and 2016 had shown a downward trend from its highest point in 1996 (53 new molecular entities (NMEs)/year) and the minimum after 1996 of 15 NMEs/year in 2010 and 2016 [Citation1]. However, 2017 and 2018 appear to be turning years in development pipelines with higher NMEs/year values (34 and 52, respectively), almost surpassing even the highest NMEs value (53 NMEs/1996) in 2018 [Citation2]. The global marine pharmaceutical clinical pipeline comprising 30 compounds, which includes eight approved drugs by the most representative approving agencies such as U.S. FDA, European Medicines Agency (EMEA), Japanese Ministry of Health and Australia’s Therapeutic Goods Administration, , and 22 drug candidates in phase III, II, or I of drug development [Citation1,Citation3–Citation5]. Interestingly, more than half of the total approvals of marine natural products (MNPs) and derivatives occurred in the 21st century (six out of eight approved drugs) [Citation1], . also highlighted the year and the approval agency for the first approval of the eight approved drugs of marine origin. In clinical trials phase III, there are six MNPs and derivatives, which includes four small molecules and two antibody-drug conjugates. In ), the chemical structures of the four small molecules in phase III were represented, which includes three anti-cancer compounds (plinabulin, lurbinectedin, and salinosporamide A) and one analgesic compound (tetrodotoxin) [Citation6,Citation7]. While in phase II there are 10 MNPs and derivatives, from those three are small molecules, one anti-cancer compound (plocabulin) and two are active against Alzheimer’s disease (DMXBA and bryostatin) according to pre-clinical studies that were previously performed, ). All the MNPs and derivatives in phase I of the clinical trials are formerly established as anti-cancer antibody-drug conjugates. However, it is important to note that the known biological activity space of MNPs and derivatives has been influenced by funding sources, e.g. 5 out of the 8 approved MNP and derivatives drugs () and 19 out of the 22 MNPs and derivatives in phase I, II, and III of the clinical trials () are set as anti-cancer drugs. The emphasis on cancer is mainly due to the fact that the leading funding agency in the U.S. for MNP and derivatives was for many years the National Institutes of Health (NIH)/National Cancer Institute (NCI), and this situation is the same in other countries [Citation4].
Figure 1. The eight MNPs and derivatives approved drugs, their chemical structures, biological sources, and clinical applications. Adapted from [Citation1] with permission of MDPI.
![Figure 1. The eight MNPs and derivatives approved drugs, their chemical structures, biological sources, and clinical applications. Adapted from [Citation1] with permission of MDPI.](/cms/asset/6e955777-55a6-4b8c-a827-ef8c28bbb7a4/iedc_a_1604675_f0001_oc.jpg)
Therefore, this corresponds to a higher success rate of marine compounds, 1 in 3,500 MNPs [Citation4], compared with the industry average of 1 in 5,000–10,000 compounds [Citation5]. In recent years, there has been a growing interest in NP-like scaffolds due to these two factors, on the one hand, the decreasing number of NMEs in development pipelines and, on the other hand, the success rate of the marine world.
2. Marine natural product chemical space
NPs remain one of the most productive sources of chemical inspiration for the development of new drugs. The structures of more than 440,000 NPs are available from public databases such as PubChem, REAXYS, ChEMBL, ZINC, NaprAlert, and Super Natural II [Citation1,Citation8]. From these, approximately 10% are commercially available compounds [Citation9], which can be obtained from commercial vendors and public research institutions to be tested experimentally, e.g. the ZINC database comprising more than 10,000 purchase NPs out of 100,000 NPs that are recorded [Citation10].In order to understand the chemical space defined by known and readily obtainable NPs as well as by individual NP databases, comparisons between these chemical spaces and those defined by drugs and other types of small molecules have been reported in several works [Citation9,Citation11,Citation12]. Ertl and Schuffenhauer performed a chemoinformatics analysis of a large collection of over 130,000 NPs and compared the physicochemical and structural properties of NPs with those of bioactive and common synthetic molecules [Citation11]. Although the authors conclude that there is a clear separation in the structural space that NPs and synthetic molecules define, they also found an overlap of the bioactive molecules in both groups [Citation11]. Chen et al. [Citation9] reported a comprehensive study with known and readily obtainable NPs from 18 databases, 9 physical libraries, and the Protein Data Bank (PDB). The authors found that the readily obtainable NPs are highly diverse and populate regions of chemical space that are of high relevance to drug discovery. In the case of PubChem database and TCM Database@ Taiwan, substantial differences in the coverage of NP classes and chemical space by the individual databases are observed, namely the high proportion of drug-like NPs and chiral NPs, respectively. Recently, Shang et al. reported a chemoinformatics comparison between the chemical spaces covered by terrestrial and marine originated NPs and concluded that MNP and terrestrial NPs (TNPs) may occupy different biologically relevant chemical spaces [Citation12]. The authors pointed out that MNPs have lower solubility and are often larger than TNPs and often contain ester bonds connected to 10-membered rings, while the scaffolds of the TNPs often have more stable ring systems and bond types. They also verified that MNPs had more nitrogen atoms and halogen atoms, especially bromine atoms, and fewer oxygen atoms, suggesting that MNPs can be synthesized by more diverse biosynthetic pathways than TNPs [Citation12].
3. Marine natural product drug discovery
In spite of marine invertebrates such as sponge, jellyfish, anemone, and a rocky coral have been the source of most bioactive MNPs, with Porifera (sponges) and Cnidaria phyla being the most prolific, the true origin of most MNPs appears to be the microorganisms who live with them in a symbiotic relationship [Citation6]. Almost all of the MNPs approved as drugs or currently in clinical trials come from bacterial and cyanobacterial biosynthetic sources, e.g. brentuximab vedotin (, cyanobacteria Symploca sp.) and salinosporamide A (, actinobacteria Salinispora tropica) [Citation5]. The Ascomycota (Fungi) and Actinobacteria have been among the four most widely collected phyla during the last years, along with Porifera and Cnidaria [Citation6]. Therefore, microbial-derived compounds will almost certainly dominate the MNP field in the coming years.
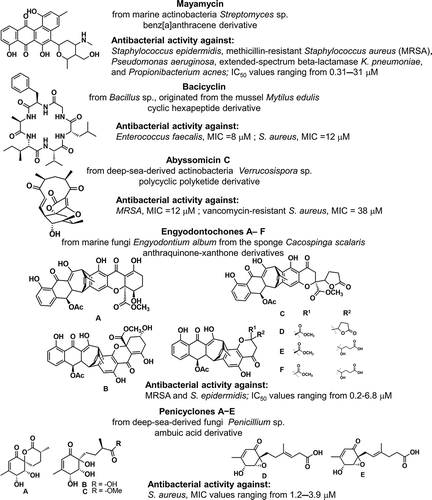
The vast majority of currently used antibiotics have been isolated from terrestrial microbes, accounting for more than 75% of all antibiotics discovered, however antimicrobial compounds from marine sources have not yet been developed into clinical trial phases [Citation3,Citation4,Citation13]. As reported above, there is a vast investment and research in the development of novel anti-cancer drugs when compared to antibacterial macromolecules. Wiese et al. reported the analyses of antibiotic activity surveys obtained in their lab during the years 2008–2017 and concluded that ca. 50% of the bacterial isolates revealed antibiotic activity against Gram-positive (Bacillus subtilis, Staphylococcus lentus) and Gram-negative strains (Escherichia coli) [Citation3]. The authors underscored the enormous potential of marine micro-organisms for the production of new anti-infective agents with some examples of MNPs, namely polyketides and cyclic peptides isolated from bacteria and marine fungi [Citation3], . Another interesting review addressed advances in the discovery of novel antibiotics isolated from deep-sea microorganisms, which are adapted to this extreme environment and therefore have the potential to produce novel secondary metabolites with potent biological activities [Citation14]. In , a few examples of deep-see-derived MNPs isolated from actinobacteria and fungi were also highlighted [Citation15].
Figure 3. A few examples of antibacterial MNPs and derivatives isolated from marine bacteria and fungi.
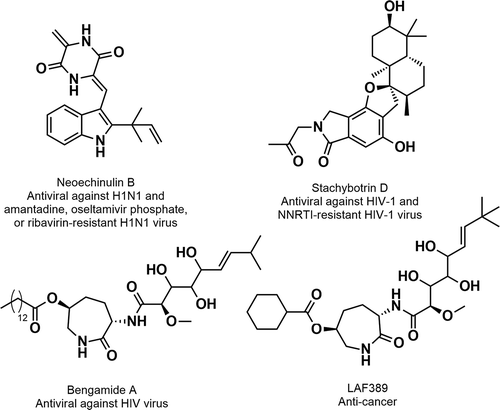
The discovery of antibiotics has undoubtedly revolutionized medicine and is considered a turning point in the history of mankind. However, the most serious consequence of the use and misuse of antibiotics is the associated development of resistance against them. The marine environment has proven to be a very rich source of diverse NPs with relevant activity against fungal, viral, and parasitic drug-resistant infections [Citation15,Citation16]. For example, the prenylated indole diketopiperazine alkaloid neoechinulin B, , isolated from a marine-derived fungus Eurotium rubrum displayed a strong inhibition against the H1N1 virus in infected MDCK cells and also inhibited several drug-resistant influenza clinical isolates, e.g. resistant against amantadine, oseltamivir phosphate, and ribavirin [Citation15].
In addition, HIV also exhibits high genetic variability, and thus develops resistance to existing drugs and escapes host immune responses elicited by AIDS vaccine candidates. In another example, the phenylspirodrimane stachybotrin D, , isolated from the marine sponge-associated fungus Stachybotrys chartarum, showed inhibition against the HIV-1 virus, as well as showed similar inhibitory effects on HIV-1 replication of wild type and several NNRTI-resistant HIV-1 strains [Citation15]. In addition, bengamide A (a seven-membered lactam), , isolated from the marine sponge-associated bacterium Myxococcus virescens, inhibits HIV replication in vitro and in primary cells with EC50s of 15–32 nM [Citation16]. Bengamide A acts by inhibiting the cellular NF-κB signaling pathway. The synthetic derivative of bengamide A (LAF389), , was tested as an anti-cancer agent in phase I of the clinical trials, but the study was terminated due to cardiovascular and pulmonary toxicities [Citation16].
4. Expert opinion
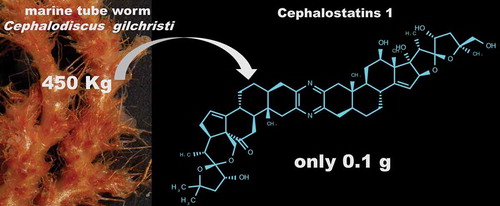
Although the field of MNP drug discovery has been fully implemented during the 21st century there are still several bottlenecks that inhibit the development of drugs from NPs and particularly MNPs. One of the most important prerequisites is the sustainable supply of NPs through the robust development of processes using fermentation procedures, chemical synthesis or genetic methods. For example, the anti-cancer clinical trial involving cephalostatin 1 (a bis-steroidal alkaloid) has been hampered because of complications in the collection of materials [Citation17], . Issues related to material harvesting can be overcome for the most prolific MNPs, the microbial derivative compounds. These MNPs present several advantages compared to the MNPs derived from macroorganisms, namely, their direct fermentation, the relative ease of their collection, and the advanced technologies for the extraction of their genomic material and their manipulation in heterologous systems, as well as the application of emerging advanced techniques [Citation6]. In general, the design and total or semi-synthetic chemical synthesis of MNPs derived from macro- or microorganisms has been the approach used to make the compounds available for clinical trials.
However, even for compounds of moderate complexity, organic synthesis remains a major challenge. Machine learning approaches have been successfully used in Computer Aided Synthesis Planning (CASP) [Citation18,Citation19] – chemical synthesis, Computer-Assisted Identification of Compounds [Citation1] – dereplication (recognizing and filtering known compounds), and Computer-Aided Drug Design (CADD) [Citation1] – ligand-based and structure-based methodologies.
Computer-aided synthetic pathways of isolated NPs such as a pentacyclic alkaloid, tacamonidine (not yet synthesized in the laboratory), have recently been reported [Citation19] and allowed to conclude that computers are currently capable of designing syntheses important to the everyday practice of organic synthesis [Citation18,Citation19] after about half a century since the first project in this field. Several advances have contributed to the beginning of the success of CASP, e.g. availability of millions of tabulated reaction examples from United States Patent and Trademark Office (USPTO), Reaxys, and SciFinder databases; machine learning techniques (Weisfeller−Lehman network, Weisfeller−Lehman difference network, graph convolutional neural network); establishment of retrosynthetic rules; predicting reaction conditions (solvent, catalyst, temperature, etc.); and in silico discovery of new reaction types and mechanisms [Citation18,Citation19]. Therefore, computational methodologies will be crucial to streamline the identification and elucidation of MNPs, design and chemical synthesis of MNPs [Citation18,Citation19], identification and validation of the biological targets, and prediction of biological activities of MNPs [Citation1].
Declaration of interest
F Pereira has no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
Reviewer Disclosures
Peer reviewers on this manuscript have no relevant financial or other relationships to disclose.
Additional information
Funding
References
- Pereira F, Aires-de-Sousa J. Computational methodologies in the exploration of marine natural product leads. Mar Drugs. 2018;16(7):236.
- FDA’s Center for Drug Evaluation and Research (CDER). Novel drug approvals for 2018. [cited 2019 Mar]. Available from: https://www.fda.gov/drugs/developmentapprovalprocess/druginnovation/ucm592464.htm
- Wiese J, Imhoff JF. Marine bacteria and fungi as promising source for new antibiotics. Drug Dev Res. 2019;80(1):24–27.
- Newman DJ, Cragg GM. Drugs and drug candidates from marine sources: an assessment of the current “State of play”. Planta Med. 2016;82:775–789.
- Gerwick WH, Moore BS. Lessons from the past and charting the future of marine natural products drug discovery and chemical biology. Chem Biol. 2012;19(1):85–98.
- Jimenez C. Marine natural products in medicinal chemistry. ACS Med Chem Lett. 2018;9(10):959–961.
- Midwestern University, Tomorrow´s Healthcare Team. Marine pharmaceuticals: the clinical pipeline. [cited 2019 Mar]. Available from: http://marinepharmacology.midwestern.edu/clinical_pipeline.html
- Banerjee P, Erehman J, Gohlke BO, et al. Super natural II: a database of natural products. Nucleic Acids Res. 2015;43(D1):D935–D939.
- Chen Y, de Lomana MG, Friedrich N-O, et al. Characterization of the chemical space of known and readily obtainable natural products. J Chem Inf Model. 2018;58(8):1518–1532.
- Natural Products Catalogs. ZINC database. [cited 2019 Mar 21] Available from: http://zinc.docking.org/browse/catalogs/natural-products
- Ertl P, Schuffenhauer A. Cheminformatics analysis of natural products: lessons from nature inspiring the design of new drugs. In: Petersen F, Amstutz R, editors. Natural compounds as drugs. progress in drug research, Birkhäuser, basel. Vol. 66. 2008. pp. 217–235.
- Shang J, Hu B, Wang J, et al. A cheminformatic insight into the differences between terrestrial and marine originated natural products. J Chem Inf Model. 2018;58(6):1182–1193.
- Choudhary A, Naughton LM, Montanchez I, et al. Current status and future prospects of Marine Natural Products (MNPs) as antimicrobials. Mar Drugs. 2017;15(9):272.
- Tortorella E, Tedesco P, Esposito FP, et al. Antibiotics from deep-sea microorganisms: current discoveries and perspectives. Mar Drugs. 2018;16(10):355.
- Abdelmohsen UR, Balasubramanian S, Oelschlaeger TA, et al. Potential of marine natural products against drug-resistant fungal, viral, and parasitic infections. Lancet Infect Dis. 2017;17(2):E30–E41.
- Andersen RJ, Ntie-Kang F, Tietjen I. Natural product-derived compounds in HIV suppression, remission, and eradication strategies. Antiviral Res. 2018;158:63–77.
- Dey P, Kundu A, Chakraborty HJ, et al. Therapeutic value of steroidal alkaloids in cancer: current trends and future perspectives. Int J Cancer. 2018. DOI:10.1002/ijc.31965.
- Coley CW, Green WH, Jensen KF. Machine learning in computer-aided synthesis planning. Acc Chem Res. 2018;51(5):1281–1289.
- Szymkuc S, Gajewska EP, Klucznik T, et al. Computer-assisted synthetic planning: the end of the beginning. Angew Chem Int Ed. 2016;55(20):5904–5937.