ABSTRACT
In our previous work, we reported comparative genomics studies on 3 Sinocyclocheilus fishes, including the surface-dwelling S. graham (Sg), the semi-cave-dewlling S. rhinocerous (Sr) and the cave-restricted S. anshuiensis (Sa). Here, we performed a genomic survey on the immunity differences among the 3 fishes by combination of the genome and transcriptome data. Our results demonstrate that immune-related genetic pathways in Sa and Sr present more similarities than Sg. The innate immune activity in Sa seems to be higher than Sg and Sr, while Sr owns more copy numbers of MHC-related genes (related with the acquired immune system) than Sg and Sa. These differential immune activities in the 3 Sinocyclocheilus species may be due to their differential habitats. Generally speaking, most immunity genes transcribe the lowest levels in Sa when compared with Sg and Sr, which may be associate with a less various microbial environment in the cave-restricted habitat. However, Sr has more members of MHC-related genes than those in Sg and Sa, suggesting a great contribution from the semi-cave-dwelling condition.
The freshwater Sinocyclocheilus (Cypriniformes: Cyprinidae) fishes are endemic to China’s massive karst area, i.e., the eastern part of Yungui Plateau and the surrounding region (including northwest Guangxi Province and east Yunnan Province).Citation1 The genus Sinocyclocheilus includes surface-dwelling, semi-cave-dwelling and cave-restricted species, which makes it a special group for investigating the molecular mechanisms of adaptive evolution in cavefishes.Citation2 In our previous study, we compared the whole genome sequences of 3 Sinocyclocheilus species, including the surface-dwelling S. graham (Sg), the semi-cave-dwelling S. rhinocerous (Sr) and the cave-restricted S. anshuiensis (Sa). By comparative genomics analyses of the 3 genomes, we provided genetic evidence for the cave adaption of cavefishes, such as eye degeneration, albinism and hearing loss.Citation3
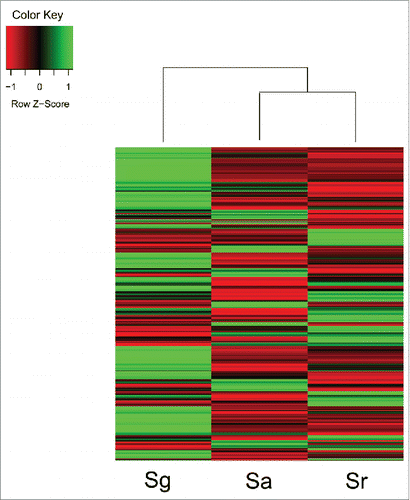
Interestingly, immune activities in the 3 Sinocyclocheilus fishes are obviously different. For example, Sa is more susceptible to disease in captivity (Jiang et al., unpublished data). In our previous work,Citation3 we annotated all predicted genes and calculated their expression levels in the skin of the 3 Sinocyclocheilus fishes. Here, we picked out 800 “1:1:1” orthologous immune-related genes, and identified 396 of them are differentially expressed genes (DEGs; changes over 2 folds). Different from Sg, Sa and Sr present similar gene expression patterns (), which is consistent with the similar cave-dwelling behaviors of Sa and Sr. This also suggests difference of immunity system between surface-dwelling and cave-dwelling fishes. Simultaneously, most immunity genes transcribe the lowest levels in Sa when compared with Sg and Sr,Citation4 which may be associate with a less various microbial environment in the cave-restricted habitat.
TLR (toll-like receptor) gene family is an important group of innate immune receptors located on the surface of membrane.Citation5 They are divided into 13 families and named as TLR1 to TLR13.Citation6 Most of them play a significant role in immunity by detecting pathogen-associated molecular patterns (PAMPs).Citation5 We analyzed the TLR gene family among the 3 Sinocyclocheilus fishes, and identified 2 copies of Tlr8 and Tlr18 genes in the Sa genome while only one copy in the Sg and Sr genomes. This difference may tell us that the innate immune activity in cave-dwelling fishes is higher than that in surface-dwelling and cave-restricted fishes.
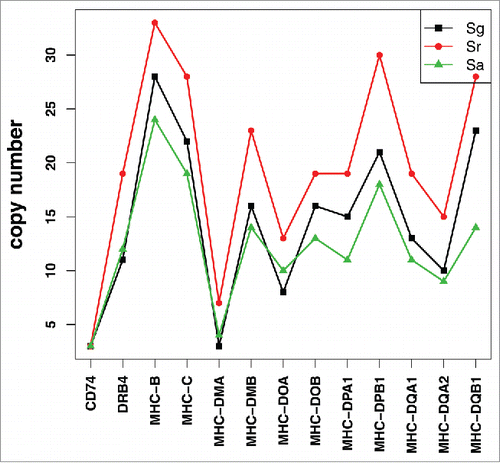
The major histocompatibility complex (MHC) is a set of cell surface proteins essential for acquired immune system to recognize foreign substances in vertebrates.Citation7 MHC genes include MHC class I and MHC class II; MHC class I genes are responsible for protection against viral pathogens, while MHC class II genes are eligible for detection of bacterial pathogens in vertebrates.Citation8 Therefore, elevation of MHC gene numbers will facilitate pathogen detection and improve immunity. Here we identified MHC gene family in the 3 Sinocyclocheilus fishes, and observed that Sr owns the highest number of MHC genes while Sa has the least (). In our previous genome paper,Citation3 we analyzed contraction and expansion of gene families in the 3 Sinocyclocheilus fishes and all the predicted proteins were subjected to GO/KEGG/IPR enrichment analyses. Among these gene families, we discovered that the GO terms of visual perception, MHC class II protein complex and antigen processing presentation were undergoing contraction in Sa (). The GO terms related to acquired immunity in Sr, such as MHC protein complex and immune response, are undergoing expansion (). These results suggest that the acquired immunity of cavefishes may be lower than that of their surface-dwelling counterparts, because they live in a subterranean habitats with less microbial and food. Sr has more members of MHC-related genes than those in Sg and Sa, suggesting a great contribution from the semi-cave-dwelling condition. Hence, we deduce that Sr should have more intense immune defense against pathogens than those encountered in cave or surface alone. This phenomenon is similar to that in the reported amphibious mudskippers.Citation9
Table 1. The GO enrichment analysis of contraction gene families in Sa.
Table 2. The GO enrichment analysis of expansion gene families in Sr.
In summary, our current results provide new insights into the genetic basis for immunity in cavefishes, and reach a novel conclusion that the 3 Sinocyclocheilus fishes have developed differential immune activities that may be related to their differential habitats. We also provide a solid foundation for further investigation of the biological differences in immunity of the 3 Sinocyclocheilus fishes.
Disclosure of potential conflicts of interest
No potential conflicts of interest were disclosed.
Funding
This work was supported by China Shenzhen Special Program for Future Industrial Development (No. JSGG20141020113728803), Special Project on the Integration of Industry, Education and Research of Guangdong Province (No. 2013B090800017), and Zhenjiang Leading Talent Program for Innovation and Entrepreneurship.
References
- Chen SY, Zhang RD, Feng JG, Xiao H, Li WX, Zan RG, Zhang YP. Exploring factors shaping population genetic structure of the freshwater fish Sinocyclocheilus grahami (Teleostei, Cyprinidae). J Fish Biol 2009; 74(8):1774-86; http://www.ncbi.nlm.nih.gov/pubmed/20735670; PMID:20735670; http://dx.doi.org/10.1111/j.1095-8649.2009.02204.x
- Jeffery WR. Cavefish as a model system in evolutionary developmental biology. Dev Biol 2001; 231(1):1-12; http://www.ncbi.nlm.nih.gov/pubmed/11180948; PMID:11180948; http://dx.doi.org/10.1006/dbio.2000.0121
- Yang J, Chen X, Bai J, Fang D, Qiu Y, Jiang W, Yuan H, Bian C, Lu J, He S, et al. The Sinocyclocheilus cavefish genome provides insights into cave adaptation. BMC Biol 2016; 14:1; http://www.ncbi.nlm.nih.gov/pubmed/26728391; PMID:26728391; http://dx.doi.org/10.1186/s12915-015-0223-4
- Trajano E, Bichuette ME, Kapoor BG. Biology of subterranean fishes. 1st ed. Enfield, N.H. Boca Raton: Science Publishers ;CRC Press; 2010. xiv, 480.
- Takeda K, Akira S. Toll-like receptors in innate immunity. Int Immunol 2005; 17(1):1-14; http://www.ncbi.nlm.nih.gov/pubmed/15585605; PMID:15585605; http://dx.doi.org/10.1093/intimm/dxh186
- Roach JC, Glusman G, Rowen L, Kaur A, Purcell MK, Smith KD, Hood LE, Aderem A. The evolution of vertebrate Toll-like receptors. Proc Natl Acad Sci U S A 2005; 102(27):9577-82; http://www.ncbi.nlm.nih.gov/pubmed/15976025; PMID:15976025; http://dx.doi.org/10.1073/pnas.0502272102
- Hughes AL, Yeager M. Natural selection at major histocompatibility complex loci of vertebrates. Annu Rev Genet 1998; 32:415-35; http://www.ncbi.nlm.nih.gov/pubmed/9928486; PMID:9928486; http://dx.doi.org/10.1146/annurev.genet.32.1.415
- Neefjes J, Jongsma ML, Paul P, Bakke O. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat Rev Immunol 2011; 11(12):823-36; http://www.ncbi.nlm.nih.gov/pubmed/22076556; PMID:22076556
- You X, Bian C, Zan Q, Xu X, Liu X, Chen J, Wang J, Qiu Y, Li W, Zhang X, et al. Mudskipper genomes provide insights into the terrestrial adaptation of amphibious fishes. Nat Commun 2014; 5:5594; http://www.ncbi.nlm.nih.gov/pubmed/25463417; PMID:25463417; http://dx.doi.org/10.1038/ncomms6594