Abstract
The myosin holoenzyme is a multimeric protein complex consisting of heavy chains and light chains. Myosin light chains are calmodulin family members which are crucially involved in the mechanoenzymatic function of the myosin holoenzyme. This review examines the diversity of light chains within the myosin superfamily, discusses interactions between the light chain and the myosin heavy chain as well as regulatory and structural functions of the light chain as a subunit of the myosin holoenzyme. It covers aspects of the myosin light chain in the localization of the myosin holoenzyme, protein-protein interactions and light chain binding to non-myosin binding partners. Finally, this review challenges the dogma that myosin regulatory and essential light chain exclusively associate with conventional myosin heavy chains while unconventional myosin heavy chains usually associate with calmodulin.
Abbreviations
| CaM | = | calmodulin |
| EDTA | = | ethylenediaminetetraacetic acid |
| ELC | = | essential light chain |
| FRET | = | Förster resonance energy transfer |
| MLCK | = | myosin light chain kinase |
| PD | = | phosphorylation domain |
| PKC | = | protein kinase C |
| RLC | = | regulatory light chain |
| ROCK | = | rho kinase |
| WT | = | wild type |
| ZIP kinase | = | zipper-interacting protein kinase. |
Introduction
Myosins are involved in a myriad of functions such as muscle contraction, cytokinesis, cargo transport, cell adhesion, cell migration and the formation and stabilization of actin-rich structures such as stereocilia.Citation1 Phylogenetic analysis groups myosins in 35 classes of which 12 are found in humans.Citation2 The myosin holoenzyme consists of heavy and light chains. The myosin heavy chain is composed of an N-terminal motor domain, a central neck domain and a C-terminal tail domain (). The motor domain mediates the ATP-dependent interaction with the F-actin cytoskeleton. The tail domain determines the oligomeric state of the heavy chain and interaction with binding partners or cellular compartments.
Figure 1. The myosin IQ motif in the heavy chain. (A) Schematic representation of a generic myosin heavy chain. The N-terminus of the heavy chain represents the catalytic motor domain (gray) that harbors the nucleotide binding site and the F-actin binding region. The central neck domain harbors the IQ motifs (orange). The C-terminal tail domain (dark gray) can contain a vast collection of domains and motifs that determine the oligomeric state of the heavy chain and interaction signatures with binding partners. The length of the myosin motor domain is very conserved, whereas the length of the myosin neck and tail domain varies considerably. Some myosins have an N-terminal extension of variable length prior to the motor domain which is indicated by the dashed line. (B) The top line represents the consensus IQ motif found in myosin heavy chains. Position 1 of the 11 amino acid long consensus IQ motif is critically occupied by a hydrophobic residue such as isoleucine (I), leucine (L) or valine (V) though methionine (M), phenylalanine (F) and lysine (K) and threonine (T) are found in some cases. Position 7 is ambiguous for several amino acids. Position 11 is preferentially occupied by a positively charged amino acid (arginine (R), histidine (H) or lysine (K)). Invariant residues are boxed in gray, ambiguous residues in peach. Non-boxed residues are highly variable. The WXW motif found in the hook region of conventional myosins-2 is colored in rose. Of note, the WXW motif is not part of the consensus IQ motif but crucial for the interaction of RLCs with the IQ2 in myosins-2 and therefore depicted for completeness. All other lines shown represent selected IQ motif found in conventional and unconventional myosin heavy chains discussed in this review. The abbreviations used are as follows: M1: IQ1 and IQ3 from human myosin-1C, M2: IQ1 and IQ2 from chicken striated muscle myosin-2; M5: IQ2 and IQ4 from budding yeast Myo2p; M6: IQ1 from pig myosin-6; M7: IQ3 from human myosin-7A; M10: IQ3 from human myosin-10; M14: IQ2 from P.yoelli. (C) Crystal structure of a prototypic myosin-2–2IQ fragment. Structural overview of the striated myosin-2–2IQ fragment from scallop (PDB ID: 1DFL). The myosin heavy chain is shown in gray cartoon representation, the nucleotide in black spheres. The associated ELC and RLC are represented in orange and cherry color, respectively. ELC and RLC bind distal from the globular motor domain to the IQ motifs of the neck domain. The myosin neck region and the associated light chains form the lever arm. The hook is indicated by a gray arrow.
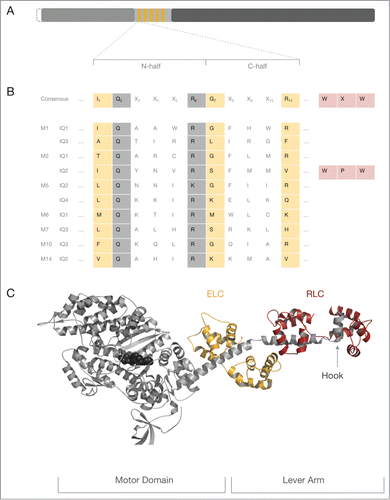
Myosin light chains are members of the calmodulin (CaM) and CaM-related gene families and associate non-covalently with the 20–25 amino acid IQ motif(s) (consensus sequence IQXXXRGXXXR)Citation3 located in the myosin neck domain ().Citation4 As reviewed previously by Bähler et al., the IQ motif is strictly α-helical and devoid of proline residues.Citation4 Position 1 of the IQ motif is occupied by a hydrophobic amino acid. The IQ motif may have a partial or no amphipathicity and a basic net charge usually between +2 and +5.Citation4 IQ motifs often display large hydrophobic residues in conserved positions (1–8–14) which face to one side of the α-helixCitation5 and usually occur in tandems separated by 20–27 amino acids measuring from the first residue of the motif.Citation4 IQ motifs of the myosin heavy chain are numbered, starting with IQ1 for the motif in closest proximity to the motor domain. The number of light chains predicted to bind to a single myosin heavy chain can vary between one and 17.Citation2
Myosin light chains are required for the structural integrity of the myosin holoenzyme. In addition, they can have regulatory functions on the mechanoenzymatic activity of the protein complex. In this review, we evaluate the interactions between myosin light chains and the myosin heavy chain, summarize recent reports of light chain heterogeneity within the myosin superfamily and discuss how the light chains participate in mechanical and regulatory functions of conventional and unconventional myosins. Further, this review briefly describes implications of myosin light chains in protein-protein interactions and the localization of the myosin holoenzyme and described non-myosin binding partners of myosin light chains.
Native Light Chain Composition of the Myosin Holoenzyme
Filament-forming class-2 myosins, historically referred to as “conventional,” were the first myosins isolated from muscles and later shown to be present in all eukaryotic cells. Biochemical and structural studies reveal that each myosin-2 heavy chain contains 2 IQ motifs and associates exclusively with 2 light chains most commonly referred to as essential light chain (ELC) and regulatory light chain (RLC), though they have been called by many names over the decades (). ELC binds the IQ1 motif closest to the motor domain; the RLC binds to the adjacent IQ2 motif.Citation6 The origin of the names ELC and RLC is somewhat curious: The term “regulatory” was chosen as this light chain is involved in the regulation of the enzymatic activity of some myosins-2.Citation7 The term “essential” was chosen because this light chain can only be removed from the myosin-2 heavy chain under harsh conditions in which the enzymatic activity of the holoenzyme is lost.Citation8 The specificity of the IQ1 and IQ2 motifs in class-2 myosins is high for ELC and RLC respectively, as demonstrated in the bay scallop Argopecten. The RLC from this species can be gently removed from IQ2 by treatment with the divalent cation chelator EDTA. The unoccupied IQ2 binds the RLCs from virtually any species presented to it in vitro, but does not bind ELCs.Citation7
Table 1. List of the commonly used historical and recent abbreviation for ELCs, RLCs and CaM in different organisms as found in the NCBI database
Myosins from all other classes are referred to as “unconventional.” Unconventional myosins typically do not form filaments and display a great diversity in their tail domains that determines their localization and interaction with binding partners.Citation1,9 The number of IQ motifs found in the neck region of unconventional myosin heavy chains is variable and not always conserved within a class.Citation10
In only a few cases has the native light chain composition of unconventional myosins been experimentally determined by co-purification coupled with proteomic or immunological analysis. Among those, myosin-1A from brush border was the first vertebrate unconventional myosin purified to homogeneity. It binds 3 molcules of the ubiquitous Ca2+-binding protein CaM as light chains.Citation11,12 Subsequently, all other vertebrate myosin-1 paralogs purified from various tissues were shown to bind CaM.Citation13,14,15 In contrast, the light chain composition of lower eukaryotic myosins-1 is more complex, as outlined below. Myosin-5A purified from chicken brain is associated with CaM and ELC, although the stoichiometry and binding order of the light chains to its 6 IQ motifs has not been firmly established.Citation16 In contrast, myosin-5A from mouse brain purifies exclusively with CaM as light chain.Citation17 Based on those early studies, a central dogma was established that conventional myosins bind ELC and RLC, whereas most unconventional myosins bind CaM as light chain.
Subsequently, less stringent methods were used to show that CaM is a light chain of less abundant or more difficult to purify myosins from various sources. These methods include (i) immunoprecipitation of the myosin heavy chain from cell or tissue extracts followed by probing for the presence of CaM in western blots or the detection of Ca2+-dependent CaM gel shifts, (ii) CaM overlay assays of blots of cell or tissue extracts which are sometimes partially enriched for the myosin heavy chain by chromatography or (iii) binding of the myosin heavy chain to a CaM-affinity column. All methods demonstrate that CaM can bind to a particular myosin heavy chain, but do not establish that CaM is the sole or preferred light chain since protein gel blots usually were not probed with antibodies to other light chains and gels were usually too lightly loaded for the direct detection of potential light chains by protein gel staining methods. Likewise, most of these studies did not quantify the molar ratio of the light chains to the heavy chain.
By using various combinations of these methods, myosins-3 from Limulus and Drosophila,Citation18-20 myosin-6 from a porcine proximal tubule cell line,Citation21 myosin-7A from mouse and bovine tissues,Citation22,23 and myosin-9B from human leukocytesCitation24 were shown to bind CaM as light chain (). More recently, myosin-18A was co-immunoprecipitated with the RLC from COS7 cellsCitation25 and a class-14 myosin, Toxoplasma myosin-A, copurified with an RLC (TgMLC1) from recombinant parasites.Citation26 Notably, the latter myosin does not copurify with CaM.Citation26
Table 2. List of selected myosin heavy chains and identified light chains discussed in this review
Finally, there are numerous studies in which a particular unconventional myosin heavy chain or its fragments are recombinantly overproduced with CaM in the baculovirus/Sf9 system, despite a lack of information of the native light chain composition. Based on this approach, previously uncharacterized myosins from classes-1, -3, -5, -6 -7, -9, -10, -14, -15, -19 and -21 were shown to be capable of binding CaM (). However, copurification does not mean that CaM is the native light chain for these myosin heavy chains as most reports do not state whether preliminary experiments were pursued to establish that CaM binds competitively with other light chains. Thus, CaM may not preferentially bind to the myosin heavy chain if coproduced along with another or the native light chain.Citation27 This hypothesis is supported by occasional reports of sub-stoichiometric CaM-binding ratios in preparations of recombinant unconventional myosin heavy chain constructs in which the addition of exogenous CaM increases the saturation of the IQ motifs, the enzymatic activity or the solubility of the myosin holoenzyme at low concentration.Citation28,29,30 These results led to the realization that CaM might not be a universal light chain for all unconventional myosins.
As outlined in greater detail below, recent reports, primarily utilizing co-expression of heavy chain and light chains genes in the baculovirus/Sf9 system, show that RLC may be a physiological subunit of unconventional myosin holoenzymes from classes-14, -15, -18 and -19, ELC a subunit of unconventional myosins from classes-1, -5, -6, -7, -14, -15 and -18 ().
Myosin Light Chains Have a Conserved Structure
Three independent genes, CALM1, CALM2 and CALM3 encode 3 identical CaM proteins in humans. CaM is a major intracellular Ca2+-sensor interacting with more than 3 hundred protein targets.Citation31 CaM is extremely conserved through phylogeny and has not evolved since the appearance of vertebrates.Citation31 CALM1–3 gene products have 148 amino acids with a molecular weight of 16 kDa. CaM has a conserved overall dumbbell-shaped structure consisting of an N-terminal (N-lobe) and a C-terminal (C-lobe) globular region separated by a central helix ().Citation32
Figure 2. Structure of CaM and its EF-hand motifs in various conformations. (A) Structures of CaM (blue) bound to a target sequence (gray) in the extended (PDB ID: 3CLN) and compact conformation (PDB ID: 2IX7). (B) EF-hand conformations found in CaM. Top row: EF-hand in an open (PDB ID: 1CLL) or closed (PDB ID: 1CFC) conformation. Bottom row: The semi-open (PDB ID 2IX7) and the uncoupled conformation of the EF-hand motif (PDB ID 4R8G). is adapted from Lu et al.Citation34
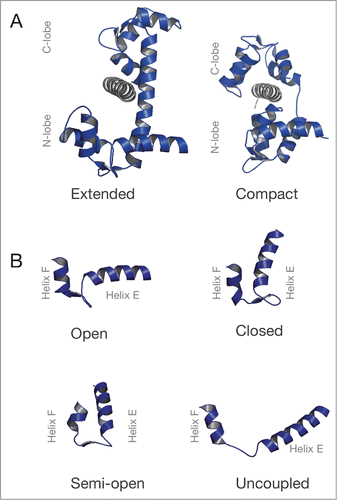
Each CaM lobe contains 2 functional helix-loop-helix motifs termed EF-hands consisting of 2 helices connected by a Ca2+-binding loop which provides coordinating oxygen or nitrogen atoms derived from the amino acid side chains or the backbone polypeptide chain. The EF-hands of CaM bind Ca2+ with high affinity and specificity compared to other divalent cations. The C-lobe has a 10-fold higher Ca2+-binding affinity (Kd ∼0.2 µM) than the N-lobe (Kd ˜2 µM).Citation5 Ca2+-binding or the interaction with a protein target can alter the conformation of the EF-hands. For example, in the absence of Ca2+ (apo), the 2 EF-hand helixes form an antiparallel, “closed” conformation. In the presence of Ca2+, the EF-hand helixes are perpendicular, the conformation “open.”Citation33 The EF-hands can also adopt a “semi-open” conformation, which is between the open and the closed conformation. The EF-hand helices are completely separated in the “uncoupled” conformation as recently described.Citation34 A compilation of EF-hand conformations found in myosin light chains is schematically depicted in . Conformational changes in the CaM EF-hand pair in a CaM lobe exposes a hydrophobic cleft which can be solvent exposed in the “open” conformation, but buried in the “closed” conformation, “semi-open” or “uncoupled.” The open, semi-open and uncoupled lobe conformation can interact with a target sequence such as the IQ motif of the myosin heavy chain, whereas the closed lobe conformation does not allow for a tight interaction.
When bound to the myosin heavy chain, the overall structure of CaM can be either compact (“canonical”) or extended (). In the compact conformation, the CaM N- and C-lobe interact with the myosin heavy chain whereas only the CaM C-lobe interacts strongly with the IQ motif when bound in the extended conformation. The extended light chain conformation results in one free light chain lobe, which is predicted to recruit binding partners in some cases and potentially determines the localization of the ternary complex.Citation26,35-38
The human RLC gene family includes MYL2, MYL5, MYL7, MYL9, MYL10, MYL11, MYL12A, MYL12B, the ELC gene family MYL1, MYL3, MYL4, MYL6 and MYL6B. RLC gene products have a high structural similarity to CaM, 172–226 amino acids and molecular weights ranging from 18.8–25.3 kDa.Citation10 RLCs are comprised of 4 EF-hand motifs, although some may be degenerated since they lack the requisite number of Ca2+-coordinating ligands. Degenerated EF-hand motifs have partially or entirely lost their Ca2+-binding properties or bind divalent cations in general with little selectivity toward Ca2+.
ELCs have 4 EF-hand motifs with some being degenerated, 150–208 amino acids and molecular weights ranging from 16.9–22.8 kDa.Citation10 Expression of ELCs from genes MYL1 and MYL3 as well as RLC from genes MYL2 and MYL7 is confined to cardiac tissue in human adults, MYL4 expression to the embryonic heart. Major expression of the RLCs from genes MYL5 and MYL11 is described in skeletal muscle. MYL10 encodes for an RLC that might be a lymphocyte specific precursor. Light chains believed to be nonmuscle myosin-2 specific are gene products of MYL6 and MYL6B as well as MYL9, MYL12A, MYL12B and have a vast, however isoform-specific tissue distribution, as described below. How a light chain recognizes the appropriate myosin heavy chains is not well understood. Based on double-epitope tagging competition experiments, Komiyama et al. propose that the ELC N-lobe is responsible for intracompartmental and isoform-specific sorting.Citation39 More specifically, another study using ELC chimeras added evidence that the second EF-hand motif of the N-lobe determines isoform-specificity for the myosin heavy chain.Citation40
Alternate splicing of the ELC genes MYL1, MYL3, MYL4, MYL6, MYL6B and the RLC gene MYL9 is reported and increases light chain diversity at the protein level. The significance of alternate splicing events of myosin light chain pre-mRNA and differences in isoform composition are largely unknown but are associated with pathological disease states in some cases, as discussed later. Further, alternate splicing events and resulting changes in ELC expression levels can be age-dependent as seen in skeletal muscle.Citation41
In fast skeletal muscle and the myocardium, MYL3 and MYL4, are alternatively spliced at the N-terminus which results in an ∼40 amino acid long extension. This extension contains a cluster of positively charged residues which interact with acidic residues on F-actin, thereby bridging the latter and the myosin-2 motor domain.Citation42,43 This may be responsible for the differences in the mechanoenzymatic activity of myosins-2 containing one or the other ELC which is only seen in the presence of F-actin and crucial for the alignment of the sarcomeric myosin-2 heavy chain on F-actin for optimal force production.Citation44,45 The long N-terminal extension of some invertebrate RLCs is thought to have similar functions.Citation46 Truncation of the N-terminal extension from Drosophila indirect flight muscle myosin regulatory light chain (DMLC2) reduces flight ability, wing beat frequency and the frequency of maximum power output.Citation47,48 Further, DMLC mutant males generate a courtship song with altered song parameters and exhibit impaired mating behavior when compared to control males.Citation47
The regulatory function of the N-terminal ELC extension on muscle performance is vertebrates is myosin-specific: Removal of the N-terminal ELC extension in the cardiac myosin-2 holoenzyme affects cross-bridge prepositioning which negatively affects contractility by generating lower force and a pathological phenotype in a mouse model.Citation45 In contrast, an artificial increase of the ELC splice variant lacking the N-terminal extension in aged rat skeletal muscle is associated with increased contractility of single muscle fibers.Citation41 Time-resolved FRET studies add evidence that the extent of the force-generating power stroke of fast skeletal muscle myosin-2 strongly depends on the ELC splice variant in its lever arm.Citation49 The extent of the myosin power stroke is smaller in the presence of an ELC splice variant that lacks the N-terminal extension when compared to myosins that associate with the ELC that harbors the N-terminal extension.Citation49 The results provide a structural explanation for the effect of the ELC splice variant on the contractile function of muscle and supports the hypothesis that the N-terminal ELC extension enhances the isometric force while slowing the speed of shortening.Citation49
Disease States Associated with ELC and RLC
Familial hypertrophic cardiomyopathy is the most common inherited heart disease and linked to mutations in genes encoding many sarcomeric proteins including the β-cardiac myosin-2 heavy chain (MYH7), ELC (MYL3) and RLC (MYL2).Citation50 Eight mutations in the RLC and 5 mutations in the ELC have been linked to the disease in humans. ELC mutations cluster around 2 of the EF-hand motifs in the ventricular ELC. Their pathological phenotypes vary significantly in severity and mostly result in sudden cardiac death at young age.Citation51 In vitro studies suggest that RLC mutations R58Q and N47K reduce the isometric force and power output. The load at which peak power is achieved is shifted toward lower loads due to changes in the strain-dependence of β-cardiac myosin-2.Citation52 The effect of light chain mutations on cardiac myosin-2 and muscle properties have been well studied and the reader is directed to 2 recent reviews for an in depth discussion.Citation51,53 MYL3 mutations are further linked to a rare myopathy of skeletal muscle.Citation54
Besides mutations in light chain genes, altered splicing events or intracellular levels of light chain proteins are associated with human disease states. An example for the former includes aberrant splicing of MYL6 pre-mRNA which is causative for altered contractility of smooth muscle myosin-2 in the corpus cavernosum in patients with erectile dysfunction.Citation55 An example for the latter is the post-transcriptional down-regulation of MYL7 in chromosome 21 chimeric mice and neonatal hearts from humans with Down syndrome which is associated with developmental abnormalities in early stages of cardiogenesis. Altered MYL7 expression and the associated downregulation of the protein level plays a key role in congenital heart disease observed in Down syndrome patients.Citation56 Down-regulation of MYL9 expression is associated with the development and metastasis of non-small cell lung cancer.Citation57
Structural Aspects of the Myosin Heavy Chain: Light Chain Interaction
The crystal structure of chicken skeletal muscle myosin-2–2IQ reveals that the N-terminus of the myosin heavy chain forms a globular motor domain and results in an extended α-helical neck domain. It harbors the 2 IQ motifs which bind the ELC and the RLC.Citation6 This sequence of domains is conserved in other myosin-2 molcules as is illustrated in the structure of scallop myosin-2 (). The neck domain with the bound light chains is also referred to as the lever arm. The myosin light chain:heavy chain interaction is critical at several levels. First, the 2 light chains are strictly required for maintenance of the structural integrity of the myosin holoenzyme and are critical to myosin function. Ablation or the reduction of the RLC expression levels leads to intracellular aggregation of the nonmuscle myosin-2 heavy chain in Drosophila.Citation58,59 Second, the myosin neck region with the bound light chains acts as a rigid lever arm that amplifies movements within the myosin motor domain into a large mechanical stroke that directionally propels the myosin along the actin filament. Evidence for this comes from in vitro motility and single molecule optical trapping studies with recombinant mutant myosins where the length of the lever arm is artificially extended, shortened or replaced with artificial lever arms.Citation60-63 Removal of a light chain from the myosin heavy chain exposes the hydrophobic IQ region to the solvent and likely results in a collapse of the α-helical neck domain. Myosins lacking the appropriate number of light chains to saturate the IQ motifs typically have a compromised power stroke but do exhibit actin-activated ATPase activity.Citation28,64,65 To circumvent this problem in functional and kinetic studies of conventional and unconventional myosins in which the native light chain composition has not been firmly established or substoichiometric light chain binding is observed, an artificial lever arm that replaces the native neck region can be used as powerful surrogate.Citation66,67 Note as will be discussed later, there is evidence that the light chain bound to IQ1 may interact with structural elements of the motor domain and influences the kinetic or mechanical properties of some myosins.Citation68 Thus caution must be exercised in truncating the myosin heavy chain, in selecting a light chain or in the use of artificial lever arms.Citation29,44,68
Third, light chains bound to the neck region of some myosins-2 are intimately involved in the regulation of its enzymatic and mechanical output as discussed later.
Structural plasticity about the central helix of myosin light chains allows them to adopt several binding conformations on different IQ motifs. For example, ELC in a compact conformation wraps around the IQ1 of myosins-2. The C-lobe is in the semi-open conformation and interacts with the N-terminal half of the IQ1. The ELC N-lobe is in the closed conformation and interacts with the C-terminal half of the IQ1 ().Citation6,69,70 The comparison between chicken skeletal muscle and smooth muscle myosin-2 crystal structures shows that the C-lobe of the ELC further interacts with the very N-terminus of the heavy chain or with a surface loop of the myosin motor domain depending on its nucleotide state, suggesting a communication between both subunits of the holoenzyme. This interaction could modulate the enzymatic activity by favoring or disfavoring conformations of the myosin motor domain during its ATPase cycle.Citation6,70 The ELC also establishes extensive interactions between its C-lobe and the myosin motor domain of the heavy chain that persists in all conformational states of the lever arm.Citation6,69,71
The RLC interacts in an extended conformation with the IQ2 of conventional myosins where the C-lobe of the light chain interacts strongly with the N-terminal half of the IQ2 motif in the semi-open conformation.Citation6 The RLC N-lobe is in the open conformation and interacts with a conserved WXW motif immediately C-terminal to the IQ2, which is also termed the “hook” region ().Citation69,72 All myosin-2 heavy chains bend sharply in the hook region. In crystal structures of myosin-2 from the sea scallop Placopecten, the heavy chains adopt different angles about the hook region which involve a 10 Å variation in the length of the light chain binding region.Citation73 The mechanical relationship between the angle of the hook region and the RLC conformation might account for some of the proposed compliance in the myosin molecule related to its force generating capabilities and be important for the regulation of myosin-2 function.Citation73
Crystal structures of unconventional myosins-1, -5, -6 highlight different interaction modes between the myosin heavy chain and the associated light chain which is usually CaM.Citation35,68,69,74 In most CaM:IQ interactions, CaM adopts a canonical conformation in which the C-lobe is in the semi-open conformation and the N-lobe remains closed and interacts weakly with a hydrophobic residue in the 8 position of the IQ motif.
A myosin-1B-1IQ structure demonstrates an interaction between CaM and the very N-terminus of the myosin heavy chain that crucially tunes the enzymatic activity and force sensitivity of the holoenzyme.Citation68 An identical fragment of myosin-1C reveals an interaction between the CaM C-lobe and a portion of the myosin motor domain involved in the communication with the nucleotide binding site. Ca2+-binding to CaM may alter this interaction and account for the Ca2+-dependence of some transient-kinetic parameters of myosin-1C's ATPase cycle which could have important implications for its role in the adaptation of the auditory system.Citation76,77 The crystal structure of a myosin-1C fragment containing its 3 IQ motifs and the remainder of the tail reveals an unprecedented mode of CaM:heavy chain interaction. The CaM C-lobe interacts with IQ3 in a semi-open conformation while the N-lobe interacts with a downstream region termed the “post-IQ” motif.Citation34 This interaction greatly distorts the CaM N-lobe compared to its canonical conformation and results in a strong bending of the structure of the tail at this region. In this sense, the CaM:heavy chain interaction resembles the interaction of the RLC with the IQ2 motif and the hook region of myosins-2.Citation69 However, the CaM bound to IQ3 of myosin-1C has a uniquely uncoupled EF-hand motif (), raising the possibility that CaM and other Ca2+-binding EF-hand proteins may recognize additional targets via uncoupled EF-hands.Citation34 Ca2+:CaM binding to IQ3 and the post-IQ induces a flexibility change in the myosin-1C tail which is predicted to alter the mechanoenzymatic properties of the motor.Citation34 In contrast, CaM interacts with the first 2 IQ motifs in myosin-1C in the canonical, compact conformation: the N-lobe is closed, the C-lobe semi-open.Citation34
For chicken myosin-5A it is still not known which IQ motif binds ELC. A crystal structure of the motor domain and the IQ1 motif in complex with a truncated ELC shows that IQ1 is capable of binding an ELC,Citation74 however another study showed that IQ1 is also capable of binding CaM.Citation29 Likewise, a proteolysis study of tissue purified chicken brain myosin-5A indicates that the ELC does not bind to either IQ1 or IQ2.Citation78 Notably, the chicken myosin-5A-1IQ:ELC and the myosin-5A-1IQ:CaM complex display slightly different kinetic parameters.Citation29 A subsequent structure of a peptide corresponding to IQ1 and IQ2 of mouse myosin-5A complexed with CaM reveals that IQ1 can bind CaM as well.Citation79 Structural comparison indicates that ELC and CaM adopt a very similar, semi-open conformation of the C-lobe that allows gripping of the first portion of the IQ1 motif in the absence of Ca2+. The N-lobe is closed and interacts weakly with the last portion of the IQ1 motif. Interestingly in the IQ1-IQ2 peptide, the fine structures of the 2 CaM molecules are strongly influenced by the non-conserved residues of the IQ motif. Treatment of myosin-5A with physiological and higher concentrations of Ca2+ results in dissociation of at least one CaM per heavy chain. This mechanism has been used by many investigators to insert a fluorescently labeled CaM into the lever arm for single molecule motility studies.Citation62,80 Several studies provide evidence that the IQ2 motif is most likely to release its CaM in the presence of Ca2+.Citation78,81
Of peripheral interest, alternate splicing of the myosin-5A tail domain results in melanocyte- and brain-specific isoforms. The brain-specific isoform binds the low molecular weight (8 kDa) dynein light chain (DLC8, DYNLL1) which has also been shown to interact with other proteins, notably dynein.Citation16 DLC8 is not a CaM gene family member, does not bind to an IQ motif and should not be mistaken for a myosin light chain.Citation82
Two studies examined the interaction of IQ motifs from a yeast myosin-5 (Myo2p) with the ELC (Mlcp1). Mlcp1 binds to IQ2 or to a peptide corresponding to IQ2-IQ3 in a compact conformation in which each of the light chain lobes interacts with the heavy chain: The C-lobe interacts with the N-terminal half of the IQ motif, the N-lobe with the C-terminal half of the IQ motif.Citation35 In contrast, Mlcp1 interacts with IQ4 in an extended conformation where only the C-lobe of the light chain interacts with the IQ motif and the N-lobe is free.Citation35 These 2 disparate interaction modes of Mlcp1 with 2 different IQ motifs from yeast Myo2p coupled with the compact binding of ELC to IQ1 of myosins-2 demonstrates that the sequence of the IQ motif plays a significant role in determining the light chain binding mode.
Whether or not a light chain adopts a compact or extended conformation may depend on the amino acid presence at position 7 of the consensus IQ motif (). A glycine residue in this position is predicted to favor the compact light chain conformation whereas the presence of a residue with a more bulky side chain such as arginine, lysine or methionine favors the extended conformation.Citation36 This is consistent with the observation that IQ4 from Myo2p and the IQ2 from all conventional myosins bind Mlcp1 or RLC in an extended conformation, respectively. Based on the crystallographic data, Terrak et al. propose a model for the yeast Myo2p neck domain which binds 2 of the 6 light chains in the extended conformation, and predict from sequence analysis that vertebrate myosin-5A would also have a subset of its light chains bound in the extended conformation.Citation36 However, a cryo-electron microscopy structure of the myosin-5A lever arm was best fit with crystal structures of light chains bound in the compact conformation to the IQ motifs.Citation83
Myosin-6 is unique in that it moves in the opposite direction on F-actin than any other myosin.Citation84 Direction reversal is mediated by an unconventionally bound CaM to a unique sequence insertion (“insert-2”) in between the motor domain and its only IQ1 motif.Citation75 The unconventional heavy chain:CaM interaction involves 4 hydrophobic residues from the heavy chain spaced at a 1–6–14 amino acid interval. This interaction effectively repositions the lever arm so that the power stroke moves in the opposite direction. The CaM associated with insert-2 tightly binds 4 Ca2+ ions.Citation27 The sole canonical IQ1 motif in the myosin-6 heavy chain binds apo-CaM.Citation27,75 Recent reports present evidence for an unfolding event of a 3-helix bundle located downstream of IQ1 into a long α-helix which exposes an additional CaM binding site. Association of CaM with this site efficiently stabilizes and extends the myosin-6 lever arm.Citation75,85
Taken together, the crystallographic findings suggest that sequence variations in both, the target IQ motif and sequences up- and downstream of it determine light chain selectivity, specificity and binding mode.Citation35 Whether the light chain binding mode in individual myosins is related to mechanoenzymatic properties remains mostly elusive.
Involvement of ELC and RLC in the Regulation of Conventional Myosins
Intrinsically regulated myosin-2 holoenzymes are responsive to RLC phosphorylation or Ca2+-binding to the ELC.Citation86 In cells, the level of smooth and nonmuscle myosin-2 RLC phosphorylation is controlled by a complex equilibrium between kinases that phosphorylate the RLC such as myosin light chain kinase (MLCK), rho-associated coiled coil-containing kinase (ROCK), zipper-interacting protein kinase (ZIP), protein kinase C (PKC) and myosin phosphatase which dephosphorylates the RLC. Myosin kinases and phosphatase are downstream effectors of various signal transduction pathways which tightly regulate RLC phosphorylation levels.Citation87,88
Phosphorylation of the RLC at S19 is a major regulatory mechanism of vertebrate smooth and nonmuscle myosins-2. It serves as an on/off-switch of the actin-activated ATPase activity of the holoenzyme and regulates filament assembly. Myosin-2 is enzymatically inactive in the absence of RLC phosphorylation.Citation89,90 In the presence of ATP, it assumes a conformation in which the 2 heads make an asymmetric interaction and the tail folds back upon itself in 2 places to form a compact conformation that inhibits myosin-2 filament formation ().Citation91,92 RLC S19 phosphorylation is associated with the adoption of an extended conformation and an increase in the actin-activated ATPase activity of the holoenzyme, the ability to translocate F-actin in the in vitro motility assay and the formation of filaments, as reviewed previously.Citation87,93 The significance of the regulation of filament assembly is clear in motile nonmuscle cells or in dividing cells in which the cytoskeleton must be continually remodeled. In contrast, it is controversial whether filament assembly is regulated by RLC phosphorylation in smooth muscle tissue.Citation94
Figure 3. Structural consequences of RLC phosphorylation on the conformation of smooth and nonmuscle myosins-2. RLC phosphorylation at S19 promotes a conformational change from a compact (left) to an extended conformation of the myosin holoenzyme (left), which readily assembles into higher order bipolar filaments (not shown). The structural transition from the compact to the expended conformation is accompanied with an increase in the mechanoenzymatic activity. RLC dephosphorylation by myosin phosphatase reverses the conformational change. Figure adapted from Heissler and Manstein.Citation87
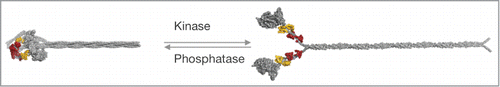
Di-phosphorylation of the RLC at T18 and S19 further increases the actin-activated ATPase activity via an effect on the F-actin affinity and results in increased filament formation of the myosin-2 holoenzyme when compared to S19 mono-phosphorylation of the RLC.Citation95-97
The first 24 amino acids of the N-terminus of the RLC have been termed the phosphorylation domain (PD). The PD is rich in positively charged amino acidsCitation98 and modeling studies assume that its structure is either fully α-helical or consists of 2 α-helices.Citation98,99 Molecular interactions between the PD, the heavy chain and ELC required to adopt the off-state of the myosin holoenzyme are not well characterized, in part because the PD is not resolved in any crystal structure. Smooth muscle myosin-2 that binds a truncated RLC that lacks the first 16 amino acids of the PD adopts the off-state, suggesting that the N-terminal portion of the PD is required for the activation of the myosin holoenzyme.Citation100 In line with this finding, several positively charged residues, in particular K11 and K12, have been shown to be important for the activation of smooth muscle myosin-2s ATPase activity following S19 phosphorylation.Citation100,101
Time resolved FRET studies suggest that the PD exists in 2 states – “open” or “closed” – characterized by the positions of probes placed in the N-terminal PD and the C-lobe of the RLC.Citation98 These studies demonstrate that the probe separation distance increased by 2 nm upon RLC phosphorylation. Molecular dynamic simulations of RLC phosphorylated smooth muscle myosin-2 predict the formation of a salt bridge between the phosphoryl group of S19 and R16 in the RLC that may stabilize the open, active conformation of the PD.Citation98,99 However, mutation of R16 to a neutral or acidic amino acid still allowed RLC phosphorylated smooth muscle myosin-2 to be activated.Citation100
A recent study demonstrates a prominent role of the ELC in the regulation the mechanoenzymatic properties of smooth muscle myosin-2. Charge reversal mutations E10K and E13R of the ELC have an inhibitory effect on the actin-activated ATPase activity and motile behavior of RLC phosphorylated smooth muscle myosin-2.Citation99 This study suggests that interactions between this region of the ELC and the positively charged residues of the RLC PD are essential for stabilization of the active myosin-2 conformation. Another study pointed out the importance of the interaction between the 2 light chains as well as the heavy chain for regulation.Citation102
The RLC of vertebrate smooth and nonmuscle myosins-2 is also a substrate for PKC which phosphorylates S1 or S2 in vivo. In vitro, PKC phosphorylates these 2 residues in addition to T9, though T9 phosphorylation of the RLC is not observed in vivo.Citation103,104 In vitro studies show that PKC phosphorylation of the RLC is inhibitory toward myosin-2 activation in 2 ways. First, myosin-2 RLC phosphorylated at the PKC sites as well as at S19 has the same maximal actin-activated ATPase activity, but requires more F-actin for half maximal activity. Second, myosin-2 pre-phosphorylated at the PKC sites is a poorer substrate for MLCK which might result in an overall decrease in RLC S19 phosphorylation in cells – under the assumption that phosphatase activity is unaffected. Notably, most in vitro studies were carried out on myosin-2 with the RLC phosphorylated at both the S1/S2 site and the T9 site. How T9 phosphorylation affects these kinetic parameters is not fully known, although one study suggests that T9 and not S1/S2 phosphorylation is inhibitory toward RLC S19 phosphorylation MLCK.Citation105 In vivo, S1/S2/T9 phosphorylation does not seem to be a substantial regulatory mechanism of nonmuscle myosin-2 function in HeLa cells and human primary keratinocytes, though a recent paper links the ability of fibroblasts to chemotact to phosphorylation at the PKC sites.Citation106,107
The RLCs of vertebrate skeletal and cardiac muscle myosins-2 are also phosphorylated at their N-termini by MLCK, but phosphorylation is only modulatory with respect to the enzymatic and motile activities of the holoenzymes in vitro and their mechanical properties in muscle.Citation108-110 Vertebrate skeletal muscle myosin-2 RLC phosphorylation increases the post-tetanic twitch potentiation, increases Ca2+-sensitivity of contraction and enhances the transition from a nonforce to a force-producing state.Citation110-112 However, RLC phosphorylation does not alter the maximum velocity of shortening or maximal isometric force.Citation113 Electron microscopy studies show that the heads of unphosphorylated skeletal muscle myosin-2 thick filaments are highly ordered with the heads held close to the filament backbone. Following RLC phosphorylation, the heads release from the thick filament backbone.Citation114 Vertebrate cardiac myosin-2 RLC phosphorylation is modulatory and associated with a 3-fold increase in force production and a 7-fold increase in peak power output.Citation115 RLC phosphorylation is reduced in hypertrophic hearts.Citation116,117 Interestingly, overexpression of the skeletal muscle MLCK gene in mouse cardiac tissue attenuates the isopreternol-induced hypertrophy. This finding makes small molecules that target the RLC phosphorylation pathways in cardiac tissue attractive for the treatment of hypertrophic cardiomyopathy.Citation118
Similarly, the RLC of myosin-2 from the lower eukaryote Dictyostelium is also phosphorylated at the N-terminus, but this phosphorylation is only modulatory with the primary regulation of ATPase activity being associated with phosphorylation of residues in the tail domain.Citation119 The enzymatic activity of myosin-2 from fission yeast Schizosaccharomyces is independent from RLC phosphorylation in vivo and in vivo.Citation120,121
RLC phosphorylation in striated muscle myosins-2 from the horseshoe crab Limulus as well as Tarantula is associated with regulation of the ATPase activity in an on/off manner, similar to the effect of RLC phosphorylation of vertebrate smooth and nonmuscle myosins-2.Citation122-126 The structure of native unphosphorylated Tarantula thick filaments in the off-state demonstrates that the myosin heads are held in close apposition to the thick filament backbone and form the same asymmetric head-head interaction observed in the off-state of smooth and nonmuscle myosin-2.Citation125
Ca2+ regulates the ATPase activity of molluscan myosin-2.Citation127 In vitro, RLC dissociation from scallop muscle myosin-2 is associated with a loss of Ca2+-dependent regulation and Ca2+-binding. However, the ATPase activity of the holoenzyme is elevated even in the absence of Ca2+, indicating that the RLC is necessary for the off-state.Citation128 The crystal structure of the scallop myosin-2 “regulatory domain,” containing both IQ motifs and light chains, revealed a surprise as Ca2+ does not bind to the RLC, but rather to the first EF-hand of the ELC.Citation129 This degenerated EF-hand is not predicted to bind Ca2+ on its own based on the lack of appropriate numbers of coordinating ligands. However, when the ELC is bound to the myosin heavy chain, the conformation of its EF-hand is altered via a direct interaction between F20 and R24 of the ELC with G117 of the RLC creating a functional Ca2+-binding site on the ELC by rotating a carbonyl oxygen in the first EF-hand into a Ca2+-coordinating position.Citation69,129 The importance of this interaction is confirmed by mutagenesis of G117 of the RLC which abolishes Ca2+-binding at this site.Citation130
Crystal structures of the scallop myosin-2 regulatory domain provide valuable mechanistic insights into the interaction between Ca2+ and the ELC and the function of the Ca2+:ELC interaction in the activation of the myosin holoenzyme. Further, the structures allow for speculations about similarities in the activation mechanisms of myosins-2 that are intrinsically regulated by Ca2+-binding or RLC phosphorylation.Citation131 It is likely that profound structural similarities between the regulatory domains of scallop and smooth muscle myosins-2 exist since it was shown that the ATPase activity of a chimeric myosin containing the scallop myosin-2 heavy chain and ELC with a vertebrate smooth muscle RLC could be activated either by Ca2+ binding or by phosphorylation of the RLC.Citation132 In both smooth muscle and scallop myosin-2 RLCs, several C-terminal residues, in particular K149 in scallop and K163 in smooth muscle, are required for the intrinsic regulation.Citation133-135 A lysine at equivalent positions is exclusively present in RLCs from intrinsically regulated myosins-2, but is not found in RLCs from myosins-2 that are not regulated by RLC phosphorylation such as the skeletal muscle myosin-2 RLCs. K149 forms a hydrogen bond with T83 in the Ca2+-bound scallop myosin-2 regulatory domain structure which, when broken in the Ca2+-free state, may help reduce the rigidity of the regulatory domain. It is also observed that fewer light chain:heavy chain interactions are present in the Ca2+-free state.Citation131 The more flexible regulatory domain may be more conducive to bending in order to adopt the off-state. In this regard, it is of note that the lever arm of solely one of the 2 heads of smooth muscle myosin-2 in the off-state shows a strong bend between the ELC and the RLC. This bend is not found in the other head or in the structure of the phosphorylated state.Citation136 Such bending may be necessary for the adoption of the head-head interaction observed in the myosin-2 off-state ().Citation91
Interestingly, the conformation of the off-states of vertebrate smooth and nonmuscle myosins-2, which are regulated by RLC phosphorylation, and the Ca2+-regulated molluscan myosins-2 are similar and involve the asymmetric intramolecular interaction between the 2 myosin heads and folding of the long tail domain into thirds.Citation137 Similarly, the asymmetric head-head interactions in the off-state is preserved in vertebrate smooth and nonmuscle myosins-2 as well as myosins-2 from molluscs, Tarantula and Limulus. This suggests that this conformational change is an ancient mechanism for regulating conventional myosins.
Nonmuscle Myosin-2 Heavy Chain Specificity and Tissue Distribution of ELC and RLC
The presence of 3 nonmuscle myosin-2 heavy chain genes (MYH9, MYH10, MYH14 – out of which MYH10 and MYH14 can be alternatively spliced at multiple loci) and 5 nonmuscle myosin-2 light chain genes in mammals create the potential for dozens of nonmuscle myosin heavy-2 chain:light chain combinations. The physiological subunit composition of the nonmuscle myosin-2 holoenzyme in most tissues and subcellular compartments remains elusive because of the lack of information on expression profiles of the myosin heavy chain and light chain subunits at the protein level.
All nonmuscle myosin-2 heavy chain genes have a vast distribution which is tissue-specific and regulated in a developmentally dependent manner.Citation138,139 RT-PCR and northern blot analysis in mice indicates the same for the 3 nonmuscle myosin-2 RLCs: Myl12a and Myl12b are abundantly expressed in most tissues but are absent in the brain (Myl12a) and striated muscle (Myl12b).Citation139 Myl9 is widely expressed with the highest transcription rates in smooth muscle and bladder tissue. Myl12a, Myl12b and Myl9 can interact with all conventional myosin-2s at the protein level but have a strong preference for the nonmuscle myosin-2 heavy chain.Citation139 This is seen for example in immunofluorescence studies on cardiomyocytes where all nonmuscle myosin-2 RLCs colocalize to F-actin stress fibers and the z-lines but do not colocalize with the α- and β-cardiac myosin heavy chains.Citation139 Immunoprecipitation studies indicate that all RLCs form a complex with nonmuscle myosins-2A and -2B in various cell types.Citation139 Similar results were obtained from affinity-purification mass spectrometry assays of human cell lines, indicating the heteromeric interaction between nonmuscle myosins-2A and -2B with the closely related myosin-18A and the association of the aforementioned myosins with both RLCs, MYL12A and MYL12B.Citation140 Nonmuscle myosins-2A and -2B interact with both ELC isoforms (MYL6 and MYL6B), as does myosin-18A. In contrast, nonmuscle myosin-2C, smooth muscle myosin-2 and unconventional myosin-1C interact exclusively with the ELC isoform MYL6.Citation140 Those data imply complementary and combinatorial expression patterns for the nonmuscle myosin-2 heavy and light chains. The physiological significance of the variations in light chain subunits of the nonmuscle myosin-2 heavy chain remains largely elusive.
CaM as Subunit of Unconventional Myosin Holoenzymes
Many unconventional myosins bind CaM as light chain(s). The binding affinity between the myosin heavy chain IQ motif and CaM can be either strengthened or weakened by the presence of Ca2+ and, in the extreme, result in the dissociation of the complex. In vitro, this may regulate myosin's mechanical properties since it could affect the stiffness of the lever arm. As a caveat, it is not clear yet whether the Ca2+-induced dissociation of CaM from the myosin heavy chain is a significant regulatory mechanism for the mechanoenzymatic properties of the holoenzyme in vivo.
Ca2+-regulation has been most critically studied in mammalian myosin-5A, a motor that moves processively along actin filaments. Myosin-5A has 6 IQ motifs in its neck which allows it to take 36 nm steps along F-actin.Citation141 In the absence of Ca2+, myosin adopts a folded off-state in which the 2 motor domains contact the 2 lobes of the globular tail domain.Citation83,142 This conformation of myosin-5A has a low affinity for F-actin and exhibits a low ATPase activity.Citation143 In the presence of Ca2+, one or more CaMs dissociate and the myosin extends, freeing the motor domains to interact with F-actin and potently activates the ATPase activity.Citation144-146 As a consequence of the Ca2+-mediated CaM-dissociation from the myosin-5A heavy chain, the molecule is mechanically incompetent since the unoccupied IQ motif does not allow for a force producing power stroke likely due to a “floppy” heavy chain segment.Citation81,147 Thus, it is unlikely that Ca2+ is a direct cellular activator of myosin-5A activity. Instead myosin-5A has been shown to be activated by the cargo adaptor protein melanophilin, which links myosin-5A to Rab27a and to melanosomes in melanocytes.Citation148,149,150 Several studies have shown that the CaM bound to IQ2 of chicken myosin-5A most easily dissociates in the presence of Ca2+.Citation78,151 Those studies support the hypothesis that the affinity of Ca2+-CaM for an IQ motif is dependent on the neighboring IQ motif as bridging may occur, where the 2 CaM domains interact with 2 adjacent IQ motifs.Citation152,153
Ca2+-regulation has also been observed for some vertebrate class-1 myosins. The neck region of myosin-1B is subject to alternative splicing events which result in isoforms with 4 to 6 IQ motifs. Laakso et al. added evidence to the concept that the myosin-1B light chain binding region is indeed a lever arm, by showing that the power stroke size measured by optical trapping increased with the lever arm length.Citation154 Biochemical studies show that myosin-1B splice variants have different kinetic, motile and force-sensing properties. The actin-activated ATPase activity of myosin-1B is activated several fold by Ca2+ whereas some transient kinetic parameters are reduced due to an effect on the IQ1:CaM interaction.Citation155-158 The interaction of myosin-1B with F-actin is very force dependent and its interaction time increases 50-fold when the working stroke is opposed by forces in the pN range.Citation154,159 Ca2+ dramatically shortens the force-dependent attachement lifetimes of myosin-1B.Citation158 The affinity of CaM for the myosin-1B IQ motifs varies significantly from 0.2 µM to 5 µM. This suggests that a subset of its IQ motifs might not be associated with CaM under physiological conditions and might be free or bind to other light chains or proteins.Citation28,154
In line with this finding, Cyr et al. show that CaM interacts very weakly with the IQ4 of unconventional myosin-1C, suggesting that only a fraction of the molecules is associated with CaM in vivo.Citation160 In contrast, IQ1, IQ2 and 1Q3 have stronger CaM affinities.Citation160 At a low intracellular Ca2+ concentration, IQ1 and IQ3 are fully saturated with CaM whereas CaM dissociates from IQ2. The unoccupied IQ2 and potentially CaM-bound IQ1 and IQ3 form a binding site in the myosin-1C heavy chain which allows the interaction with stereociliary receptors.Citation161 A drastic increase in Ca2+ results in a loss of all CaM molecules and abolishes the myosin-1C:receptor interaction which was initially suggested as a mechanism for the adaptation of hair-cell mechanoelectrical transduction important for hearing.Citation161 However, subsequent work shows that the rate of CaM dissociation from myosin-1C is too slow to account for the adaptation rate in stereocelia.Citation162 While Ca2+ has only modest effects on the steady-state actin-activated ATPase activity of myosin-1C, transient kinetic analysis shows that it dramatically decreases the rate of ATP hydrolysis and increases the rate of ADP dissociation from actomyosin.Citation77 Ca2+ also increases the size of the working stroke of myosin-1C perhaps by stiffening or effectively increasing the length of the lever arm.Citation163 It should be noted that Ca2+ has no effect on the working stroke of myosin-1B or myosin-5.Citation147,158
The myosin-9B lever arm contains 4 to 6 IQ motifs that bind CaM when purified from human leukocytes or recombinantly overproduced in Sf9 insect cells.Citation24,164 Myosin-9B contains an additional CaM binding site that resides in an actin-binding surface loop in the motor domain that is highly conserved and unique to class-9 myosins.Citation164 The potential regulatory effect of Ca2+ on the enzymatic activity of myosin-9B is difficult to decipher since a myosin-9B-4IQ construct has distinct mechanoenzymatic properties from a motor domain construct that lacks the IQ motifs with respect to the steady-state ATPase activity and in vitro motile behavior.Citation164 Interestingly, a point mutation in the third IQ motif of human unconventional myosin-9B predisposes to inflammatory bowel disease and might affect the interaction with CaM.Citation165 Similar, a loss of the myosin heavy chain:CaM interaction or impaired CaM binding due to a missense mutation in the fifth IQ motif of unconventional myosin-7A represents a pathomechanism for genetic hearing loss.Citation166
Apart from regulatory functions on the myosin holoenzyme, CaM also determines the oligomeric state of myosin-21 from the parasite Leishmania. CaM-binding to its IQ motif prevents dimerization of 2 myosin-21 heavy chains via a coiled-coil forming sequence. Vice versa, myosin-21 dimerizes in the absence of CaM. Notably, monomeric myosin-21 is mechanoenzymatically active, the dimer inactive.Citation167
Unconventional Myosins as Targets of ELC and RLC
Recent biochemical reports on unconventional myosin heavy chains recombinantly overproduced along with various light chains in the baculovirus/Sf9 expression system, immunoprecipitation and proteomic studies strongly contradict the dogma of the myosin-2 heavy chain specificity of ELC and RLC. ELC and/or RLC may be subunits of the unconventional myosin holoenzyme of classes-1, -5, -6, -7, -14, -15, -18, and -19 ().Citation16,27,140,168-171 The interaction signatures can be identical or distinct from those reported for conventional myosin-2s: ELC and RLC can bind together, independently from each other or in conjunction with CaM and potentially other light chains in various sequential orders to the IQ motifs in the unconventional myosin heavy chain.
As described above, tissue purified myosin-5A from chicken brain is associated with CaM and ELC. In vitro studies with recombinant chicken myosin-5A-IQ1 fragments show that it can bind the ELCs MYL6, MYL6B or CaM, but not RLC.Citation29 In vitro studies show no differences in kinetic parameters under steady-state conditions in the presence of different ELCs. The presence of CaM has subtle effects on the ATPase activity.Citation29 Pre-steady-state kinetics indicate that most parameters are independent from the light chain bound to the IQ motifs, making it unlikely that the light chain isoform determines the functional properties of myosin-5A under physiological conditions as long as the IQ motifs are saturated.Citation29 Notably, mouse myosin-5A fragments do not appreciably bind ELC when overproduced in Sf9 insect cells and native myosin-5A purified from mouse brain does not bind ELC but copurifies with CaM.Citation17 In contrast, Drosophila myosin-5 copurifies with ELC and CaM when recombinantly overproduced with the 2 light chains, however the binding order is unexplored.Citation172 Those findings suggest species- (and tissue)-specific associations of a myosin heavy chain and its light chain subunits, as described above for nonmuscle myosins-2.
The dimeric pseudoenzyme myosin-18A, a close relative of conventional nonmuscle myosins-2, forms a complex with both ELC and RLC when coproduced in the baculovirus/Sf9 system.Citation171 Myosin-18A also co-immunoprecipitates with RLC from COS7 cells.Citation25 The binding sites for the ELC and RLC on myosin-18A are believed to occur in identical (“conventional”) order to those on the myosin-2 heavy chain with the ELC binding to the IQ1 and RLC to IQ2, based on sequence analysis of the 2 IQ motifs.Citation171 Similar, recombinant overproduction of Toxoplasma myosin-A with RLC (TgMLC1) and ELC (TgELC) reveals the formation of a ternary complex. Sequence analysis of the IQ motifs suggests that ELC and RLC bind in conventional order to the myosin heavy chain.Citation169
Recombinant myosin-15A also binds both ELC and RLC. However, the binding order, as determined by expression of differential length fragments, is reversed when compared to myosins-2: RLC binds to IQ1, ELC binds to IQ2.Citation168
Porcine myosin-6–1IQ copurifies in complex with 2 CaMs from Sf9 insect cells when it is solely overproduced with CaM: One CaM binds to the unique insert-2, the second CaM to the adjacent IQ1 motif.Citation27 Myosin-6–1IQ forms a ternary complex with CaM and ELC when coproduced along with both light chains: CaM binds to the unique insert-2, ELC to the IQ1 motif.Citation27 Interestingly, when only the myosin heavy chain is overproduced in the Sf9 cells, myosin-6–1IQ copurifies with endogenous insect cell CaM,Citation27 supporting the hypothesis that the association of a light chain with the IQ motif is driven by affinity and mass action.
The unconventional myosin-19 heavy chain associates with both CaM and RLC subunits.Citation173,174 Interestingly, the myosin-19:RLC holoenzyme binds MYL9 and MYL12B in a 2:1 ratio to its 3 IQ motifs, suggesting differential affinities which determine light chain selectivity and specificity of the IQ motif.Citation174 Independent of subunit composition, both myosin-19 holoenzymes are functional and only display subtle enzymatic differences suggesting that light chain heterogeneity does not automatically imply changes in mechanoenzymatic properties.Citation173,174
The RLCs bound to unconventional myosins are in some cases phosphorylatable at the T18/S19 activation sites by MLCK, but to date no effect of this phosphorylation event on the enzymatic activity of the myosin holoenzyme has been demonstrated.Citation171,174 This suggests that phosphorylation is – so far – an exclusive regulatory mechanism for the aforementioned conventional myosins. Moreover, the presence of Ca2+ does not influence the mechanoenzymatic properties of characterized myosins from classes-14 and -18 which bind ELC and RLC in “conventional” order.Citation169,171
Non-Myosin Binding Partners of ELC and RLC
An increasing number of non-myosin binding partners of ELC and RLC have been discovered in the past years. The interaction partners do not share common features or obvious similarities between IQ motifs. ELC and RLC non-myosin binding partners are involved in cellular functions such as translation, signaling, cell division and the regulation of the actomyosin cytoskeleton; their localizations are cytosolic or membrane-bound. A compilation of selected binding partners can be found in . It remains mostly elusive if the interaction between a non-myosin binding partner occurs between a heavy chain associated or isolated light chain. A pool of free RLC has been demonstrated in cell biological experiments showing that T18/S19 di-phosphorylated RLC localizes independently from S19 mono-phosphorylated RLC and the nonmuscle myosin-2 heavy chain to the midzone during cytokinesis.Citation175 A pool of free light chains or light chains bound to non-myosin binding partners is also proposed after extensive immunoprecipitation studies of the nonmuscle myosin-2 heavy chain in Drosophila.Citation170 The NMDA-type glutamate receptor for example favorably binds free RLC with nanomolar affinity, but does not bind the nonmuscle myosin-2 heavy chain associated RLC. This suggests that the NMDA:RLC interaction is not a mechanism for localization of nonmuscle myosin-2. RLC phosphorylation disrupts the interaction with NMDA-type glutamate receptor. Disruption of the NMDA-type glutamate receptor:RLC interaction results in a delay in delivery of the receptors to the cell membrane, indicating trafficking functions.Citation176
Table 3. Compilation of selected myosin and non-myosin binding partners of ELC and RLC. It is of note that not all kinases and potentially phosphatases that interact with the RLC are listed due to space limitations
If non-myosin binding partners interact with the myosin heavy chain bound light chain, then 2 different scenarios are plausible: Transient ternary complexes can be formed with the conventional or unconventional myosin holoenzyme, similar to the interaction of the nonmuscle myosin-2 holoenzyme with kinases and phosphatases. An example for such a ternary complex is the CaM-mediated interaction between the G-protein RalA and the unconventional myosin-1C heavy chain during the insulin-stimulated glucose uptake.Citation37
As discussed above, light chains can bind in an extended conformation to the IQ motif which could allow the free N-lobe to simultaneously interact with another protein, thereby forming a ternary complex. A CaM-mediated dimerization has for example been shown for the Ca2+-activated K+ channel.Citation177 In the case of myosin light chains, such an interaction would result either in a ternary complex formed by a myosin holoenzyme and a non-myosin binding partners or potentially the exclusive light chain-mediated dimerization of 2 myosins or 2 non-myosin binding partners (). An example for the former is the potential RLC mediated ternary complex between the nonmuscle myosin-2 heavy chain and the bile salt export protein BSEP that is required for BSEP trafficking and delivery to the apical membrane.Citation178 Also, a ternary complex of yeast ELC with the myosin-2 heavy chain and another, so far unknown binding partner, is suggested.Citation179,180
Figure 4. Consequences of different binding modes of myosin light chains. (A) Schematic representation of the open and closed conformation of CaM (blue) bound to an exclusive target IQ-motif (gray). CaM-dependent dimerization of 2 domains of the Ca2+-activated K+ channel (gray), each bound to one lobe of CaM (PDB ID: 1G4Y). (B) Extended and compact light chain conformation exemplarily shown for the ELC (orange) on the IQ motifs (gray) of the yeast myosin-5 (PDB IDs: 1M45, 1M46). Potential functional consequences of the extended and compact light chain conformation are valid for ELC, RLC and CaM.
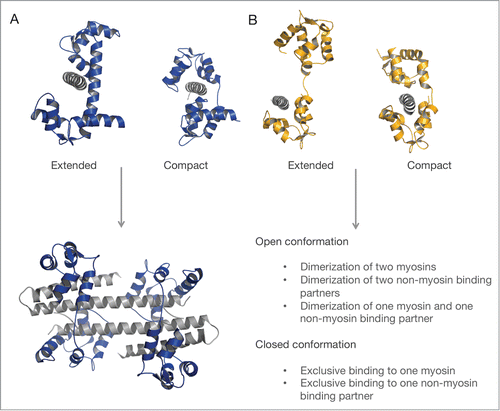
Other Myosin Light Chains and Potential Light Chains
Besides ELC, RLC and CaM, other EF-hand proteins are described as potential myosin light chains. As demonstrated by Kollmar, numerous CaM-related Ca2+-binding proteins from the centrin/caltractin, troponin C, frequinin/hippocalcin, and calcineurin families and other unconventional proteins might potentially function as myosin light chains.Citation181 Experimentally verified examples include the lower eukaryotic light chains MIMLC, MlcB, MlcC and MlcD that associate with the IQ motifs of Acanthamoeba myosin-1C and Dictyostelium myosins-1B, -1C and -1D.Citation182-184 Dictyostelium MlcD is a CaM-like protein with a molecular weight of 16.5 kDa. MlcD has 4 EF-hands, though all of them have lost high affinity Ca2+-binding sites.Citation182 In contrast, MlcB and MlcC are low molecular (8.3–8.6 kDa), single-lobe light chains.Citation185,186 MlcC EF-hands have lost the ability to bind Ca2+.Citation186 In contrast, MlcB can bind Ca2+ and undergoes a Ca2+-dependent conformational change, however its tight, submicromolar affinity for the myosin-1B IQ motif is Ca2+-independent suggesting that structural plasticity is not critical for the MlcB:IQ interaction.Citation184,185 A recent NMR-structure of the globular, 4-helix bundle MlcB indicates a significantly different IQ motif recognition mode when compared to bi-lobed light chains that involves extensive hydrophobic and electrostatic interactions.Citation185 The structure of single-lobed MlcB is most reminiscent to the closed conformation of the CaM C-lobe under Ca2+-free conditions.Citation185
Androcam is a CaM-like protein expressed in limited tissues in Drosophila which associates with the myosin-6 heavy chain in testes as determined by immunolocalization, co-immunoprecipitation, yeast-2-hybrid and peptide studies.Citation187 The myosin-6:androcam complex localizes to F-actin cones and does not co-localize with CaM by immunofluorescence microscopy, strongly suggesting that androcam is the physiological light chain for myosin-6 in fly testes. Recent studies demonstrate that androcam has an unusual structure: Its N-lobe noncanonically binds a single Ca2+ weakly through a site created from both EF-hands.Citation188 The site remains in the closed position regardless of whether Ca2+ is present. The C-lobe binds 2 Ca2+ with high affinity and may be saturated with Ca2+ over the entire physiological concentration range. Androcam binds to peptides from both, the insert-2 region and the canonical IQ motif of the fly myosin-6 heavy chain. Binding of androcam to the insert-2 peptide occurs only via the C-lobe and thus differs from the binding mode of CaM to the insert-2 region in the mammalian myosin-6 heavy chain.Citation188 Notably, peptides of the insert-2 region and the canonical IQ motif also bind Drosophila CaM, which is probably the physiological myosin-6 light chain in most other tissues.Citation188
The native light chain composition of unconventional myosin-10 is unknown, though recombinant heavy chain interacts with CaM.Citation189,190 CaM-like protein (CLP) is expressed in epithelial tissues where it associates with the 3 IQ motifs of myosin-10.Citation191 The binding of CLP to myosin-10 was first shown by CLP overlays of cell extracts and subsequent yeast-2-hybrid studies and confirmed by co-immunoprecipitation of overproduced proteins in mammalian cells and later by direct binding studies using bacterially produced IQ motif fragments. Direct binding studies with IQ1-IQ2 and IQ1-IQ2-IQ3 peptides suggest that IQ3 preferentially associates with CLP in the presence of Ca2+, though subsequent studies using the individual IQ motifs do not confirm the CLP specificity for IQ3 in the presence of Ca2+, but do confirm the strong effect of Ca2+ on the CLP:IQ interaction.Citation192
As mentioned above, apicomplexan-specific class-14 myosins interact with their light chains via strongly degenerated IQ motifs. The association between myosin heavy and light chains is critically dependent for parasite motility and invasion.Citation26 T. gondii has a repertoire of 7 putative myosin light chains (TgMLC1–7) which all contain 4 degenerated EF-hand domains.Citation38 Six out of the 7 light chains contain a long N-terminal extension when compared to CaM which in case of TgMLC3 is predicted to form a coiled-coil forming sequence.Citation38
Toxoplasma myosin-A copurifies with TgMLC1 from parasites. The heavy chain:light chain interaction is unique in that TgMLC1 anchors the myosin holoenzyme to the plasma membrane.Citation26 Similarly, the N-terminal extension of TgMLC2 is required for the localization of the myosin-D holoenzyme to the parasite pellicle and anchoring to the plasma membrane, thereby substituting for the function of the lacking tail domain to determine the localization of the myosin-D holoenzyme.Citation38 The association between TgMLC2 and the plasma membrane is predicted to be mediated by putative palmitoylation sites in the N-terminal domain of the light chain, whereas the C-terminal CaM-like domain of TgMLC2 is sufficient for the interaction with the myosin-D heavy chain. The small molecule tachypleginA and its analogs covalently bind to a cysteine residue (C58) in the N-terminus of TgMLC1. The covalent light chain modification inhibits the enzymatic activity of the holoenzyme, parasite motility and invasion and might be a first step toward the design of anti-parasitic drugs.Citation193-195
In Plasmodium species, light chain MLC1 is also called myosin-A tail-interacting protein (MTIP). MTIP localizes the heavy chain in complex with other proteins of the host cell invasion machinery to the inner membrane complex. P. knowlesi MTIP interacts with the myosin-A tail of P. yoelli in an extended conformation in which the C-lobe makes extensive hydrophobic contacts with the tail domain.Citation196 P. falciparum MTIP interacts in a compact conformation with the P. yoelli tail sequence.Citation197 The C-lobe still establishes hydrophobic interactions with the tail domain but the central helix is kinked, which allows the N-lobe to interact with the myosin-A tail helix. The compact conformation of MTIP is reminiscent to the structure of the ELC bound to scallop myosin-2.Citation69 Intriguingly, lysine K813 at the seventh position of the myosin-A IQ motif is crucially involved in the formation of the compact conformation.Citation197 This is counter to the results from studies on the P. knowlesi myosin-A and yeast myosin-5 which predict that a lysine at this position would interfere with binding of the N-lobe to the IQ motif, thereby preventing the formation of a compact conformation, as discussed above.Citation36,196 This exception to the rule indicates impressively the structural plasticity of the light chain:heavy chain interaction which – in the case of P. falciparum – is crucially involved in complex formation, as suggested by yeast-2-hybrid studies.Citation197
Plasmodium myosin-B localization and function differs substantially from myosin-A. Myosin-B does not associate with MTIP but binds the 78.7 kDa MLC-B, the largest myosin light chain identified in any species.Citation198 MLC-B harbors a CaM-like C-terminus and an extended α-helical N-terminus which is predicted to form a dimeric or trimeric coiled-coil.Citation198 Those findings support the idea that class-14 myosins compensate for the lack of a tail domain by a light chain that localizes the holoenzyme to its intracellular destination. Further, the extended N-terminus of MLC-B and its ability to form a coiled-coiled hypothetically results in a light chain mediated dimerization or oligomerization of 2 or more monomeric myosin-B heavy chains.
CaM competes with the CaM-like proteins calcium-binding protein 1 (CaBP1) and the calcium- and integrin-binding-protein-1 (CIB1) for the IQ motifs of mammalian myosin-1C.Citation199 In myosin-5A, the t-SNARE syntaxin-1A competes with CaM for the first IQ-motif at high Ca2+, an interaction required for tethering the motor complex to the plasma membrane during the early stages of exocytosis.Citation200 The effects of light chain substitutions on myosin's mechanochemical activity are not yet elucidated.
Caveats for the Use of Fluorescent Protein (FP) Tagged RLC and Phosphomimetics in Cell Biological Studies
Because of the supposed exclusive association to conventional myosins, FP fusion to the N- or C-terminus of RLC and the use of RLC phosphomimetics is a popular tool to study nonmuscle myosin-2 localization, function and regulation in live cells.Citation106,201-207 As reviewed recently, there are a few caveats associated with it.Citation208 The first major caveat to this approach – besides the widely accepted FP-driven mislocalization and aggregation of its fusion protein – are the above mentioned numerous unconventional myosin and non-myosin binding partners of ELC and RLC (). Further, FP-labeled light chains will bind to the 3 nonmuscle myosin-2 isoforms and their splice variants in virtually all mammalian cell types and tissues which excludes a strict distinction of any functional diversity that may exists among these myosins.
The second major caveat is that FP-RLC adversely affects nonmuscle myosin-2 regulation and enzymology. A recombinant nonmuscle myosin-2 with a GFP fused to the RLC N-terminus is regulated by light chain phosphorylation, however, the maximum catalytic activity of the holoenzyme under steady-state conditions is about half of the control and the in vitro motility is reduced by a smaller amount.Citation209 These effects are relatively small in terms of absolute rates, but are large compared to the effect of disease-associated point mutations in cardiac myosin-2 where the effects of potentially fatal mutations are subtle when probed enzymatically.Citation210,211 Therefore, it is possible that even the small effects seen for nonmuscle myosin-2 might be adversely affecting its performance in cells in ways that might be difficult to interpret. The in vitro phosphorylation rate of GFP-RLC bound to the nonmuscle myosin-2A heavy chain by MLCK is reduced, which suggests a lower overall phosphorylation level inside cells with unpredictable consequences for cellular homeostasis.Citation209
The third caveat is perhaps the most serious. To mimic the constitutively active, phosphorylated state of nonmuscle myosin-2, RLC T18/S19 are often replaced with aspartate/glutamate residues – despite a considerable chemical difference between the phosphorylated amino acid and the side chain of the surrogate, including size and charge at neutral pH. Strikingly, phosphomimetics do not fully phenocopy the phosphorylated state of the RLC in vivo.Citation58,59 Further, T18/S19 phosphomimetic RLCs interact differently with nonmuscle myosin-2A and -2B. Whereas phosphomimetic RLC for the mono- and di-phosphorylated RLC regulate the assembly and stability of nonmuscle myosin-2B filaments, nonmuscle myosin-2A filaments are not impacted.Citation212 In Drosophila, some phosphomimetic RLC mutants accumulate nonmuscle myosin-2 in aggregates, which reduce in size when the native RLC is present at endogenous levels.Citation59 Nonmuscle myosin-2 aggregation is also observed in hypomorphic flies that have a 90% reduced RLC level.Citation213 The cause for nonmuscle myosin-2 aggregation is unknown but it is suggested that the aggregates, that do not contain F-actin, could result from constitutive nonmuscle myosin-2 filament assembly.Citation59
In vitro, biochemical experiments show that a single replacement of either T18 or S19 with a charged amino acid does not fully activate smooth muscle myosin-2 in the absence of T18 or T19 phosphorylation. RLC T18D/E and S19D/E myosin holoenzymes have less than 10% of the actin-activated ATPase activity of wild type (WT) phosphorylated myosins and the S19E mutant does not move actin filaments in the in vitro motility assay in the absence of phosphorylation by MLCK. In contrast, the unphosphorylated S19E myosin exhibits 94% of the filament forming ability of the WT phosphorylated smooth muscle myosin-2 in the presence of ATP, indicating an uncoupling of the regulation of filament formation and enzymatic activation. Even simultaneous replacement of both, RLC S19 and T18, with charged amino acids only results in 16–30% of the ATPase activity and 45–60% of the actin sliding speed of phosphorylated WT myosin. This T18E/S19E double mutant forms filaments nearly as well as WT phosphorylated smooth muscle myosin-2.Citation96,101 Note that a recent molecular dynamics simulation of the effect of phosphomimetic substitutions of the light chain on its conformation also support the idea that single phosphomimetic amino acid replacement of T18 or S19 does not activate, whereas the double substitution might be partially activating the myosin-2 holoenzyme.Citation214
To mimic the dephosphorylated RLC state, alanine is sometimes used as a surrogate for T18/S19. Biochemical studies with smooth muscle-2 indicate that a single alanine replacement at S19 or at T18 does not drastically affect regulation or activity since MLCK will phosphorylate the remaining phosphorylatable residue. These mutant smooth muscle myosins-2 are inactive when unphosphorylated and have WT-like activities when phosphorylated.Citation96,101
There are no in vitro data to address the question of whether phosphomimetics of the S1/S2 PKC sites actually mimic the phosphorylation.
In summary, none of the biochemical studies supports the hypothesis that RLC phosphomimetics faithfully mimic the phosphorylated or dephosphorylated RLC state and could bona fide regulate the activity of the myosin-2 holoenzyme. Further, RLC mutants that simultaneously mimic the phosphorylated or dephosphorylated state on both, T18 and S19 are completely uncoupled from upstream signaling pathways whereas RLC mutants carrying solely amino acid surrogate at either T18 or S19 are still integrated. This may lead – apart from the predicted misregulation - to myosin-2 mislocalization.
Summary and Perspectives
Myosin light chains are as diverse as the associated heavy chains. They are more than passive subunits of the myosin holoenzyme and have structural and regulatory functions. Light chains are themselves regulated by cation binding or phosphorylation and allosterically communicate with the myosin heavy chain. Further, light chains can mediate the association of a myosin heavy chain with a binding partner to form a ternary complex or determine the intracellular localization of a myosin heavy chain. Mutations of some myosin light chain genes, alternate splicing events of pre-mRNA and aberrant expression levels are disease causing in humans. As a target for small molecules, myosin light chains are druggable and represent promising targets for example for the design of anti-parasitic drugs.
The historically strict categorization of ELC and RLC as exclusive light chains for conventional myosins and CaM as the light chain for unconventional myosins is obsolete. We are beginning to understand that ELC and RLC – as CaM – can exist as free light chains in the cytosol, bind with nanomolar affinity to the heavy chain of unconventional myosins and non-myosin binding partners in both, a phosphorylation dependent and independent manner. Therefore, the term “myosin holoenzyme” per se is needs to be re-defined, given the subunit heterogeneity for both, the heavy and the light chains. A challenge in the future will be to delineate the native heavy and light chain composition of conventional and unconventional myosins, identify how myosin heavy chains select their light chain in a spatiotemporal manner, and research on the physiological significance the heavy:light chain composition in health and disease.
As an editorial comment, we would like to point out that many of the publications, including ones from our own laboratory, have often used myosin heavy and light chain genes from different species when recombinantly overexpressing myosin fragments and holoenzymes in Sf9 insect cells. Likewise, the same mix-and-match strategy is frequently used in cell biological experiments where further the species of the host cell can be different. It is very tedious and sometimes impossible to determine the exact origin and isoform of the genes used for in vitro and in vivo studies. For better comparison of the results in the “post-genomic” era, it would be extremely useful for authors to provide the accession numbers for the myosin heavy and light chain genes to create their myosin holoenzyme of interest in biochemical and cell biological studies.
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
Acknowledgments
We thank Earl Homsher for critical reading of the manuscript and Anne Houdusse for valuable discussion of the definition of EF-hands. Neil Billington contributed the structural model of myosin-2 in the off-state.
Searching only the terms “myosin essential light chain” and “myosin regulatory light chain” on PubMed results in ∼1000 hits, which is a great underestimation given the light chain diversity within the myosin superfamily. Many more excellent publications deserve the inclusion in this review but could not be cited due to space limitations.
Funding
JRS is funded by the Intramural Program of the National Heart, Lung, and Blood Institute (HL001786).
References
- Sellers JR. Myosins: a diverse superfamily. Biochim Biophys Acta 2000; 1496:3-22; PMID:10722873; http://dx.doi.org/10.1016/S0167-4889(00)00005-7
- Odronitz F, Kollmar M. Drawing the tree of eukaryotic life based on the analysis of 2,269 manually annotated myosins from 328 species. Genome Biol 2007; 8:R196; PMID:17877792; http://dx.doi.org/10.1186/gb-2007-8-9-r196
- Cheney RE, Mooseker MS. Unconventional myosins. Curr Opin Cell Biol 1992; 4:27-35; PMID:1558751; http://dx.doi.org/10.1016/0955-0674(92)90055-H
- Bahler M, Rhoads A. Calmodulin signaling via the IQ motif. FEBS Lett 2002; 513:107-13; PMID:11911888; http://dx.doi.org/10.1016/S0014-5793(01)03239-2
- Vetter SW, Leclerc E. Novel aspects of calmodulin target recognition and activation. Eur J Biochem 2003; 270:404-14; PMID:12542690; http://dx.doi.org/10.1046/j.1432-1033.2003.03414.x
- Rayment I, Rypniewski WR, Schmidt-Base K, Smith R, Tomchick DR, Benning MM, Winkelmann DA, Wesenberg G, Holden HM. Three-dimensional structure of myosin subfragment-1: a molecular motor. Science 1993; 261:50-8; PMID:8316857; http://dx.doi.org/10.1126/science.8316857
- Sellers JR, Chantler PD, Szent-Gyorgyi AG. Hybrid formation between scallop myofibrils and foreign regulatory light-chains. J Mol Biol 1980; 144:223-45; PMID:6454788; http://dx.doi.org/10.1016/0022-2836(80)90088-1
- Kominz DR, Carroll WR, Smith EN, Mitchell ER. A Subunit of Myosin. Arch Biochem Biophys 1959; 79:191-9; http://dx.doi.org/10.1016/0003-9861(59)90395-9
- Hartman MA, Spudich JA. The myosin superfamily at a glance. J Cell Sci 2012; 125:1627-32; PMID:22566666; http://dx.doi.org/10.1242/jcs.094300
- Odronitz F, Kollmar M. Pfarao: a web application for protein family analysis customized for cytoskeletal and motor proteins (CyMoBase). BMC Genomics 2006; 7:300; PMID:17134497; http://dx.doi.org/10.1186/1471-2164-7-300
- Coluccio LM, Bretscher A. Mapping of the microvillar 110K-calmodulin complex (brush border myosin I). Identification of fragments containing the catalytic and F-actin-binding sites and demonstration of a calcium ion dependent conformational change. Biochemistry 1990; 29:11089-94; PMID:2271696; http://dx.doi.org/10.1021/bi00502a011
- Collins K, Sellers JR, Matsudaira P. Calmodulin dissociation regulates brush border myosin I (110-kD-calmodulin) mechanochemical activity in vitro. J Cell Biol 1990; 110:1137-47; PMID:2139032; http://dx.doi.org/10.1083/jcb.110.4.1137
- Coluccio LM, Conaty C. Myosin-I in mammalian liver. Cell Motil Cytoskeleton 1993; 24:189-99; PMID:8467525; http://dx.doi.org/10.1002/cm.970240306
- Bahler M, Kroschewski R, Stoffler HE, Behrmann T. Rat myr 4 defines a novel subclass of myosin I: identification, distribution, localization, and mapping of calmodulin-binding sites with differential calcium sensitivity. J Cell Biol 1994; 126:375-89; PMID:8034741; http://dx.doi.org/10.1083/jcb.126.2.375
- Ruppert C, Kroschewski R, Bahler M. Identification, characterization and cloning of myr 1, a mammalian myosin-I. J Cell Biol 1993; 120:1393-403; PMID:8449985; http://dx.doi.org/10.1083/jcb.120.6.1393
- Espindola FS, Suter DM, Partata LB, Cao T, Wolenski JS, Cheney RE, King SM, Mooseker MS. The light chain composition of chicken brain myosin-Va: calmodulin, myosin-II essential light chains, and 8-kDa dynein light chain/PIN. Cell Motil Cytoskeleton 2000; 47:269-81; PMID:11093248; http://dx.doi.org/10.1002/1097-0169(200012)47:4%3c269::AID-CM2%3e3.0.CO;2-G
- Wang F, Chen L, Arcucci O, Harvey EV, Bowers B, Xu Y, Hammer JA, 3rd, Sellers JR. Effect of ADP and ionic strength on the kinetic and motile properties of recombinant mouse myosin V. J Biol Chem 2000; 275:4329-35; PMID:10660602; http://dx.doi.org/10.1074/jbc.275.6.4329
- Battelle BA, Andrews AW, Calman BG, Sellers JR, Greenberg RM, Smith WC. A myosin III from Limulus eyes is a clock-regulated phosphoprotein. J Neurosci 1998; 18:4548-59; PMID:9614231
- Porter JA, Yu M, Doberstein SK, Pollard TD, Montell C. Dependence of calmodulin localization in the retina on the NINAC unconventional myosin. Science 1993; 262:1038-42; PMID:8235618; http://dx.doi.org/10.1126/science.8235618
- Porter JA, Minke B, Montell C. Calmodulin binding to Drosophila NinaC required for termination of phototransduction. Embo J 1995; 14:4450-9; PMID:7556088
- Hasson T, Mooseker MS. Porcine myosin-VI: characterization of a new mammalian unconventional myosin. J Cell Biol 1994; 127:425-40; PMID:7929586; http://dx.doi.org/10.1083/jcb.127.2.425
- Todorov PT, Hardisty RE, Brown SD. Myosin VIIA is specifically associated with calmodulin and microtubule-associated protein-2B (MAP-2B). Biochem J 2001; 354:267-74; PMID:11171103; http://dx.doi.org/10.1042/0264-6021:3540267
- Udovichenko IP, Gibbs D, Williams DS. Actin-based motor properties of native myosin VIIa. J Cell Sci 2002; 115:445-50; PMID:11839794
- Post PL, Bokoch GM, Mooseker MS. Human myosin-IXb is a mechanochemically active motor and a GAP for rho. J Cell Sci 1998; 111 (Pt 7):941-50; PMID:9490638
- Tan I, Yong J, Dong JM, Lim L, Leung T. A tripartite complex containing MRCK modulates lamellar actomyosin retrograde flow. Cell 2008; 135:123-36; PMID:18854160; http://dx.doi.org/10.1016/j.cell.2008.09.018
- Herm-Gotz A, Weiss S, Stratmann R, Fujita-Becker S, Ruff C, Meyhofer E, Soldati T, Manstein DJ, Geeves MA, Soldati D. Toxoplasma gondii myosin A and its light chain: a fast, single-headed, plus-end-directed motor. Embo J 2002; 21:2149-58; PMID:11980712; http://dx.doi.org/10.1093/emboj/21.9.2149
- Bahloul A, Chevreux G, Wells AL, Martin D, Nolt J, Yang Z, Chen LQ, Potier N, Van Dorsselaer A, Rosenfeld S, et al. The unique insert in myosin VI is a structural calcium-calmodulin binding site. Proc Natl Acad Sci U S A 2004; 101:4787-92; PMID:15037754; http://dx.doi.org/10.1073/pnas.0306892101
- Lin T, Tang N, Ostap EM. Biochemical and motile properties of Myo1b splice isoforms. J Biol Chem 2005; 280:41562-7; PMID:16254000; http://dx.doi.org/10.1074/jbc.M508653200
- De La Cruz EM, Wells AL, Sweeney HL, Ostap EM. Actin and light chain isoform dependence of myosin V kinetics. Biochemistry 2000; 39:14196-202; PMID:11087368; http://dx.doi.org/10.1021/bi001701b
- Yang Y, Baboolal TG, Siththanandan V, Chen M, Walker ML, Knight PJ, Peckham M, Sellers JR. A FERM domain autoregulates Drosophila myosin 7a activity. Proc Natl Acad Sci U S A 2009; 106:4189-94; PMID:19255446; http://dx.doi.org/10.1073/pnas.0808682106
- Sorensen AB, Sondergaard MT, Overgaard MT. Calmodulin in a heartbeat. Febs J 2013; 280:5511-32; PMID:23663249; http://dx.doi.org/10.1111/febs.12337
- Babu YS, Bugg CE, Cook WJ. Structure of calmodulin refined at 2.2 A resolution. J Mol Biol 1988; 204:191-204; PMID:3145979; http://dx.doi.org/10.1016/0022-2836(88)90608-0
- Grabarek Z. Structural basis for diversity of the EF-hand calcium-binding proteins. J Mol Biol 2006; 359:509-25; PMID:16678204; http://dx.doi.org/10.1016/j.jmb.2006.03.066
- Lu Q, Li J, Ye F, Zhang M. Structure of myosin-1c tail bound to calmodulin provides insights into calcium-mediated conformational coupling. Nat Struct Mol Biol 2015; 22:81-8; PMID:25437912; http://dx.doi.org/10.1038/nsmb.2923
- Terrak M, Wu G, Stafford WF, Lu RC, Dominguez R. Two distinct myosin light chain structures are induced by specific variations within the bound IQ motifs-functional implications. Embo J 2003; 22:362-71; PMID:12554638; http://dx.doi.org/10.1093/emboj/cdg058
- Terrak M, Rebowski G, Lu RC, Grabarek Z, Dominguez R. Structure of the light chain-binding domain of myosin V. Proc Natl Acad Sci U S A 2005; 102:12718-23; PMID:16120677; http://dx.doi.org/10.1073/pnas.0503899102
- Chen XW, Leto D, Chiang SH, Wang Q, Saltiel AR. Activation of RalA is required for insulin-stimulated Glut4 trafficking to the plasma membrane via the exocyst and the motor protein Myo1c. Dev Cell 2007; 13:391-404; PMID:17765682; http://dx.doi.org/10.1016/j.devcel.2007.07.007
- Polonais V, Javier Foth B, Chinthalapudi K, Marq JB, Manstein DJ, Soldati-Favre D, Frenal K. Unusual anchor of a motor complex (MyoD-MLC2) to the plasma membrane of Toxoplasma gondii. Traffic 2011; 12:287-300; PMID:21143563; http://dx.doi.org/10.1111/j.1600-0854.2010.01148.x
- Komiyama M, Soldati T, von Arx P, Perriard JC. The intracompartmental sorting of myosin alkali light chain isoproteins reflects the sequence of developmental expression as determined by double epitope-tagging competition. J Cell Sci 1996; 109 (Pt 8):2089-99; PMID:8856505
- Khan MM, Komiyama M. The second EF-hand is responsible for the isoform-specific sorting of myosin essential light chain. Cell Struct Funct 2001; 26:243-51; PMID:11699641; http://dx.doi.org/10.1247/csf.26.243
- Kim JH, Torgerud WS, Mosser KH, Hirai H, Watanabe S, Asakura A, Thompson LV. Myosin light chain 3f attenuates age-induced decline in contractile velocity in MHC type II single muscle fibers. Aging Cell 2012; 11:203-12; PMID:…; http://dx.doi.org/10.1111/j.1474-9726.2011.00774.x
- Petzhold D, Simsek B, Meissner R, Mahmoodzadeh S, Morano I. Distinct interactions between actin and essential myosin light chain isoforms. Biochem Biophys Res Commun 2014; 449:284-8; PMID:24857983; http://dx.doi.org/10.1016/j.bbrc.2014.05.040
- Timson DJ, Trayer HR, Trayer IP. The N-terminus of A1-type myosin essential light chains binds actin and modulates myosin motor function. Eur J Biochem 1998; 255:654-62; PMID:9738905; http://dx.doi.org/10.1046/j.1432-1327.1998.2550654.x
- Wagner PD, Slater CS, Pope B, Weeds AG. Studies on the actin activation of myosin subfragment-1 isoezymes and the role of myosin light chains. Eur J Biochem 1979; 99:385-94; PMID:159176; http://dx.doi.org/10.1111/j.1432-1033.1979.tb13267.x
- Muthu P, Wang L, Yuan CC, Kazmierczak K, Huang W, Hernandez OM, Kawai M, Irving TC, Szczesna-Cordary D. Structural and functional aspects of the myosin essential light chain in cardiac muscle contraction. Faseb J 2011; 25:4394-405; PMID:21885653; http://dx.doi.org/10.1096/fj.11-191973
- Zhu J, Sun Y, Zhao FQ, Yu J, Craig R, Hu S. Analysis of tarantula skeletal muscle protein sequences and identification of transcriptional isoforms. BMC Genomics 2009; 10:117; PMID:19298669; http://dx.doi.org/10.1186/1471-2164-10-117
- Chakravorty S, Vu H, Foelber V, Vigoreaux JO. Mutations of the Drosophila myosin regulatory light chain affect courtship song and reduce reproductive success. PLoS One 2014; 9:e90077; PMID:24587213; http://dx.doi.org/10.1371/journal.pone.0090077
- Miller MS, Farman GP, Braddock JM, Soto-Adames FN, Irving TC, Vigoreaux JO, Maughan DW. Regulatory light chain phosphorylation and N-terminal extension increase cross-bridge binding and power output in Drosophila at in vivo myofilament lattice spacing. Biophys J 2011; 100:1737-46; PMID:21463587; http://dx.doi.org/10.1016/j.bpj.2011.02.028
- Guhathakurta P, Prochniewicz E, Thomas DD. Amplitude of the actomyosin power stroke depends strongly on the isoform of the myosin essential light chain. Proc Natl Acad Sci U S A 2015; 112(15):4660-5; PMID:25825773
- Alcalai R, Seidman JG, Seidman CE. Genetic basis of hypertrophic cardiomyopathy: from bench to the clinics. J Cardiovasc Electrophysiol 2008; 19:104-10; PMID:17916152
- Hernandez OM, Jones M, Guzman G, Szczesna-Cordary D. Myosin essential light chain in health and disease. Am J Physiol Heart Circ Physiol 2007; 292:H1643-54; PMID:17142342; http://dx.doi.org/10.1152/ajpheart.00931.2006
- Greenberg MJ, Kazmierczak K, Szczesna-Cordary D, Moore JR. Cardiomyopathy-linked myosin regulatory light chain mutations disrupt myosin strain-dependent biochemistry. Proc Natl Acad Sci U S A 2010; 107:17403-8; PMID:20855589; http://dx.doi.org/10.1073/pnas.1009619107
- Moore JR, Leinwand L, Warshaw DM. Understanding cardiomyopathy phenotypes based on the functional impact of mutations in the myosin motor. Circ Res 2012; 111:375-85; PMID:22821910; http://dx.doi.org/10.1161/CIRCRESAHA.110.223842
- Poetter K, Jiang H, Hassanzadeh S, Master SR, Chang A, Dalakas MC, Rayment I, Sellers JR, Fananapazir L, Epstein ND. Mutations in either the essential or regulatory light chains of myosin are associated with a rare myopathy in human heart and skeletal muscle. Nat Genet 1996; 13:63-9; PMID:8673105; http://dx.doi.org/10.1038/ng0596-63
- Koi PT, Milhoua PM, Monrose V, Melman A, DiSanto ME. Expression of myosin isoforms in the smooth muscle of human corpus cavernosum. Int J Impot Res 2007; 19:62-8; PMID:16885993; http://dx.doi.org/10.1038/sj.ijir.3901503
- Nishigaki R, Shinohara T, Toda T, Omori A, Ichinose S, Itoh M, Shirayoshi Y, Kurimasa A, Oshimura M. An extra human chromosome 21 reduces mlc-2a expression in chimeric mice and Down syndrome. Biochem Biophys Res Commun 2002; 295:112-8; PMID:12083776; http://dx.doi.org/10.1016/S0006-291X(02)00640-X
- Tan X, Chen M. MYLK and MYL9 expression in non-small cell lung cancer identified by bioinformatics analysis of public expression data. Tumour Biol 2014; 35(12):12189-200; PMID:25179839
- Jordan P, Karess R. Myosin light chain-activating phosphorylation sites are required for oogenesis in Drosophila. J Cell Biol 1997; 139:1805-19; PMID:9412474; http://dx.doi.org/10.1083/jcb.139.7.1805
- Vasquez CG, Tworoger M, Martin AC. Dynamic myosin phosphorylation regulates contractile pulses and tissue integrity during epithelial morphogenesis. J Cell Biol 2014; 206:435-50; PMID:25092658; http://dx.doi.org/10.1083/jcb.201402004
- Uyeda TQ, Abramson PD, Spudich JA. The neck region of the myosin motor domain acts as a lever arm to generate movement. Proc Natl Acad Sci U S A 1996; 93:4459-64; PMID:8633089; http://dx.doi.org/10.1073/pnas.93.9.4459
- Warshaw DM, Guilford WH, Freyzon Y, Krementsova E, Palmiter KA, Tyska MJ, Baker JE, Trybus KM. The light chain binding domain of expressed smooth muscle heavy meromyosin acts as a mechanical lever. J Biol Chem 2000; 275:37167-72; PMID:10945998; http://dx.doi.org/10.1074/jbc.M006438200
- Sakamoto T, Yildez A, Selvin PR, Sellers JR. Step-size is determined by neck length in myosin V. Biochemistry 2005; 44:16203-10; PMID:16331980; http://dx.doi.org/10.1021/bi0512086
- Ruff C, Furch M, Brenner B, Manstein DJ, Meyhofer E. Single-molecule tracking of myosins with genetically engineered amplifier domains. Nat Struct Biol 2001; 8:226-9; PMID:11224566; http://dx.doi.org/10.1038/84962
- VanBuren P, Waller GS, Harris DE, Trybus KM, Warshaw DM, Lowey S. The essential light chain is required for full force production by skeletal muscle myosin. Proc Natl Acad Sci U S A 1994; 91:12403-7; PMID:7809049; http://dx.doi.org/10.1073/pnas.91.26.12403
- Lowey S, Waller GS, Trybus KM. Skeletal muscle myosin light chains are essential for physiological speeds of shortening. Nature 1993; 365:454-6; PMID:8413589; http://dx.doi.org/10.1038/365454a0
- Heissler SM, Manstein DJ. Functional characterization of the human myosin-7a motor domain. Cell Molecular Life Sci 2012; 69:299-311; PMID:21687988; http://dx.doi.org/10.1007/s00018-011-0749-8
- Hachikubo Y, Ito K, Schiefelbein J, Manstein DJ, Yamamoto K. Enzymatic activity and motility of recombinant Arabidopsis myosin XI, MYA1. Plant Cell Physiol 2007; 48:886-91; PMID:17504816; http://dx.doi.org/10.1093/pcp/pcm054
- Shuman H, Greenberg MJ, Zwolak A, Lin T, Sindelar CV, Dominguez R, Ostap EM. A vertebrate myosin-I structure reveals unique insights into myosin mechanochemical tuning. Proc Natl Acad Sci U S A 2014; 111:2116-21; PMID:24469830; http://dx.doi.org/10.1073/pnas.1321022111
- Houdusse A, Cohen C. Structure of the regulatory domain of scallop myosin at 2 A resolution: implications for regulation. Structure 1996; 4:21-32; PMID:8805510; http://dx.doi.org/10.1016/S0969-2126(96)00006-8
- Dominguez R, Freyzon Y, Trybus KM, Cohen C. Crystal structure of a vertebrate smooth muscle myosin motor domain and its complex with the essential light chain: visualization of the pre-power stroke state. Cell 1998; 94:559-71; PMID:9741621; http://dx.doi.org/10.1016/S0092-8674(00)81598-6
- Houdusse A, Szent-Gyorgyi AG, Cohen C. Three conformational states of scallop myosin S1. Proc Natl Acad Sci U S A 2000; 97:11238-43; PMID:11016966; http://dx.doi.org/10.1073/pnas.200376897
- Houdusse A, Cohen C. Target sequence recognition by the calmodulin superfamily: implications from light chain binding to the regulatory domain of scallop myosin. Proc Natl Acad Sci U S A 1995; 92:10644-7; PMID:7479857; http://dx.doi.org/10.1073/pnas.92.23.10644
- Brown JH, Kumar VS, O'Neall-Hennessey E, Reshetnikova L, Robinson H, Nguyen-McCarty M, Szent-Gyorgyi AG, Cohen C. Visualizing key hinges and a potential major source of compliance in the lever arm of myosin. Proc Natl Acad Sci U S A 2011; 108:114-9; PMID:21149681; http://dx.doi.org/10.1073/pnas.1016288107
- Coureux PD, Wells AL, Menetrey J, Yengo CM, Morris CA, Sweeney HL, Houdusse A. A structural state of the myosin V motor without bound nucleotide. Nature 2003; 425:419-23; PMID:14508494; http://dx.doi.org/10.1038/nature01927
- Menetrey J, Bahloul A, Wells AL, Yengo CM, Morris CA, Sweeney HL, Houdusse A. The structure of the myosin VI motor reveals the mechanism of directionality reversal. Nature 2005; 435:779-85; PMID:15944696; http://dx.doi.org/10.1038/nature03592
- Munnich S, Taft MH, Manstein DJ. Crystal structure of human myosin 1c–the motor in GLUT4 exocytosis: implications for Ca2+ regulation and 14-3-3 binding. J Mol Biol 2014; 426:2070-81; PMID:24636949; http://dx.doi.org/10.1016/j.jmb.2014.03.004
- Adamek N, Coluccio LM, Geeves MA. Calcium sensitivity of the cross-bridge cycle of Myo1c, the adaptation motor in the inner ear. Proc Natl Acad Sci U S A 2008; 105:5710-5; PMID:18391215; http://dx.doi.org/10.1073/pnas.0710520105
- Koide H, Kinoshita T, Tanaka Y, Tanaka S, Nagura N, Meyer zu Horste G, Miyagi A, Ando T. Identification of the single specific IQ motif of myosin V from which calmodulin dissociates in the presence of Ca2+. Biochemistry 2006; 45:11598-604; PMID:16981719; http://dx.doi.org/10.1021/bi0613877
- Houdusse A, Gaucher JF, Krementsova E, Mui S, Trybus KM, Cohen C. Crystal structure of apo-calmodulin bound to the first two IQ motifs of myosin V reveals essential recognition features. Proc Natl Acad Sci U S A 2006; 103:19326-31; PMID:17151196; http://dx.doi.org/10.1073/pnas.0609436103
- Sakamoto T, Webb MR, Forgacs E, White HD, Sellers JR. Direct observation of the mechanochemical coupling in myosin Va during processive movement. Nature 2008; 455:128-32; PMID:18668042; http://dx.doi.org/10.1038/nature07188
- Lu H, Krementsova EB, Trybus KM. Regulation of myosin V processivity by calcium at the single molecule level. J Biol Chem 2006; 281:31987-94; PMID:16920704; http://dx.doi.org/10.1074/jbc.M605181200
- Wagner W, Fodor E, Ginsburg A, Hammer JA 3rd. The binding of DYNLL2 to myosin Va requires alternatively spliced exon B and stabilizes a portion of the myosin's coiled-coil domain. Biochemistry 2006; 45:11564-77; PMID:16981716; http://dx.doi.org/10.1021/bi061142u
- Liu J, Taylor DW, Krementsova EB, Trybus KM, Taylor KA. Three-dimensional structure of the myosin V inhibited state by cryoelectron tomography. Nature 2006; 442:208-11; PMID:16625208
- Wells AL, Lin AW, Chen LQ, Safer D, Cain SM, Hasson T, Carragher BO, Milligan RA, Sweeney HL. Myosin VI is an actin-based motor that moves backwards. Nature 1999; 401:505-8; PMID:10519557; http://dx.doi.org/10.1038/46835
- Liu Y, Hsin J, Kim H, Selvin PR, Schulten K. Extension of a three-helix bundle domain of myosin VI and key role of calmodulins. Biophys J 2011; 100:2964-73; PMID:21689530; http://dx.doi.org/10.1016/j.bpj.2011.05.010
- Bresnick AR. Molecular mechanisms of nonmuscle myosin-II regulation. Curr Opin Cell Biol 1999; 11:26-33; PMID:10047526; http://dx.doi.org/10.1016/S0955-0674(99)80004-0
- Heissler SM, Manstein DJ. Nonmuscle myosin-2: mix and match. Cell Mol Life Sci 2013; 70:1-21; PMID:22565821; http://dx.doi.org/10.1007/s00018-012-1002-9
- Amin E, Dubey BN, Zhang SC, Gremer L, Dvorsky R, Moll JM, Taha MS, Nagel-Steger L, Piekorz RP, Somlyo AV, et al. Rho-kinase: regulation, (dys)function, and inhibition. Biol Chem 2013; 394:1399-410; PMID:23950574; http://dx.doi.org/10.1515/hsz-2013-0181
- Trybus KM. Filamentous smooth muscle myosin is regulated by phosphorylation. J Cell Biol 1989; 109:2887-94; PMID:2531749; http://dx.doi.org/10.1083/jcb.109.6.2887
- Sellers JR. Mechanism of the phosphorylation-dependent regulation of smooth muscle heavy meromyosin. J Biol Chem 1985; 260:15815-9; PMID:2933403
- Wendt T, Taylor D, Messier T, Trybus KM, Taylor KA. Visualization of head-head interactions in the inhibited state of smooth muscle myosin. J Cell Biol 1999; 147:1385-90; PMID:10613897; http://dx.doi.org/10.1083/jcb.147.7.1385
- Jung HS, Billington N, Thirumurugan K, Salzameda B, Cremo CR, Chalovich JM, Chantler PD, Knight PJ. Role of the tail in the regulated state of myosin 2. J Mol Biol 2011; 408:863-78; PMID:21419133; http://dx.doi.org/10.1016/j.jmb.2011.03.019
- Trybus KM. Assembly of cytoplasmic and smooth muscle myosins. Curr Opin Cell Biol 1991; 3:105-11; http://dx.doi.org/10.1016/0955-0674(91)90172-U
- Somlyo AV. Smooth muscle myosin filament controversy, once again? J Physiol 2015; 593:473-5; PMID:25630266; http://dx.doi.org/10.1113/jphysiol.2014.285866
- Umemoto S, Bengur AR, Sellers JR. Effect of multiple phosphorylations of smooth muscle and cytoplasmic myosins on movement in an in vitro motility assay. J Biol Chem 1989; 264:1431-6; PMID:2521481
- Ikebe M, Koretz J, Hartshorne DJ. Effects of phosphorylation of light chain residues threonine 18 and serine 19 on the properties and conformation of smooth muscle myosin. J Biol Chem 1988; 263:6432-7; PMID:2966156
- Ikebe M, Hartshorne DJ. Phosphorylation of smooth muscle myosin at two distinct sites by myosin light chain kinase. J Biol Chem 1985; 260:10027-31; PMID:3839510
- Kast D, Espinoza-Fonseca LM, Yi C, Thomas DD. Phosphorylation-induced structural changes in smooth muscle myosin regulatory light chain. Proc Natl Acad Sci U S A 2010; 107:8207-12; PMID:20404208; http://dx.doi.org/10.1073/pnas.1001941107
- Taylor KA, Feig M, Brooks CL, 3rd, Fagnant PM, Lowey S, Trybus KM. Role of the essential light chain in the activation of smooth muscle myosin by regulatory light chain phosphorylation. J Struct Biol 2014; 185:375-82; PMID:24361582; http://dx.doi.org/10.1016/j.jsb.2013.12.008
- Ikebe M, Ikebe R, Kamisoyama H, Reardon S, Schwonek JP, Sanders CR, 2nd, Matsuura M. Function of the NH2-terminal domain of the regulatory light chain on the regulation of smooth muscle myosin. J Biol Chem 1994; 269:28173-80; PMID:7961753
- Sweeney HL, Yang Z, Zhi G, Stull JT, Trybus KM. Charge replacement near the phosphorylatable serine of the myosin regulatory light chain mimics aspects of phosphorylation. Proc Natl Acad Sci U S A 1994; 91:1490-4; PMID:…; http://dx.doi.org/10.1073/pnas.91.4.1490
- Ni S, Hong F, Haldeman BD, Baker JE, Facemyer KC, Cremo CR. Modification of interface between regulatory and essential light chains hampers phosphorylation-dependent activation of smooth muscle myosin. J Biol Chem 2012; 287:22068-79; PMID:22549781; http://dx.doi.org/10.1074/jbc.M112.343491
- Ikebe M, Hartshorne DJ, Elzinga M. Phosphorylation of the 20,000-dalton light chain of smooth muscle myosin by the calcium-activated, phospholipid-dependent protein kinase. Phosphorylation sites and effects of phosphorylation. J Biol Chem 1987; 262:9569-73; PMID:3036866
- Nishikawa M, Sellers JR, Adelstein RS, Hidaka H. Protein kinase C modulates in vitro phosphorylation of the smooth muscle heavy meromyosin by myosin light chain kinase. J Biol Chem 1984; 259:8808-14; PMID:6235218
- Turbedsky K, Pollard TD, Bresnick AR. A subset of protein kinase C phosphorylation sites on the myosin II regulatory light chain inhibits phosphorylation by myosin light chain kinase. Biochemistry 1997; 36:2063-7; PMID:9047304; http://dx.doi.org/10.1021/bi9624651
- Beach JR, Licate LS, Crish JF, Egelhoff TT. Analysis of the role of Ser1/Ser2/Thr9 phosphorylation on myosin II assembly and function in live cells. BMC Cell Biol 2011; 12:52; PMID:22136066; http://dx.doi.org/10.1186/1471-2121-12-52
- Asokan SB, Johnson HE, Rahman A, King SJ, Rotty JD, Lebedeva IP, Haugh JM, Bear JE. Mesenchymal chemotaxis requires selective inactivation of myosin II at the leading edge via a noncanonical PLCgamma/PKCalpha pathway. Dev Cell 2014; 31:747-60; PMID:25482883; http://dx.doi.org/10.1016/j.devcel.2014.10.024
- Vandenboom R, Gittings W, Smith IC, Grange RW, Stull JT. Myosin phosphorylation and force potentiation in skeletal muscle: evidence from animal models. J Muscle Res Cell Motil 2013; 34:317-32; PMID:24162313; http://dx.doi.org/10.1007/s10974-013-9363-8
- Stull JT, Kamm KE, Vandenboom R. Myosin light chain kinase and the role of myosin light chain phosphorylation in skeletal muscle. Arch Biochem Biophys 2011; 510:120-8; PMID:21284933; http://dx.doi.org/10.1016/j.abb.2011.01.017
- Moore RL, Houston ME, Iwamoto GA, Stull JT. Phosphorylation of rabbit skeletal muscle myosin in situ. J Cell Physiol 1985; 125:301-5; PMID:4055914; http://dx.doi.org/10.1002/jcp.1041250219
- Persechini A, Stull JT, Cooke R. The effect of myosin phosphorylation on the contractile properties of skinned rabbit skeletal muscle fibers. J Biol Chem 1985; 260:7951-4; PMID:3839239
- Sweeney HL, Stull JT. Alteration of cross-bridge kinetics by myosin light chain phosphorylation in rabbit skeletal muscle: implications for regulation of actin-myosin interaction. Proc Natl Acad Sci U S A 1990; 87:414-8; PMID:2136951; http://dx.doi.org/10.1073/pnas.87.1.414
- Sweeney HL, Stull JT. Phosphorylation of myosin in permeabilized mammalian cardiac and skeletal muscle cells. Am J Physiol 1986; 250:C657-60; PMID:3754389
- Levine RJ, Kensler RW, Yang Z, Stull JT, Sweeney HL. Myosin light chain phosphorylation affects the structure of rabbit skeletal muscle thick filaments. Biophys J 1996; 71:898-907; PMID:8842229; http://dx.doi.org/10.1016/S0006-3495(96)79293-7
- Toepfer C, Caorsi V, Kampourakis T, Sikkel MB, West TG, Leung MC, Al-Saud SA, MacLeod KT, Lyon AR, Marston SB, et al. Myosin regulatory light chain (RLC) phosphorylation change as a modulator of cardiac muscle contraction in disease. J Biol Chem 2013; 288:13446-54; PMID:23530050; http://dx.doi.org/10.1074/jbc.M113.455444
- Jacques AM, Briceno N, Messer AE, Gallon CE, Jalilzadeh S, Garcia E, Kikonda-Kanda G, Goddard J, Harding SE, Watkins H, et al. The molecular phenotype of human cardiac myosin associated with hypertrophic obstructive cardiomyopathy. Cardiovasc Res 2008; 79:481-91; PMID:18411228; http://dx.doi.org/10.1093/cvr/cvn094
- Abraham TP, Jones M, Kazmierczak K, Liang HY, Pinheiro AC, Wagg CS, Lopaschuk GD, Szczesna-Cordary D. Diastolic dysfunction in familial hypertrophic cardiomyopathy transgenic model mice. Cardiovasc Res 2009; 82:84-92; PMID:19150977; http://dx.doi.org/10.1093/cvr/cvp016
- Huang J, Shelton JM, Richardson JA, Kamm KE, Stull JT. Myosin regulatory light chain phosphorylation attenuates cardiac hypertrophy. J Biol Chem 2008; 283:19748-56; PMID:18474588; http://dx.doi.org/10.1074/jbc.M802605200
- Bosgraaf L, van Haastert PJ. The regulation of myosin II in Dictyostelium. Eur J Cell Biol 2006; 85:969-79; PMID:16814425; http://dx.doi.org/10.1016/j.ejcb.2006.04.004
- Lord M, Pollard TD. UCS protein Rng3p activates actin filament gliding by fission yeast myosin-II. J Cell Biol 2004; 167:315-25; PMID:15504913; http://dx.doi.org/10.1083/jcb.200404045
- McCollum D, Feoktistova A, Gould KL. Phosphorylation of the myosin-II light chain does not regulate the timing of cytokinesis in fission yeast. J Biol Chem 1999; 274:17691-5; PMID:10364209; http://dx.doi.org/10.1074/jbc.274.25.17691
- Wang F, Martin BM, Sellers JR. Regulation of actomyosin interactions in Limulus muscle proteins. J Biol Chem 1993; 268:3776-80; PMID:8429052
- Sellers JR. Phosphorylation-dependent regulation of Limulus myosin. J Biol Chem 1981; 256:9274-8; PMID:6114959
- Hidalgo C, Craig R, Ikebe M, Padron R. Mechanism of phosphorylation of the regulatory light chain of myosin from tarantula striated muscle. J Muscle Res Cell Motil 2001; 22:51-9; PMID:11563549; http://dx.doi.org/10.1023/A:1010388103354
- Alamo L, Wriggers W, Pinto A, Bartoli F, Salazar L, Zhao FQ, Craig R, Padron R. Three-dimensional reconstruction of tarantula myosin filaments suggests how phosphorylation may regulate myosin activity. J Mol Biol 2008; 384:780-97; PMID:18951904; http://dx.doi.org/10.1016/j.jmb.2008.10.013
- Brito R, Alamo L, Lundberg U, Guerrero JR, Pinto A, Sulbaran G, Gawinowicz MA, Craig R, Padron R. A molecular model of phosphorylation-based activation and potentiation of tarantula muscle thick filaments. J Mol Biol 2011; 414:44-61; PMID:21959262; http://dx.doi.org/10.1016/j.jmb.2011.09.017
- Kendrick-Jones J, Lehman W, Szent-Gyorgyi AG. Regulation in molluscan muscles. J Mol Biol 1970; 54:313-26; PMID:4250215; http://dx.doi.org/10.1016/0022-2836(70)90432-8
- Kendrick-Jones J, Szentkiralyi EM, Szent-Gyorgyi AG. Regulatory light chains in myosins. J Mol Biol 1976; 104:747-75; PMID:134163; http://dx.doi.org/10.1016/0022-2836(76)90180-7
- Xie X, Harrison DH, Schlichting I, Sweet RM, Kalabokis VN, Szent-Gyorgyi AG, Cohen C. Structure of the regulatory domain of scallop myosin at 2.8 A resolution. Nature 1994; 368:306-12; PMID:8127365; http://dx.doi.org/10.1038/368306a0
- Jancso A, Szent-Gyorgyi AG. Regulation of scallop myosin by the regulatory light chain depends on a single glycine residue. Proc Natl Acad Sci U S A 1994; 91:8762-6; PMID:8090720; http://dx.doi.org/10.1073/pnas.91.19.8762
- Himmel DM, Mui S, O'Neall-Hennessey E, Szent-Gyorgyi AG, Cohen C. The on-off switch in regulated myosins: different triggers but related mechanisms. J Mol Biol 2009; 394:496-505; PMID:19769984; http://dx.doi.org/10.1016/j.jmb.2009.09.035
- Kendrick-Jones J, Tooth P, Taylor KA, Scholey JM. Regulation of myosin-filament assembly by light-chain phosphorylation. Cold Spring Harb Symp Quant Biol 1982; 46 Pt 2:929-38; PMID:6955111; http://dx.doi.org/10.1101/SQB.1982.046.01.087
- Rowe T, Kendrick-Jones J. The C-terminal helix in subdomain 4 of the regulatory light chain is essential for myosin regulation. Embo J 1993; 12:4877-84; PMID:8223496
- Goodwin EB, Leinwand LA, Szent-Gyorgyi AG. Regulation of scallop myosin by mutant regulatory light chains. J Mol Biol 1990; 216:85-93; PMID:2146399; http://dx.doi.org/10.1016/S0022-2836(05)80062-2
- Trybus KM, Chatman TA. Chimeric regulatory light chains as probes of smooth muscle myosin function. J Biol Chem 1993; 268:4412-9; PMID:8440724
- Baumann BA, Taylor DW, Huang Z, Tama F, Fagnant PM, Trybus KM, Taylor KA. Phosphorylated smooth muscle heavy meromyosin shows an open conformation linked to activation. J Mol Biol 2012; 415:274-87; PMID:22079364; http://dx.doi.org/10.1016/j.jmb.2011.10.047
- Jung HS, Burgess SA, Billington N, Colegrave M, Patel H, Chalovich JM, Chantler PD, Knight PJ. Conservation of the regulated structure of folded myosin 2 in species separated by at least 600 million years of independent evolution. Proc Natl Acad Sci U S A 2008; 105:6022-6; PMID:18413616; http://dx.doi.org/10.1073/pnas.0707846105
- Ma X, Jana SS, Conti MA, Kawamoto S, Claycomb WC, Adelstein RS. Ablation of nonmuscle myosin II-B and II-C reveals a role for nonmuscle myosin II in cardiac myocyte karyokinesis. Mol Biol Cell 2010; 21:3952-62; PMID:20861308; http://dx.doi.org/10.1091/mbc.E10-04-0293
- Park I, Han C, Jin S, Lee B, Choi H, Kwon JT, Kim D, Kim J, Lifirsu E, Park WJ, et al. Myosin regulatory light chains are required to maintain the stability of myosin II and cellular integrity. Biochem J 2011; 434:171-80; PMID:21126233; http://dx.doi.org/10.1042/BJ20101473
- Maliga Z, Junqueira M, Toyoda Y, Ettinger A, Mora-Bermudez F, Klemm RW, Vasilj A, Guhr E, Ibarlucea-Benitez I, Poser I, et al. A genomic toolkit to investigate kinesin and myosin motor function in cells. Nat Cell Biol 2013; 15:325-34; PMID:23417121; http://dx.doi.org/10.1038/ncb2689
- Sellers JR, Veigel C. Walking with myosin V. Curr Opin Cell Biol 2006; 18:68-73; PMID:16378722; http://dx.doi.org/10.1016/j.ceb.2005.12.014
- Thirumurugan K, Sakamoto T, Hammer JA, 3rd, Sellers JR, Knight PJ. The cargo-binding domain regulates structure and activity of myosin 5. Nature 2006; 442:212-5; PMID:16838021; http://dx.doi.org/10.1038/nature04865
- Sato O, Li XD, Ikebe M. Myosin Va becomes a low duty ratio motor in the inhibited form. J Biol Chem 2007; 282:13228-39; PMID:17363376; http://dx.doi.org/10.1074/jbc.M610766200
- Wang F, Thirumurugan K, Stafford WF, Hammer JA, 3rd, Knight PJ, Sellers JR. Regulated conformation of myosin V. J Biol Chem 2004; 279:2333-6; PMID:14634000; http://dx.doi.org/10.1074/jbc.C300488200
- Li XD, Mabuchi K, Ikebe R, Ikebe M. Ca2+-induced activation of ATPase activity of myosin Va is accompanied with a large conformational change. Biochem Biophys Res Commun 2004; 315:538-45; PMID:14975734; http://dx.doi.org/10.1016/j.bbrc.2004.01.084
- Krementsov DN, Krementsova EB, Trybus KM. Myosin V: regulation by calcium, calmodulin, and the tail domain. J Cell Biol 2004; 164:877-86; PMID:15007063; http://dx.doi.org/10.1083/jcb.200310065
- Nguyen H, Higuchi H. Motility of myosin V regulated by the dissociation of single calmodulin. Nat Struct Mol Biol 2005; 12:127-32; PMID:15665867; http://dx.doi.org/10.1038/nsmb894
- Wu X, Rao K, Bowers MB, Copeland NG, Jenkins NA, Hammer JA 3rd. Rab27a enables myosin Va-dependent melanosome capture by recruiting the myosin to the organelle. J Cell Sci 2001; 114:1091-100; PMID:11228153
- Wu X, Wang F, Rao K, Sellers JR, Hammer JA, 3rd. Rab27a is an essential component of melanosome receptor for myosin Va. Mol Biol Cell 2002; 13:1735-49; PMID:12006666; http://dx.doi.org/10.1091/mbc.01-12-0595
- Sckolnick M, Krementsova EB, Warshaw DM, Trybus KM. More than just a cargo adapter, melanophilin prolongs and slows processive runs of myosin Va. J Biol Chem 2013; 288:29313-22; PMID:23979131; http://dx.doi.org/10.1074/jbc.M113.476929
- Trybus KM, Krementsova E, Freyzon Y. Kinetic characterization of a monomeric unconventional myosin V construct. J Biol Chem 1999; 274:27448-56; PMID:10488077; http://dx.doi.org/10.1074/jbc.274.39.27448
- Martin SR, Bayley PM. Calmodulin bridging of IQ motifs in myosin-V. FEBS Lett 2004; 567:166-70; PMID:15178316; http://dx.doi.org/10.1016/j.febslet.2004.04.053
- Martin SR, Bayley PM. Regulatory implications of a novel mode of interaction of calmodulin with a double IQ-motif target sequence from murine dilute myosin V. Protein Sci 2002; 11:2909-23; PMID:12441389; http://dx.doi.org/10.1110/ps.0210402
- Laakso JM, Lewis JH, Shuman H, Ostap EM. Control of myosin-I force sensing by alternative splicing. Proc Natl Acad Sci U S A 2010; 107:698-702; PMID:20080738; http://dx.doi.org/10.1073/pnas.0911426107
- Perreault-Micale C, Shushan AD, Coluccio LM. Truncation of a mammalian myosin I results in loss of Ca2+-sensitive motility. J Biol Chem 2000; 275:21618-23; PMID:10777479; http://dx.doi.org/10.1074/jbc.M000363200
- Geeves MA, Perreault-Micale C, Coluccio LM. Kinetic analyses of a truncated mammalian myosin I suggest a novel isomerization event preceding nucleotide binding. J Biol Chem 2000; 275:21624-30; PMID:10781577; http://dx.doi.org/10.1074/jbc.M000342200
- Coluccio LM, Geeves MA. Transient kinetic analysis of the 130-kDa myosin I (MYR-1 gene product) from rat liver. A myosin I designed for maintenance of tension? J Biol Chem 1999; 274:21575-80; PMID:10419463; http://dx.doi.org/10.1074/jbc.274.31.21575
- Lewis JH, Greenberg MJ, Laakso JM, Shuman H, Ostap EM. Calcium regulation of myosin-I tension sensing. Biophys J 2012; 102:2799-807; PMID:22735530; http://dx.doi.org/10.1016/j.bpj.2012.05.014
- Laakso JM, Lewis JH, Shuman H, Ostap EM. Myosin I can act as a molecular force sensor. Science 2008; 321:133-6; PMID:18599791; http://dx.doi.org/10.1126/science.1159419
- Gillespie PG, Cyr JL. Calmodulin binding to recombinant myosin-1c and myosin-1c IQ peptides. BMC Biochem 2002; 3:31; PMID:12453307; http://dx.doi.org/10.1186/1471-2091-3-31
- Cyr JL, Dumont RA, Gillespie PG. Myosin-1c interacts with hair-cell receptors through its calmodulin-binding IQ domains. J Neurosci 2002; 22:2487-95; PMID:11923413
- Manceva S, Lin T, Pham H, Lewis JH, Goldman YE, Ostap EM. Calcium regulation of calmodulin binding to and dissociation from the myo1c regulatory domain. Biochemistry 2007; 46:11718-26; PMID:17910470; http://dx.doi.org/10.1021/bi700894h
- Batters C, Arthur CP, Lin A, Porter J, Geeves MA, Milligan RA, Molloy JE, Coluccio LM. Myo1c is designed for the adaptation response in the inner ear. Embo J 2004; 23:1433-40; PMID:15014434; http://dx.doi.org/10.1038/sj.emboj.7600169
- Liao W, Elfrink K, Bahler M. Head of myosin IX binds calmodulin and moves processively toward the plus-end of actin filaments. J Biol Chem 2010; 285:24933-42; PMID:20538589; http://dx.doi.org/10.1074/jbc.M110.101105
- van Bodegraven AA, Curley CR, Hunt KA, Monsuur AJ, Linskens RK, Onnie CM, Crusius JB, Annese V, Latiano A, Silverberg MS, et al. Genetic variation in myosin IXB is associated with ulcerative colitis. Gastroenterology 2006; 131:1768-74; PMID:17087940; http://dx.doi.org/10.1053/j.gastro.2006.09.011
- Bolz H, Bolz SS, Schade G, Kothe C, Mohrmann G, Hess M, Gal A. Impaired calmodulin binding of myosin-7A causes autosomal dominant hearing loss (DFNA11). Hum Mutat 2004; 24:274-5; PMID:15300860; http://dx.doi.org/10.1002/humu.9272
- Batters C, Ellrich H, Helbig C, Woodall KA, Hundschell C, Brack D, Veigel C. Calmodulin regulates dimerization, motility, and lipid binding of Leishmania myosin XXI. Proc Natl Acad Sci U S A 2014; 111:E227-36; PMID:24379364; http://dx.doi.org/10.1073/pnas.1319285110
- Bird JE, Takagi Y, Billington N, Strub MP, Sellers JR, Friedman TB. Chaperone-enhanced purification of unconventional myosin 15, a molecular motor specialized for stereocilia protein trafficking. Proc Natl Acad Sci U S A 2014; 111:12390-5; PMID:25114250; http://dx.doi.org/10.1073/pnas.1409459111
- Bookwalter CS, Kelsen A, Leung JM, Ward GE, Trybus KM. A Toxoplasma gondii Class XIV Myosin, Expressed in Sf9 Cells with a Parasite Co-chaperone, Requires Two Light Chains for Fast Motility. J Biol Chem 2014; 289(44):30832-41; PMID:25231988
- Franke JD, Boury AL, Gerald NJ, Kiehart DP. Native nonmuscle myosin II stability and light chain binding in Drosophila melanogaster. Cell Motil Cytoskeleton 2006; 63:604-22; PMID:16917818; http://dx.doi.org/10.1002/cm.20148
- Guzik-Lendrum S, Heissler SM, Billington N, Takagi Y, Yang Y, Knight PJ, Homsher E, Sellers JR. Mammalian myosin-18A, a highly divergent myosin. J Biol Chem 2013; 288:9532-48; PMID:23382379; http://dx.doi.org/10.1074/jbc.M112.441238
- Toth J, Kovacs M, Wang F, Nyitray L, Sellers JR. Myosin V from Drosophila reveals diversity of motor mechanisms within the myosin V family. J Biol Chem 2005; 280:30594-603; PMID:15980429; http://dx.doi.org/10.1074/jbc.M505209200
- Adikes RC, Unrath WC, Yengo CM, Quintero OA. Biochemical and bioinformatic analysis of the myosin-XIX motor domain. Cytoskeleton (Hoboken) 2013; 70:281-95; PMID:23568824; http://dx.doi.org/10.1002/cm.21110
- Lu Z, Ma XN, Zhang HM, Ji HH, Ding H, Zhang J, Luo D, Sun Y, Li XD. Mouse myosin-19 is a plus-end-directed, high-duty ratio molecular motor. J Biol Chem 2014; 289:18535-48; PMID:24825904; http://dx.doi.org/10.1074/jbc.M114.569087
- Kondo T, Isoda R, Uchimura T, Sugiyama M, Hamao K, Hosoya H. Diphosphorylated but not monophosphorylated myosin II regulatory light chain localizes to the midzone without its heavy chain during cytokinesis. Biochem Biophys Res Commun 2012; 417:686-91; PMID:22166199; http://dx.doi.org/10.1016/j.bbrc.2011.11.151
- Bajaj G, Zhang Y, Schimerlik MI, Hau AM, Yang J, Filtz TM, Kioussi C, Ishmael JE. N-methyl-D-aspartate receptor subunits are non-myosin targets of myosin regulatory light chain. J Biol Chem 2009; 284:1252-66; PMID:18945678; http://dx.doi.org/10.1074/jbc.M801861200
- Schumacher MA, Rivard AF, Bachinger HP, Adelman JP. Structure of the gating domain of a Ca2+-activated K+ channel complexed with Ca2+/calmodulin. Nature 2001; 410:1120-4; PMID:11323678; http://dx.doi.org/10.1038/35074145
- Chan W, Calderon G, Swift AL, Moseley J, Li S, Hosoya H, Arias IM, Ortiz DF. Myosin II regulatory light chain is required for trafficking of bile salt export protein to the apical membrane in Madin-Darby canine kidney cells. J Biol Chem 2005; 280:23741-7; PMID:15826951; http://dx.doi.org/10.1074/jbc.M502767200
- Boyne JR, Yosuf HM, Bieganowski P, Brenner C, Price C. Yeast myosin light chain, Mlc1p, interacts with both IQGAP and class II myosin to effect cytokinesis. J Cell Sci 2000; 113 Pt 24:4533-43; PMID:11082046
- Shannon KB, Li R. A myosin light chain mediates the localization of the budding yeast IQGAP-like protein during contractile ring formation. Curr Biol 2000; 10:727-30; PMID:10873803; http://dx.doi.org/10.1016/S0960-9822(00)00539-X
- Kollmar M. Thirteen is enough: the myosins of Dictyostelium discoideum and their light chains. BMC Genomics 2006; 7:183; PMID:16857047; http://dx.doi.org/10.1186/1471-2164-7-183
- De La Roche MA, Lee SF, Cote GP. The Dictyostelium class I myosin, MyoD, contains a novel light chain that lacks high-affinity calcium-binding sites. Biochem J 2003; 374:697-705; PMID:12826013; http://dx.doi.org/10.1042/BJ20030656
- Wang ZY, Sakai J, Matsudaira PT, Baines IC, Sellers JR, Hammer JA, 3rd, Korn ED. The amino acid sequence of the light chain of Acanthamoeba myosin IC. J Muscle Res Cell Motil 1997; 18:395-8; PMID:9172081; http://dx.doi.org/10.1023/A:1018686428955
- Crawley SW, de la Roche MA, Lee SF, Li Z, Chitayat S, Smith SP, Cote GP. Identification and characterization of an 8-kDa light chain associated with Dictyostelium discoideum MyoB, a class I myosin. J Biol Chem 2006; 281:6307-15; PMID:16415352; http://dx.doi.org/10.1074/jbc.M508670200
- Liburd J, Chitayat S, Crawley SW, Munro K, Miller E, Denis CM, Spencer HL, Cote GP, Smith SP. Structure of the small Dictyostelium discoideum myosin light chain MlcB provides insights into MyoB IQ motif recognition. J Biol Chem 2014; 289:17030-42; PMID:24790102; http://dx.doi.org/10.1074/jbc.M113.536532
- Crawley SW, Liburd J, Shaw K, Jung Y, Smith SP, Cote GP. Identification of calmodulin and MlcC as light chains for Dictyostelium myosin-I isozymes. Biochemistry 2011; 50:6579-88; PMID:21671662; http://dx.doi.org/10.1021/bi2007178
- Frank DJ, Martin SR, Gruender BN, Lee YS, Simonette RA, Bayley PM, Miller KG, Beckingham KM. Androcam is a tissue-specific light chain for myosin VI in the Drosophila testis. J Biol Chem 2006; 281:24728-36; PMID:16790438; http://dx.doi.org/10.1074/jbc.M602094200
- Joshi MK, Moran S, Beckingham KM, MacKenzie KR. Structure of androcam supports specialized interactions with myosin VI. Proc Natl Acad Sci U S A 2012; 109:13290-5; PMID:22851764; http://dx.doi.org/10.1073/pnas.1209730109
- Homma K, Saito J, Ikebe R, Ikebe M. Motor function and regulation of myosin X. J Biol Chem 2001; 276:34348-54; PMID:11457842; http://dx.doi.org/10.1074/jbc.M104785200
- Kovacs M, Wang F, Sellers JR. Mechanism of action of myosin X, a membrane-associated molecular motor. J Biol Chem 2005; 280:15071-83; PMID:15705568; http://dx.doi.org/10.1074/jbc.M500616200
- Rogers MS, Strehler EE. The tumor-sensitive calmodulin-like protein is a specific light chain of human unconventional myosin X. J Biol Chem 2001; 276:12182-9; PMID:11278607; http://dx.doi.org/10.1074/jbc.M010056200
- Caride AJ, Bennett RD, Strehler EE. Kinetic analysis reveals differences in the binding mechanism of calmodulin and calmodulin-like protein to the IQ motifs of myosin-10. Biochemistry 2010; 49:8105-16; PMID:20731332; http://dx.doi.org/10.1021/bi100644q
- Leung JM, Tran F, Pathak RB, Poupart S, Heaslip AT, Ballif BA, Westwood NJ, Ward GE. Identification of T. gondii myosin light chain-1 as a direct target of TachypleginA-2, a small-molecule inhibitor of parasite motility and invasion. PLoS One 2014; 9:e98056; PMID:24892871; http://dx.doi.org/10.1371/journal.pone.0098056
- Carey KL, Westwood NJ, Mitchison TJ, Ward GE. A small-molecule approach to studying invasive mechanisms of Toxoplasma gondii. Proc Natl Acad Sci U S A 2004; 101:7433-8; PMID:15123807; http://dx.doi.org/10.1073/pnas.0307769101
- Heaslip AT, Leung JM, Carey KL, Catti F, Warshaw DM, Westwood NJ, Ballif BA, Ward GE. A small-molecule inhibitor of T. gondii motility induces the posttranslational modification of myosin light chain-1 and inhibits myosin motor activity. PLoS Pathog 2010; 6:e1000720; PMID:20084115; http://dx.doi.org/10.1371/journal.ppat.1000720
- Bosch J, Turley S, Daly TM, Bogh SM, Villasmil ML, Roach C, Zhou N, Morrisey JM, Vaidya AB, Bergman LW, et al. Structure of the MTIP-MyoA complex, a key component of the malaria parasite invasion motor. Proc Natl Acad Sci U S A 2006; 103:4852-7; PMID:16547135; http://dx.doi.org/10.1073/pnas.0510907103
- Bosch J, Turley S, Roach CM, Daly TM, Bergman LW, Hol WG. The closed MTIP-myosin A-tail complex from the malaria parasite invasion machinery. J Mol Biol 2007; 372:77-88; PMID:17628590; http://dx.doi.org/10.1016/j.jmb.2007.06.016
- Yusuf NA, Green JL, Wall RJ, Knuepfer E, Moon RW, Schulte-Huxel C, Stanway RR, Martin SR, Howell SA, Douse CH, et al. The Plasmodium Class XIV Myosin, MyoB has a Distinct Subcellular Location in Invasive and Motile Stages of the Malaria Parasite, and an Unusual Light Chain. J Biol Chem 2015; 290(19):12147-64; PMID:25802338
- Tang N, Lin T, Yang J, Foskett JK, Ostap EM. CIB1 and CaBP1 bind to the myo1c regulatory domain. J Muscle Res Cell Motil 2007; 28:285-91; PMID:17994197; http://dx.doi.org/10.1007/s10974-007-9124-7
- Watanabe M, Nomura K, Ohyama A, Ishikawa R, Komiya Y, Hosaka K, Yamauchi E, Taniguchi H, Sasakawa N, Kumakura K, et al. Myosin-Va regulates exocytosis through the submicromolar Ca2+-dependent binding of syntaxin-1A. Mol Biol Cell 2005; 16:4519-30; PMID:16030255; http://dx.doi.org/10.1091/mbc.E05-03-0252
- Castoreno AB, Smurnyy Y, Torres AD, Vokes MS, Jones TR, Carpenter AE, Eggert US. Small molecules discovered in a pathway screen target the Rho pathway in cytokinesis. Nat Chem Biol 2010; 6:457-63; PMID:20436488; http://dx.doi.org/10.1038/nchembio.363
- Mizutani T, Kawabata K, Koyama Y, Takahashi M, Haga H. Regulation of cellular contractile force in response to mechanical stretch by diphosphorylation of myosin regulatory light chain via RhoA signaling cascade. Cell Motil Cytoskeleton 2009; 66:389-97; PMID:19444895; http://dx.doi.org/10.1002/cm.20378
- Miyauchi K, Yamamoto Y, Kosaka T, Hosoya H. Myosin II activity is not essential for recruitment of myosin II to the furrow in dividing HeLa cells. Biochem Biophys Res Commun 2006; 350:543-8; PMID:17022938; http://dx.doi.org/10.1016/j.bbrc.2006.09.065
- Neco P, Fernandez-Peruchena C, Navas S, Gutierrez LM, de Toledo GA, Ales E. Myosin II contributes to fusion pore expansion during exocytosis. J Biol Chem 2008; 283:10949-57; PMID:18283106; http://dx.doi.org/10.1074/jbc.M709058200
- Burghardt TP, Josephson MP, Ajtai K. Single myosin cross-bridge orientation in cardiac papillary muscle detects lever-arm shear strain in transduction. Biochemistry 2011; 50:7809-21; PMID:21819137; http://dx.doi.org/10.1021/bi2008992
- Chen Z, Huang W, Dahme T, Rottbauer W, Ackerman MJ, Xu X. Depletion of zebrafish essential and regulatory myosin light chains reduces cardiac function through distinct mechanisms. Cardiovasc Res 2008; 79:97-108; PMID:18343897; http://dx.doi.org/10.1093/cvr/cvn073
- Komatsu S, Yano T, Shibata M, Tuft RA, Ikebe M. Effects of the regulatory light chain phosphorylation of myosin II on mitosis and cytokinesis of mammalian cells. J Biol Chem 2000; 275:34512-20; PMID:10944522; http://dx.doi.org/10.1074/jbc.M003019200
- Heissler SM, Sellers JR. Four things to know about myosin light chains as reporters for nonmuscle myosin-2 dynamics in live cells. Cytoskeleton (Hoboken) 2015
- Kengyel A, Wolf WA, Chisholm RL, Sellers JR. Nonmuscle myosin IIA with a GFP fused to the N-terminus of the regulatory light chain is regulated normally. J Muscle Res Cell Motil 2010; 31:163-70; PMID:20711642; http://dx.doi.org/10.1007/s10974-010-9220-y
- Sommese RF, Sung J, Nag S, Sutton S, Deacon JC, Choe E, Leinwand LA, Ruppel K, Spudich JA. Molecular consequences of the R453C hypertrophic cardiomyopathy mutation on human beta-cardiac myosin motor function. Proc Natl Acad Sci U S A 2013; 110:12607-12; PMID:23798412; http://dx.doi.org/10.1073/pnas.1309493110
- Bloemink M, Deacon J, Langer S, Vera C, Combs A, Leinwand L, Geeves MA. The hypertrophic cardiomyopathy myosin mutation R453C alters ATP binding and hydrolysis of human cardiac beta-myosin. J Biol Chem 2014; 289:5158-67; PMID:24344137; http://dx.doi.org/10.1074/jbc.M113.511204
- Vicente-Manzanares M, Horwitz AR. Myosin light chain mono- and di-phosphorylation differentially regulate adhesion and polarity in migrating cells. Biochem Biophys Res Commun 2010; 402:537-42; PMID:20971064; http://dx.doi.org/10.1016/j.bbrc.2010.10.071
- Royou A, Field C, Sisson JC, Sullivan W, Karess R. Reassessing the role and dynamics of nonmuscle myosin II during furrow formation in early Drosophila embryos. Mol Biol Cell 2004; 15:838-50; PMID:14657248; http://dx.doi.org/10.1091/mbc.E03-06-0440
- Espinoza-Fonseca LM, Colson BA, Thomas DD. Effects of pseudophosphorylation mutants on the structural dynamics of smooth muscle myosin regulatory light chain. Mol Biosyst 2014; 10:2693-8; PMID:25091814; http://dx.doi.org/10.1039/C4MB00364K
- El Mezgueldi M, Tang N, Rosenfeld SS, Ostap EM. The kinetic mechanism of Myo1e (human myosin-IC). J Biol Chem 2002; 277:21514-21; PMID:11940582; http://dx.doi.org/10.1074/jbc.M200713200
- Yang Y, Kovacs M, Sakamoto T, Zhang F, Kiehart DP, Sellers JR. Dimerized Drosophila myosin VIIa: a processive motor. Proc Natl Acad Sci U S A 2006; 103:5746-51; PMID:16585515; http://dx.doi.org/10.1073/pnas.0509935103
- Luo J, Vallen EA, Dravis C, Tcheperegine SE, Drees B, Bi E. Identification and functional analysis of the essential and regulatory light chains of the only type II myosin Myo1p in Saccharomyces cerevisiae. J Cell Biol 2004; 165:843-55; PMID:15210731; http://dx.doi.org/10.1083/jcb.200401040
- Ledee DR, Tripathi BK, Zelenka PS. The CDK5 activator, p39, binds specifically to myosin essential light chain. Biochem Biophys Res Commun 2007; 354:1034-9; PMID:17276406; http://dx.doi.org/10.1016/j.bbrc.2007.01.112
- Desautels M, Den Haese JP, Slupsky CM, McIntosh LP, Hemmingsen SM. Cdc4p, a contractile ring protein essential for cytokinesis in Schizosaccharomyces pombe, interacts with a phosphatidylinositol 4-kinase. J Biol Chem 2001; 276:5932-42; PMID:11087749; http://dx.doi.org/10.1074/jbc.M008715200
- Valouev IA, Urakov VN, Kochneva-Pervukhova NV, Smirnov VN, Ter-Avanesyan MD. Translation termination factors function outside of translation: yeast eRF1 interacts with myosin light chain, Mlc1p, to effect cytokinesis. Mol Microbiol 2004; 53:687-96; PMID:15228544; http://dx.doi.org/10.1111/j.1365-2958.2004.04157.x
- Nagano K, Bornhauser BC, Warnasuriya G, Entwistle A, Cramer R, Lindholm D, Naaby-Hansen S. PDGF regulates the actin cytoskeleton through hnRNP-K-mediated activation of the ubiquitin E3-ligase MIR. Embo J 2006; 25:1871-82; PMID:16619033; http://dx.doi.org/10.1038/sj.emboj.7601059
