ABSTRACT
Coronavirus disease 2019 (COVID-19) pandemic continues to constitute a public health emergency of international concern. Multiple vaccine candidates for COVID-19, which is caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), have entered clinical trials. However, some evidence suggests that patients who have recovered from COVID-19 can be reinfected. For example, in China, two discharged COVID-19 patients who had recovered and fulfilled the discharge criteria for COVID-19 were retested positive to a reverse transcription polymerase chain reaction (RT-PCR) assay for the virus. This finding is critical and could hamper COVID-19 vaccine development. This review offers literature-based evidence of reinfection with SARS-CoV-2, provides explanation for the possibility of SARS-CoV-2 reinfection both from the agent and host points of view, and discusses its implication for COVID-19 vaccine development.
Introduction
A novel coronavirus disease (COVID-19), caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), emerged in central China in December 2019.Citation1,Citation2 Since then, the virus has spread worldwide, and more than 30 million cases and more than one million deaths have been reported, based on the COVID-19 Dashboard database.Citation3 SARS-CoV-2, an enveloped and positive-sense single-stranded RNA virus, is a member of the genus Betacoronavirus, together with two highly pathogenic human viruses – severe acute respiratory syndrome coronavirus (SARS-CoV) and Middle East respiratory syndrome coronavirus (MERS-CoV).Citation2 Different studies have suggested that batsCitation4,Citation5 and pangolinCitation6,Citation7 might be the original hosts of SARS-CoV-2. Because of the genomic similarity between SARS-CoV-2 and SARS-CoV,Citation4,Citation5 these viruses are also suggested to have similar immunopathology.Citation8
With no current specific treatments for COVID-19,Citation9–11 a vaccine is expected to provide protection against SARS-CoV-2 infection. In an animal model study in which rhesus macaques after recovering from SARS-CoV-2 infection were re-exposed to SARS-CoV-2, viral replication was not detected in anal and nasopharyngeal swabs, suggesting the protective effect of the primary infection.Citation12 This promising result is important for vaccine development, as it suggests that vaccination could be an effective protective measure against SARS-CoV-2 infection. Currently, multiple vaccine candidates have entered clinical trials; more than 100 are in the vaccine development pipeline,Citation13,Citation14 and their consumer acceptanceCitation15 as well as the willingness to purchaseCitation16 the vaccine candidates have been assessed. However, recent studies have reported that some COVID-19 patients that have recovered and fulfilled the discharge criteria for COVID-19 continued to show a positive result to reverse transcription polymerase chain reaction (RT-PCR) test for the virus.Citation17–21 Since these findings are critical to the design of a vaccine against the virus, scientists are still debating the authenticity of these results to determine whether reinfection is possible. In SARS-CoV and MERS-CoV infections, conflicting findings have been reported,Citation22–29 and hence, no licensed vaccine is currently available against the viruses.Citation30,Citation31 In this review, we systematically review the evidence of repositive RT-PCR test results for those who have been declared free of COVID-19, explain the possibility of COVID-19 reinfection from the virus and host points of view, and discuss the implication of this re-infection possibility for the development of COVID-19 vaccine.
SARS-CoV-2 reinfection: the evidence from animal models and patients
MERS-CoV has been reported to reinfect camels, and neutralizing antibodies (nAbs), while not providing full immunity, could still reduce the viral load.Citation25,Citation26 In humans, MERS-CoV infection induced immunoglobulin G (IgG) production and sustained antibody levels against spike protein to clear the viral load.Citation22,Citation23 For SARS-CoV infection, studies on mice, Syrian hamsters, as well as rhesus and cynomolgus monkeys reported that SARS-CoV reexposure conferred resistance and did not enhance the severity of the disease.Citation27–29 In SARS patients, IgG antibodies produced after SARS-CoV infection could neutralize the virus and prevent reinfection by the same virus for up to 2 years.Citation24 In a small scale animal model study on rhesus macaques, animals that had previously tested positive for SARS-CoV-2 and then had a negative RT-PCR test result after treatment, showed no viral replication by RT-PCR assay of anal and nasopharyngeal swabs after SARS-CoV-2 reexposure.Citation12 Besides, none of the COVID-19 symptoms was observed, which is why scientists believe that people who recover from SARS-CoV-2 infection will produce antibodies and be immune to reinfection.
Currently, the detection of SARS-CoV-2 from patients’ samples relies on the use of a molecular-based diagnostic approach, such as RT-PCR.Citation32 With a high level of precision and definitive speed to produce reliable results, this method is recognized as the current gold standard for SARS-CoV-2 detection.Citation33 However, despite its decisive role in the detection of viral genomes, the RT-PCR method has some limitations that may lead to misdiagnosis in the state of infection.Citation33 Therefore, other clinical characteristics need to be evaluated and chest computed tomography scan needs to be performed before patients are discharged from hospitals.Citation20,Citation34–36 According to China CDC,Citation20 a COVID-19 patient should meet the following discharge criteria before discharge from hospital: (1) afebrile state for more than three consecutive days; (2) improved respiratory symptoms (no cough and expectoration, normal ranges of interleukin-6 (IL-6) and C-reactive protein (CRP) as well as oxygenation index ≥ 350); (3) improved chest radiography; and (4) negative RT-PCR results for two consecutive tests with sampling interval of at least 24 h. Similar criteria are also issued by the US CDC,Citation37 which include negative real-time RT-PCR results of nasopharyngeal and throat swabs for at least two consecutive tests with sampling interval of at least 24 h, afebrile state, and improvement in signs and symptoms of COVID-19.
The incidence of reinfection in COVID-19 patients who have recovered and had a negative RT-PCR test has been brought to public attention. In China, two discharged COVID-19 patients (39-year-old woman and 50-year-old man) were retested positive to the RT-PCR assay for the virus.Citation19 Both patients had been treated with lopinavir-ritonavir, provided with supportive care, and were discharged after meeting the discharge criteria set by CDC China. Some medical workers also showed positive RT-PCR test approximately 5–13 days after being discharged from hospital in the Hubei province of China.Citation21 Another report from China showed that 14.5% (n = 172) patients tested positive by RT-PCR after having been discharged from hospital (the median age was 28 years; patients included children below 12 years (n = 6)).Citation20 Korean CDC reported repositive RT-PCR test results for recovered COVID-19 patients in Sejong city (25.9%, n = 27), Daegu city (27.2%, n = 195), and Gyeongbuk province (48.9%, n = 47).Citation18 Approximately 59.6% of the re-positive cases had no symptoms, while the rest presented symptoms such as sore throat and cough.Citation18 A list of studies that have reported reinfection with COVID-19 is presented in . In short, there is clear evidence that reinfection with SARS-CoV-2 is possible in humans, which should be considered in the development of an effective vaccine.
Table 1. List of reinfection or reactivation cases of COVID-19 reported worldwide as of July 25, 2020
While the concern of SARS-CoV-2 reinfection cases have been raised, it is important to note that misdiagnosis for re-positivity may occur due to laboratory errors. Many steps during SARS-CoV-2 detection by RT-qPCR are prone to human and/or technical faults. Samples mishandling, problems in sensitivity and/or specificity of reagents and/or techniques used to detect the viral RNA, or even record keeping are examples of error sources that may cause atypical pattern of laboratory results, as suggested.Citation38 Indeed, the newly developed COVID-19 reagents or even primer sets currently used in the RT-qPCR detection of SARS-CoV-2 RNA are factors that cannot be ruled out from the error equation.
SARS-CoV-2 reinfection: evidence in favor of and against
As an obligate intracellular parasite, a virus heavily relies on robust activity of host organelles, particularly ribosomes, to propagate and produce new virions that are ready to infect other healthy cells.Citation39 To facilitate efficient transmission and its continuous existence, a virus requires sequential interaction between the infected host and other potential healthy host(s). Unfortunately, human cells are among the victims of pathogenic organisms with insidious viral life cycle, which include SARS-CoV-2.Citation40,Citation41 To survive infection, humans are equipped with two arms of immune responses, innate and adaptive immunity, which provide protection from invasion by pathogenic microbes.Citation42 Several studies have been performed worldwide to reveal the molecular nature and characteristics of SARS-CoV-2 and the clinical manifestations of COVID-19.Citation32 However, despite massive global cooperation, information about the host immune response characteristics against SARS-CoV-2 remains limited.
Fortunately, accumulative experience from other pathogenic human coronavirus infections, which include SARS-CoV and MERS-CoV as well as from other closely related animal coronaviruses, has provided valuable insights into the viral structure, replication, potential ways of transmission, organ targets, clinical features, and possible pathogenic mechanisms of SARS-CoV-2.Citation32,Citation43,Citation44 This information is important in fostering our efforts to mitigate COVID-19 transmission, accelerate disease management, and promote empirical yet rational pharmacological as well as nonpharmacological interventions.
Cellular and humoral antiviral immune responses, particularly adaptive immunity, are important players in the continuous host protection against cytopathic viruses.Citation45,Citation46 The presence of memory T cells and B cells with the ability to produce antibodies immediately upon reintroduction of a pathogen provides steadfast response and a high level of protection to the host,Citation39,Citation45 which in turn leads to continuous protective immunity. However, in the event of reinfection , this particular immunological concept is profoundly challenged.
Recent reports showed that B cells play an important role in the clearance of SARS-CoV-2, mostly through the production of nAbs.Citation47,Citation48 However, the duration of host protection by these nAbs remains unknown. With the detection of SARS-CoV-2 in discharged patients, which is currently considered reinfection,Citation20,Citation32,Citation49–52 many have questioned whether prolonged immune protection against SARS-CoV-2 truly prevails. In this section, the results and evidence from different fields are discussed and possible explanations on the caveats of host defenses that may lead to relapse and/or reinfection are provided. The possible explanations or arguments are provided based on two factors: the host and the agent.
Host point of view
Accumulated evidence has demonstrated that COVID-19 patients with moderate to severe disease level suffer from different types of symptoms, including decreased number of lymphocytes known as lymphopenia.Citation51,Citation53,Citation54 Lymphocytes, which comprise T and B cells, are the sole players of human adaptive immunity,Citation45,Citation46 and therefore, any deleterious effect on the quality and quantity of these cells may cause serious consequences on the integrity of host anti-SARS-CoV-2 immune responses.
A recent report suggested that lymphopenia occurs because of the extensive killing of lymphocytes in COVID-19 patients, and this event is apparently linearly correlated with IL-6 level and Fas-FasL interactions.Citation55 Lymphopenia may occur as a collateral damage because of increasing pro-inflammatory cytokine levels in COVID-19 patientsCitation41,Citation56,Citation57 or because of direct consequence of SARS-CoV-2 infection on lymphocytes.Citation58 Both these proposed mechanisms in cell death-mediated lymphocyte reduction in COVID-19 patients are illustrated in . However, whether SARS-CoV-2 virions can infect lymphocytes and directly promote the destruction of infected lymphocytes remains an interesting aspect of investigation.
Figure 1. Potential causes of lymphopenia in COVID-19 patients. (A) Lymphopenia is possible to occur as a result of T cell depletion due to either SARS-CoV-2-mediated or IL-6-mediated pyroptotic cell death. (B) Alternatively, lymphocyte count in circulation may be reduced due to massive infiltration of lymphocytes into infected tissues (e.g. alveolus) (created with BioRender)
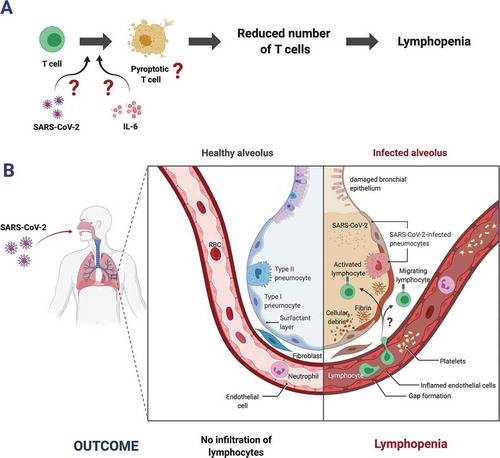
Lymphopenia in COVID-19 patients correlates with disease severity,Citation51,Citation56 probably owing to the important role of lymphocytes in providing adaptive protection against viral infections such as SARS-CoV-2.Citation8 In general, patients with lymphopenia most likely have low levels of B and/or T memory cells.Citation59 Indeed, extensive infiltration of lymphocytes into infected tissues or organs, including the lungs, as suggested by recent reports,Citation60,Citation61 may result in low lymphocyte numbers . Nevertheless, owing to the roles of memory lymphocytes in providing prolonged antiviral protection,Citation45 the decreased quantity (as well as quality) of these cells will dampen the host immune responses against reintroduction of previously encountered pathogenic microbes. In the context of SARS-CoV-2 infection, lymphopenia may lead to suboptimal production of anti-SARS-CoV-2 nAbs and/or reduced activities of CD4 helper T cells as well as CD8 cytotoxic T cells.Citation41 Taking that into account, it is possible that recovered COVID-19 patients with lymphopenia history may have increased vulnerability to SARS-CoV-2 reinfection.
As mentioned before, the occurrence of lymphopenia is associated with increased levels of proinflammatory cytokines such as IL-6.Citation55 Normally, proinflammatory cytokines are expressed as an initial response to the presence of foreign materials, including viral particles, and the expression decreases gradually during resolution of the inflammation.Citation46 These foreign materials can be characterized as pathogen-associated molecular patterns (PAMPs) and damage-associated molecular patters (DAMPs).Citation62 Both PAMPs and DAMPs are detected by pattern recognition receptors (PRRs) in a manner dependent on their unique structuresCitation63 leading to the activation of a signaling cascade via NF-kB and/or IRF pathways to produce antiviral molecules/effectors and/or pro-inflammatory cytokines.Citation62,Citation64 Elevated levels of proinflammatory cytokines have been suggested as one of the drivers of lymphocytes killing in COVID-19 patients.Citation40,Citation41,Citation65
As the expression of proinflammatory cytokines is one of the most tightly regulated processes in the human body,Citation42,Citation46 it is important to investigate the reasons for the breach of such tightly regulated processes upon SARS-CoV-2 infection. Nonetheless, based on the well-known tripartite correlation between increased level of proinflammatory cytokines, cell death, and tissue injury,Citation46 investigational studies to answer these questions will probably uncover the fundamental aspects on why reinfections occur and patients that are likely to experience reinfection, if reinfection is possible in the first place.
Another striking report regarding the characteristics of COVID-19 patients and lymphopenia is the presence of PD1 and TIM3, which are markers of T cell exhaustion.Citation56 In general, T cell exhaustion is characterized by inadequate effector function, persistent expression of inhibitory markers such as PD1 and TIM3, and a distinctive transcriptional profile compared with that of normal effector and/or memory T cells.Citation66 It has been suggested that these exhaustion characteristics cause insufficient T cell-mediated control of chronic infection,Citation66,Citation67 which may provoke relapse and/or recurrent infection.
In general, host immune responses are rapidly activated in the event of viral infection,Citation45 and the induction of humoral and cellular immune responses has been documented during human coronavirus 229E,Citation68 SARS-CoV,Citation69,Citation70 and MERS-CoV infections.Citation71,Citation72 This adaptive immunity might persist in the recovered patients for up to several years,Citation73 thus implying that reinfection is plausible after the levels of memory cells and antibodies of adaptive immunity are diminished. However, in the context of COVID-19, this alternative scenario might not be the case as reinfections were reported soon after the patients were discharged from hospitals .Citation20,Citation32,Citation49–52
Thus, while all of our speculations remain to be demonstrated experimentally, reintroduction of SARS-CoV-2 into convalescent individuals with lymphopenia history may not only cause reinfection but also provide clues about the unexplored potential of convalescent plasma therapy in the management of COVID-19.
Figure 2. Possible cause(s) of re-positivity of COVID-19 patients. Status of SARS-CoV-2 negative patients are decided based on at least two consecutive negative results on SARS-CoV-2 presence in patients’ samples using reverse transcriptase-quantitative polymerase chain reaction (RT-qPCR). Plausible assumptions of patients who turned positive RT-qPCR after discharged from hospital are based on the status of patients’ adaptive immunity (exhaustive memory cells and/or depleted memory cells) and the origin of SARS-CoV-2 present in the patients’ samples (either already present in a low copy number or obtained by reexposure). The scarce evidence is unbale to conclude either this as reinfection or relapse cases (created with BioRender)
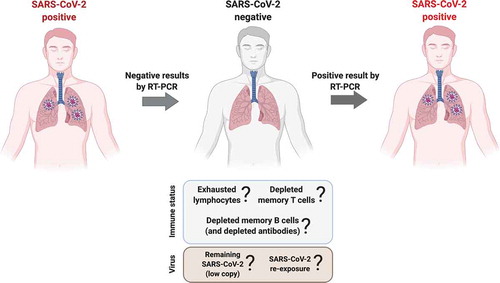
Virus point of view
In response to the COVID-19 pandemic, rapid and reliable methods for the detection of SARS-CoV-2 are urgently needed. Currently, the RT-PCR technique can be adopted to vigorously assess samples within several hours. The fundamental principle of this method is the detection of SARS-CoV-2 genetic material in a manner dependent on the action of reverse transcriptase and optimal amplification by DNA polymerase.Citation74 However, false negative results can occur owing to a really low level of starting genetic materials present in the examined samples , leading to misinterpretation of supposedly positive results as negative for SARS-CoV-2.
To this date, most of reinfection cases were reported from China despite the fact that COVID-19 pandemic has been occurred for several months in more than 200 countries. Many factors, including laboratory errors as described above, may play a role in the detection of false-positive reinfection cases. In addition to that, variation of swab sites for SARS-CoV-2 sample detection, oral, anal, sputum, saliva, and nasopharyngeal swabs, may yield different results, as suggested in recent publications.Citation75,Citation76 In respect to the immunogenicity data, there is a possibility that patients may yield different results; tested negative from samples collected in one site but potentially positive in samples taken from other swab sites. With this in mind, it might be important to compare laboratory data using the same swab sites and diagnostic parameters before jumping to conclusion.
SARS-CoV-2, with ssRNA as its genome, uses RNA-dependent RNA polymerase (RdRp) in its replication, and owing to its error-prone tendency, RdRp may produce progenies with slightly mutated genes.Citation77 A recent report has suggested the presence of mutations in SARS-CoV-2 genome, particularly in ORF1ab, ORF8, and N genes.Citation78 The RT-PCR method used for the detection of SARS-CoV-2 heavily relies on the amplification of particular sequences in SARS-CoV-2 genes; hence, mutation at primer and/or probe sites may cause false-negative detection, and SARS-CoV-2-infected persons may show negative results. However, with several uncertainties in the characteristics of the SARS-CoV-2 genome, it remains difficult to conclude whether the mutations identified in the viral genome are responsible for the improper detection of SARS-CoV-2 in the clinical setting.
Alternatively, certain mutations in the viral RNA genome may lead to changes in epitope structure and/or characteristics and thus result in the production of progenies with new antigenic determinants, as shown in the case of influenza virus.Citation79 If reinfection occurs, the antibodies produced in the previous (primary) infection may not be able to recognize the epitope of the new virus. In this case, host defense towards infection will start from the beginning (from innate to adaptive immune activation), which eventually will result in either a successful recovery or reinfection.
Previous studies have pointed out the possible involvement of viral evasion strategies in the pathogenesis of SARS-CoV-2 in humans.Citation8,Citation80 As seen in cases of HIV, HBV, and HCV, evasion of host defense is one of the strategies used by the virus to initiate a chronic state of infection.Citation81,Citation82 Chronic infection has been linked to the reemergence of infection or relapse state. Relapse is different from reinfection; relapse is described as a recurrent infection with the same type of pathogen that was present beforehand, whereas reinfection is defined as the emergence of infection with a different species or serologic strain of pathogen, as seen in other infectious diseases.Citation83,Citation84 It is yet unclear whether relapse or reinfection occurs in COVID-19 patients.
An experimental study to assess the possible occurrence of reinfection using rhesus macaque showed that primary SARS-CoV-2 infection could provide adequate protection against subsequent exposure.Citation85,Citation86 Nevertheless, while the current data from a SARS-CoV-2-infected animal model are likely to support protective immunity against reinfection, related studies are still in early stages. Based on experience from animal coronaviruses,Citation87 including other human coronaviruses,Citation88 it is tempting to speculate that reinfection is likely to occur in some people. This is practically possible in immunodeficient individuals with limited capacity to mount proper adaptive immunity and/or those that failed to produce sufficient memory cells.Citation89 However, recurrent infection may not be a general feature of COVID-19, as the cases reported so far are not particularly high.Citation20,Citation49,Citation50 Further experiments are needed to provide a definitive answer.
Implication in vaccine development
A vaccine aims to stimulate the cellular and humoral immunity of the adaptive immune system to recognize a pathogen and mount sufficient numbers of memory T and B cells as well as long lasting nAbs against the pathogen. Therefore, it is expected to protect the vaccinated individual from severe disease when infected by the specific pathogen the vaccine was designed for. Unlike cancer vaccines, which have also been used for immunotherapy (postphylaxis), vaccines against infectious disease-related pathogens are usually administered as a means of prevention (prephylaxis).Citation90
Vaccine candidates currently developed against SARS-CoV-2 come in different forms, which include the classic whole-virus vaccines (both inactivated and live attenuated vaccines), genetic vaccines in the form of DNA or RNA vaccines, viral vector vaccines, and protein-based vaccines in the forms of virus-like particles (VLPs) or subunit vaccines.Citation91 In order for a vaccine to be effective, several immunological factors have to be taken into account.
COVID-19 vaccine should induce protective and durable immunity
An effective vaccine ideally aims to prevent vaccinated individuals from target pathogens by stimulating the memory mechanism of protective humoral and cellular immunity; this protection should be long lasting. However, natural infection by coronaviruses does not usually result in long-lasting protective immunity, and antibodies wane within a few years or even months.Citation92 A similar phenomenon has been observed with immunity to SARS-CoV-2. Short duration of immune protection may allow reinfection by the same virus once the stimulated protective memory component of the adaptive immune system has depleted. Impaired immunity and disease severity in COVID-19 result from viral interference with type I interferon (IFN-I) synthesis.Citation93 IFN-I plays a critical role in the activation and maturation of the adaptive immune system, Citation94 and inhibition of IFN-I synthesis hinders the activation and maturation of B cells,Citation95–97 dendritic cells (DC),Citation98 and T cells.Citation99
Consequently, vaccine design should not only mimic natural SARS-CoV-2 infection through the administration of appropriate antigens, but vaccine components should also be engineered to stimulate stronger immune response both in magnitude and durability than what is achievable by natural infection. One possible way is by enhancing the innate immune response, especially by pathways regulating the adaptive immune response, such as, but not limited to, IFN-I synthesis.Citation100
Vaccine adjuvants have been shown to increase the durability of the immune response elicited by a SARS-CoV whole virus vaccine candidate. Different adjuvants including alum, CpG, Adva, and delta-inulin-based polysaccharide increased serum nAb titers and reduced lung virus titers in mice.Citation92 The use of alum in a yeast recombinant Hepatitis B (HBV), for example, had an increasing seroprotective effect in adults ≥40 years old.Citation101
Several antigen delivery systems currently used in the development of SARS-CoV-2 vaccine candidates may also enhance the innate and adaptive immune responses. Liposomes, for example, are synthetic phospholipid bilayers mimicking the plasma membrane of living cells. Liposomal delivery systems may be engineered to constitute phospholipid types, which act as intracellular messengers in modulating innate and adaptive immunity. Liposome surfaces can also be decorated with adjuvants or PAMP entities such as lipopolysaccharides (LPS) to stimulate the corresponding pattern recognition receptors (PRR), such as the toll-like receptors (TLR), which in return induces IFN-I expression.Citation102
DNA vaccine encoding SARS-CoV-2 protein utilizes the adenoviral vector (Ad) as a delivery system.Citation103 The Ad vector, for more or less than a decade, has been shown to activate or transduce DCs, macrophages, and natural killer (NK) cells. Ad vector also plays a role in IFN-I induction, which is important for the efficacy of Ad-based vaccines, and has a critical role in antigen presenting cell (APC) maturation and proinflammatory cytokine induction and regulation. IFN-I induction by Ad vector is mediated by the TLR9 and RIG-I receptors of the innate immune system.Citation104
VLP and other protein-based nanocages (PNC) are composed of an assembly of monomeric subunit proteins such as viral capsid proteins, or other protein subunits such as the small heat shock protein, forming a cage-like nanostructure.Citation105 VLP-based vaccines are either composed of native SARS-CoV-2 capsid proteinsCitation91 or a heterologous VLP platform presenting the SARS-CoV-2 spike protein on the surface.Citation106 The assembly of monomeric subunit proteins to form the VLP or PNC structure is advantageous in immune response enhancement in several ways. Firstly, the structural assembly provides a repetitive antigen presentation motif with high spatio-geometric density, which is known to enhance the immunogenicity and responsiveness of B cells.Citation107 Secondly, the repetitive nature of VLP and NPC assembly provides a PAMP motif allowing recognition by PRR, such as the TLR, and leading to IFN-I synthesis.Citation108 Thirdly, VLPs can activate DCs and enter APCs, including DC, to present antigens via the MHC Class I and MHC Class II pathways and therefore mediate the cytotoxic T cell and helper T cell immune response, respectively.Citation108–111 Small heat shock protein nanocages, without any additional antigen, induced the formation of bronchus-associated lymphoid tissue (iBALT) in the lung when inoculated intranasally and showed protection against lethal challenge induced by viral respiratory pathogens, including SARS-CoV.Citation112 A similar phenomenon was also observed when the inoculation with antigen-free papaya mosaic virus (PapMV) VLP showed protection against influenza or Streptococcus pneumoniae challenge in mice.Citation113 The observed protective immune response was mediated by the innate immunity, most likely involving neutrophils and CD11c+ cells.Citation113
HPV vaccines are made of virus-like particles (VLP) assembled from recombinant HPV coat proteins (L1). The fact that the vaccine is highly immunogenic and is very effective in preventing infection long-termCitation114,Citation115 may be due to the immunogenic properties of the close geometrical spacing of the repetitive antigen presented on the VLP.Citation90,Citation116 However, the L1-based VLP vaccine does not have an effective post-exposure therapeutic effect to clear HPV infected tumour cells. HPV-associated tumour progression is the result of HPV DNA integration into the host cell DNA, inactivating some HPV genes, including L1, and upregulating HPV E6 and E7 protein expression, which play a role in host cell transformation into tumour cells. The inactivation of L1 protein hinders their presentation via the MHC Class I and MHC Class II pathways on the infected host cells, and as a consequence, also the L1-based vaccine stimulated cytotoxic and helper T cell clearance.Citation117 To circumvent this, efforts have been made to develop vaccines using the continuously expressed E6 and E7 as antigens.Citation118 In the context of VLP-based vaccine, the encapsidation technologyCitation119 may be useful to ensure entry of the E6 and E7 antigens into APCs and to prevent extracellular antibody neutralisation.
The ability of pathogenic viral proteins to interfere with the innate and adaptive immune response in the form of altered gene expression conferring tolerance or other forms of immune evasion has been observed.Citation120,Citation121 For the coronaviruses, several structural and non-structural proteins which play a role in pathways leading to IFN-I and III expression have been identified,Citation122 including nsp16,Citation123 membrane,Citation124,Citation125 nucleocapsid,Citation126 amongst others. Some of these immune evasive proteins have a high homology with SARS-CoV-2 proteins.Citation127 Recent literature suggested that the non-structural proteinCitation128 and the ORF3 proteinCitation129 of the virus may interfere with the immune response. Therefore, the inclusion of these proteins in a vaccine design should be evaluated.
In addition, a strong antibody response and T cell response to ensure protective and enduring immunity requires prolonged antigen exposure to B and T cells in a dose-escalating manner.Citation130,Citation131 Accordingly, COVID-19 vaccine engineering may utilize a controlled-release delivery technology, and repeated vaccination in a dose-escalating manner may also provide a similar benefit.
COVID-19 vaccine should stimulate humoral and cellular immunity
Protective immunity against a pathogen usually depends on the availability of nAbs and effector T cells at the time of infection.Citation132 To induce protective immunity, vaccines are primarily designed to induce nAb expression by the adaptive immune system and the corresponding formation of memory B cells that can be rapidly activated after antigen reexposure.Citation130 The function of nAbs is to prevent viral interaction with the host, and hence, much work has been done in vaccine development to evaluate the generation of antibodies capable of binding to the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein. The RBD plays a role in viral binding to the angiotensin-converting enzyme-2 (ACE-2) receptor in a variety of host cell types.Citation133,Citation134 Antibodies that specifically bind to this domain prevent the entry of the SARS-CoV-2 into the host cells and therefore exhibit neutralizing properties.Citation135
The generation of memory B cells and nondividing bone marrow-resident plasma cells for long-term antibody production protective for the next bout of infection requires antigenic multivalency and antigenic threshold. Repetitive antigen and a longer duration of antigen exposure during natural infection or vaccination may increase the activation of germinal centers and thus the number of activated T and B cells, and hence, increased amount of long-lived plasma cells as well as memory B and T cells.Citation90,Citation116
A recent study also pointed out that SARS-CoV-2 could enter T cells with an alternative receptor, possibly CD147, through a yet unidentified region on its spike protein.Citation136 Hence, identification of this binding region and its corresponding nAbs would be important in preventing T cell infection and consequently T cell response impairment.
Stimulation of cellular immunity through corresponding T cell activation should be targeted concurrently with the generation of antibody-related responses against SARS-CoV-2. Single-cell RNA sequencing analysis of the bronchoalveolar lavage fluid (BALF) from COVID-19 patients demonstrated the crucial role of cytotoxic T cells (Tc) during recovery from COVID-19.Citation137 Clonally expanded effector Tc of homogenous transcriptional profile with tissue-resident characteristics and the upregulation of genes associated with activation, migration, and the cytokine pathway are the hallmarks of Tc population in the BALF from moderate COVID-19 patients. However, severe/critical patients display an ensemble of highly proliferative T cells with heterogeneous transcriptional profiles and upregulation of genes associated with translation initiation, cell homeostasis, and nucleoside metabolic pathway.Citation137 Moderate COVID-19 cases are also marked by elevated T cell recruitment into the lung, whereas chemokine recruitment favors inflammatory monocytes and neutrophils into the lungs in severe/critical cases.Citation137 Analysis of Tc responses in COVID-19 patients revealed that the majority of convalescent individuals generate Tc responses, responding to Tc epitopes from the spike (26%), membrane (22%), nonstructural protein (15%), nucleocapsid protein (12%), ORF8 (10%), and ORF3a (7%).Citation138 Consequently, in addition to the spike protein, a vaccine designed to concurrently target Tc response should also consider the delivery of nonspike SARS-CoV-2 proteins related to Tc responses.
Analysis of helper T cell response revealed their important role in patient recovery from COVID-19, with T helper 1 (Th1) showing the prominent response.Citation138,Citation139 Unlike the non-spike Th response, spike-specific Th response correlated well with the magnitude of anti-RBD spike protein IgG and IgA responses. Th responses were mostly directed towards the spike (27%), membrane (21%), and nucleocapsid protein (11%) of SARS-CoV-2.Citation138 As Th response plays a significant role in B cell maturation to generate nAbs, the raising of Th response should be considered in vaccine design for COVID-19.
In order for a vaccine to generate a T cell response, T cell-associated antigens need to be delivered into the host cellular compartment to stimulate the cellular immune response. Moreover, it would be advantageous to minimize the generation of antibodies against T-cell associated antigens, which may prevent the corresponding antigens from reaching the host cellular compartment during natural infection. This may be achieved by encapsulating the T cell-associated antigen vaccine component inside the relevant delivery system (such as in the use of VLPsCitation119 and liposomes) or engineer their expression inside the host cell (such as in the use of RNA and DNA vaccines). Moreover, T cell epitopes are associated with HLA, and therefore, T-cell based vaccine candidates should be designed to encompass the majority of the global population.Citation140
Immunity works by exerting both the humoral (antibody-based) and cellular (T cell-based) immunity. In vaccine development, the correlate of protection should be determined and the presence of immune-related adverse events should be tested and avoided.
The RNA polymerase of RNA vaccines such as SARS-CoV-2 are known to be error-prone, and therefore, may introduce frequent mutations.Citation141 Although the mutational rate of SARS-CoV-2 is lower than influenza,Citation142 a future mutation altering the conformational structure of nAb recognition sites of the SARS-CoV-2 protein may result in possible reinfection. Furthermore, a vaccine designed to stimulate cross-protective immunity would be a good strategy to prevent or reduce the severity of disease from other possible pandemic coronavirus strains in the future.Citation143
A cross-protective vaccine may utilize conserved T and B cell epitopes to stimulate cross-protective immunity.Citation144,Citation145 Conserved or partially conserved T cell epitopes are commonly easier to identify and engineer than conserved B cell epitopes due to the conformation-dependent nature of B cell epitopes.Citation146,Citation147
A natural T cell response may not prevent infection but may reduce the severity of the disease.Citation148 Therefore, even if reinfection occurs in a state of reduced antibody levels, there is a possibility that pre-existing T cell immunity may prevent clinically severe disease. Moreover, conserved or partially conserved T cell epitopes from other circulating coronaviruses including those that cause the common cold may have provided a broadly cross-protective T cell immunity against SARS-CoV-2 in unexposed individuals. This may explain why a significant number of SARS-CoV-2 positively tested individuals are asymptomatic/presymptomatic or show only mild symptoms or moderate disease in SARS-CoV-2.Citation149 However, it has to be kept in mind that during the period of natural SARS-CoV-2 infection, with or without symptoms, a patient may still transmit the virus to other people in their vicinity, who may be at higher risk in contracting a more severe disease.
A T cell-based SARS-CoV-2 vaccine utilizing conserved T cell epitopes shared amongst coronavirus strains (those causing common cold and also the more severe disease such as SARS and MERS), may not only confer some protection against SARS-CoV-2 in terms of reducing disease severity but may also protect from other potential coronavirus pandemic strains in the future. These T cell-based vaccines can be used as a prepandemic vaccine or administered in the early phase of a pandemic to reduce health and disease burden before an antibody-based vaccine becomes available.Citation30,Citation150
Studies on Th responses have identified SARS-CoV-2 epitopes that have also been found in previously circulating human coronaviruses,Citation149,Citation151 which may play a role in the immune protection of unexposed and recovered patients of COVID-19. Conserved antibody recognition siteCitation152 and cross-protective Th epitopes on the SARS-CoV-2 S proteinCitation149,Citation151 have been identified and their use in vaccine design may help from severe re-infection of SARS-CoV-2 or infection from a future coronavirus related pandemic.
Nonetheless, T cell immunity may only protect to a certain extent, and therefore, a combination with antibody-based immunity will be the best approach in vaccine development. Moreover, there are additional factors to consider, such as ruling out immune-related adverse events (irAE)Citation153 and determining the correlates of protection of both T cell-based and antibody-based immunity.Citation90 Epidemiologically, a current estimate of 70% vaccinated individual in the population will result in herd immunity and thus reduce the transmission of the virus in the population to a negligible level.Citation154–156
Conclusion and future perspectives
The transmission of infectious viral particles from one host to another usually triggers vigorous host immune responses leading to the clearance of viral particles. However, in some cases, the virus cannot be cleared off, resulting in either the death of the infected host or the emergence of persistent infection. In another scenario, host immunity can remove the viral particles in the primary infection but fails to protect the host from reinfection. There is still limited knowledge about whether the reported reinfection cases of COVID-19 were accurately described as reinfection or whether they were simply due to the failure of currently available methods to detect the low copy number of SARS-CoV-2 in certain patients, which led to a relapse. To improve our understanding on this issue, suitable model organisms to investigate the nature of reinfection and elucidate its possible mechanistic basis are urgently required.
Due to its insidious life cycle in the host cells, SARS-CoV-2 can achieve continuous transmission among humans. For susceptible hosts such as immunodeficient, older people, and those with associated comorbidities, SARS-CoV-2 primary infection increases the risk of death. To avoid such outcome, effective and safe vaccination to achieve herd immunity in the given population is the best option to adopt. However, with the challenges associated with development of an effective COVID-19 vaccine and the probability of reinfection by SARS-CoV-2, if that is truly possible, the risk of death of susceptible hosts may persist. In such conditions, avoidance of reinfection is the only available option, however difficult it might be to achieve.
Disclosure of potential conflicts of interest
No potential conflicts of interest were disclosed.
Human subjects approval statement
Not required.
Additional information
Funding
References
- Rodriguez-Morales AJ, Cardona-Ospina JA, Gutierrez-Ocampo E, Villamizar-Pena R, Holguin-Rivera Y, Escalera-Antezana JP, Alvarado-Arnez LE, Bonilla-Aldana DK, Franco-Paredes C, Henao-Martinez AF, et al. Clinical, laboratory and imaging features of COVID-19: a systematic review and meta-analysis. Travel Med Infect Dis. 2020;34:101623. doi:https://doi.org/10.1016/j.tmaid.2020.101623.
- Harapan H, Itoh N, Yufika A, Winardi W, Keamg S, Te H, Megawati D, Hayati Z, Wagner AL, Mudatsir M, et al. Coronavirus disease 2019 (COVID-19): a literature review. J Infect Public Health. 2020;13(5):667–73. doi:https://doi.org/10.1016/j.jiph.2020.03.019.
- Dong E, Du H, Gardner L. An interactive web-based dashboard to track COVID-19 in real time. Lancet Infect Dis. 2020;20(5):533–34. doi:https://doi.org/10.1016/S1473-3099(20)30120-1.
- Chan JF-W, Zhang AJ, Yuan S, Poon VK-M, Chan CC-S, Lee AC-Y, Chan W-M, Fan Z, Tsoi H-W, Wen L, et al. Simulation of the clinical and pathological manifestations of Coronavirus Disease 2019 (COVID-19) in golden Syrian hamster model: implications for disease pathogenesis and transmissibility. Clin Infect Dis. 2020. doi:https://doi.org/10.1093/cid/ciaa325.
- Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si H-R, Zhu Y, Li B, Huang C-L, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270–73. doi:https://doi.org/10.1038/s41586-020-2012-7.
- Lam TT, Shum MH, Zhu HC, Tong YG, Ni XB, Liao YS, Tong Y-G, Shi Y-X, Ni X-B, Liao Y-S, et al. Identifying SARS-CoV-2 related coronaviruses in Malayan pangolins. Nature. 2020;583(7815):282–85. doi:https://doi.org/10.1038/s41586-020-2169-0.
- Zhang T, Wu Q, Zhang Z. Probable pangolin origin of SARS-CoV-2 associated with the COVID-19 outbreak. Curr Biol. 2020;30(8):1578. doi:https://doi.org/10.1016/j.cub.2020.03.063.
- Keam S, Megawati D, Patel S, Tiwari R, Dhama K, Harapan H. Immunopathology and immunotherapeutic strategies in SARS-CoV-2 infection. Rev Medical Virol. 2020;30(5). doi:https://doi.org/10.1002/rmv.2123.
- Frediansyah A, Nainu F, Dhamad K, Mudatsir M, Harapan H. Remdesivir and its antiviral activity against COVID-19: a systematic review. Clin Epidemiol Glob Health. 2020. (In press) doi:https://doi.org/10.1016/j.cegh.2020.07.011
- Frediansyah A, Tiwari R, Sharun K, Dhama K, Harapan H. Antivirals for COVID-19: a critical review. Clin Epidemiol Global Health. 2020. doi:https://doi.org/10.1016/j.cegh.2020.07.006.
- Sharun K, Tiwari R, Iqbal Yatoo M, Patel SK, Natesan S, Dhama J, Malik YS, Harapan H, Singh RK, Dhama K, et al. Antibody-based immunotherapeutics and use of convalescent plasma to counter COVID-19: advances and prospects. Expert Opin Biol Ther. 2020;20(9):1033–46. doi:https://doi.org/10.1080/14712598.2020.1796963.
- Bao L, Deng W, Gao H, Xiao C, Liu J, Xue J, Lv Q, Liu J, Yu P, Xu Y, et al. Reinfection could not occur in SARS-CoV-2 infected rhesus macaques. BioRxiv. 2020. doi:https://doi.org/10.1101/2020.03.13.990226.
- Le T T, Andreadakis Z, Kumar A, Gomez Roman R, Tollefsen S, Saville M, Mayhew S. The COVID-19 vaccine development landscape. Nat Rev Drug Discov. 2020; 19:305-306. doi:https://doi.org/10.1038/d41573-020-00073-5.
- Lurie N, Saville M, Hatchett R, Halton J. Developing Covid-19 vaccines at pandemic speed. N Engl J Med. 2020;382(21):1969–73. (In press). doi:https://doi.org/10.1056/NEJMp2005630.
- Harapan H, Wagner AL, Yufika A, Winardi W, Anwar S, Gan AK, Setiawan AM, Rajamoorthy Y, Sofyan H, Mudatsir M, et al. Acceptance of a COVID-19 vaccine in Southeast Asia: a cross-sectional study in Indonesia. Front Public Health. 2020;8:381. doi:https://doi.org/10.3389/fpubh.2020.00381.
- Harapan H, Wagner A, Yufika A, Winardi W, Anwar S, Gan A, Setiawan AM, Rajamoorthy Y, Sofyan H, Vo TQ, et al. Willingness to pay for a COVID-19 vaccine in Indonesia. Hum Vacc Immunother. 2020;1–7. (Sumbitted). doi:https://doi.org/10.1080/21645515.2020.1819741.
- Smith J South Korea reports more recovered coronavirus patients testing positive again. April 13, 2020.
- Korean CDC. Findings from investigation and analysis of re-positive cases. 19 May 2020.
- Ni L, Ye F, Cheng M-L, Feng Y, Deng Y-Q, Zhao H, Wei P, Ge J, Gou M, Li X, et al. Detection of SARS-CoV-2-specific humoral and cellular immunity in COVID-19 convalescent individuals. Immunity. 2020;52(6):971–977.e3. doi:https://doi.org/10.1016/j.immuni.2020.04.023.
- Qin C, Zhou L, Hu Z, Zhang S, Yang S, Tao Y, Xie C, Ma K, Shang K, Wang W, et al. Dysregulation of immune response in patients with coronavirus 2019 (COVID-19) in Wuhan, China. Clin Infect Dis. 2020;71(15):762–68. doi:https://doi.org/10.1093/cid/ciaa248.
- Lan L, Xu D, Ye G, Xia C, Wang S, Li Y, Xu H. Positive RT-PCR test results in patients recovered from COVID-19. Jama. 2020;323(15):1502–03. doi:https://doi.org/10.1001/jama.2020.2783.
- Da Guan W, Mok CKP, Chen ZL, Feng LQ, Li ZT, Huang JC, Ke CW, Deng X, Ling Y, Wu SG. Characteristics of traveler with Middle East respiratory syndrome, China, 2015. Emerg Infect Dis. 2015;21(12):2278. doi:https://doi.org/10.3201/eid2112.151232.
- Liu Z-Y, Li T-S, Wang Z, Xu Z-J, Wang H-L, Yu Y, Du T-K, Bai Y, Qiu Z-F, Lu W, et al. Clinical features and therapy of 106 cases of severe acute respiratory syndrome. Zhonghua Nei Ke Za Zhi. 2003;42:373–77.
- MO H, ZENG G, REN X, LI H, KE C, TAN Y, CAI C, LAI K, CHEN R, CHAN-YEUNG M, et al. Longitudinal profile of antibodies against SARSΓÇÉcoronavirus in SARS patients and their clinical significance. Respirology. 2006;11(1):49–53. doi:https://doi.org/10.1111/j.1440-1843.2006.00783.x.
- Alagaili AN, Briese T, Mishra N, Kapoor V, Sameroff SC, Burbelo PD, de Wit E, Munster VJ, Hensley LE, Zalmout IS, et al. Middle East respiratory syndrome coronavirus infection in dromedary camels in Saudi Arabia. MBio. 2014;5(2):e00884–14. doi:https://doi.org/10.1128/mBio.01002-14.
- Hemida MG, Chu DK, Poon LL, Perera RA, Alhammadi MA, Ng H-Y, Siu LY, Guan Y, Alnaeem A, Peiris M, et al. MERS coronavirus in dromedary camel herd, Saudi Arabia. Emerg Infect Dis. 2014;20(7):1231. doi:https://doi.org/10.3201/eid2007.140571.
- Roberts A, Vogel L, Guarner J, Hayes N, Murphy B, Zaki S, Subarao K. Severe acute respiratory syndrome coronavirus infection of golden Syrian hamsters. J Virol. 2005;79(1):503–11. doi:https://doi.org/10.1128/JVI.79.1.503-511.2005.
- Subbarao K, McAuliffe J, Vogel L, Fahle G, Fischer S, Tatti K, Packard M, Shieh W-J, Zaki S, Murphy B, et al. Prior infection and passive transfer of neutralizing antibody prevent replication of severe acute respiratory syndrome coronavirus in the respiratory tract of mice. J Virol. 2004;78(7):3572–77. doi:https://doi.org/10.1128/JVI.78.7.3572-3577.2004.
- McAuliffe J, Vogel L, Roberts A, Fahle G, Fischer S, Shieh W-J, Butler E, Zaki S, St. Claire M, Murphy B, et al. Replication of SARS coronavirus administered into the respiratory tract of African Green, rhesus and cynomolgus monkeys. Virology. 2004;330(1):8–15. doi:https://doi.org/10.1016/j.virol.2004.09.030.
- Woodland DL. Progress towards a SARS vaccine. Viral Immunol. 2010;23(5):455. doi:https://doi.org/10.1089/vim.2010.ed.23.5.
- Excler JL, Delvecchio CJ, Wiley RE, Williams M, Yoon IK, Modjarrad K, Boujelal M, Moorthy VS, Hersi AS, Kim JH, et al. Toward developing a preventive MERS-CoV vaccine-report from a workshop organized by the Saudi Arabia ministry of health and the international vaccine institute, Riyadh, Saudi Arabia, November 14–15, 2015. Emerg Infect Dis. 2016;22(8): e160229. doi: https://doi.org/10.3201/eid2208.160229.
- Chen D, Xu W, Lei Z, Huang Z, Liu J, Gao Z, Peng L. Recurrence of positive SARS-CoV-2 RNA in COVID-19: a case report. Int J Infect Dis. 2020;93:297–99. doi:https://doi.org/10.1016/j.ijid.2020.03.003.
- Lippi G, Simundic A-M, Plebani M. Potential preanalytical and analytical vulnerabilities in the laboratory diagnosis of coronavirus disease 2019 (COVID-19). Clin Chem Lab Med. 2020;58(7):1070-1076. doi:https://doi.org/10.1515/cclm-2020-0285.
- Chan JF, Yuan S, Kok KH, To KK, Chu H, Yang J, Xing F, Liu J, Yip CCY, Poon RWS, et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020;395(10223):514–23. doi:https://doi.org/10.1016/S0140-6736(20)30154-9.
- Menggi HYT, Wang S, Chen S, Zhou W, Chen D, Zhou L, Wang M, Zhao Y, Zeng W, Huang Q, et al. Clinical characteristics of laboratory confirmed positive cases of SARS-CoV-2 infection in Wuhan, China: a retrospective single center analysis. Travel Med Infect Dis. 2020;36:101606. doi:https://doi.org/10.1016/j.tmaid.2020.1016062020.
- Xie X, Zhong Z, Zhao W, Zheng C, Wang F, Liu J. Chest CT for typical 2019-nCoV pneumonia: relationship to negative RT-PCR testing. Radiology. 2020;296(2):E41-E45. doi: https://doi.org/10.1148/radiol.2020200343.
- CDC. Coronavirus Disease 2019 (COVID-19). https://www.cdc.gov/coronavirus/2019-ncov/hcp/disposition-hospitalized-patients.html, 2020.
- Lippi G, Simundic A-M, Plebani M. Potential preanalytical and analytical vulnerabilities in the laboratory diagnosis of coronavirus disease 2019 (COVID-19). Clin Chem Lab Med. 2020;58(7):1070. doi:https://doi.org/10.1515/cclm-2020-0285.
- Pulendran B, Oh JZ, Nakaya HI, Ravindran R, Kazmin DA. Immunity to viruses: learning from successful human vaccines. Immunol Rev. 2013;255(1):243–55. doi:https://doi.org/10.1111/imr.12099.
- Tay MZ, Poh CM, Rénia L, MacAry PA, Ng LFP. The trinity of COVID-19: immunity, inflammation and intervention. Nat Rev Immunol. 2020;20(6):363–74. doi:https://doi.org/10.1038/s41577-020-0311-8.
- Vabret N, Britton GJ, Gruber C, Hegde S, Kim J, Kuksin M, Levantovsky R, Malle L, Moreira A, Park MD, et al. Immunology of COVID-19: current state of the science. Immunity. 2020;52(6):910–41. doi:https://doi.org/10.1016/j.immuni.2020.05.002.
- Chaplin DD. Overview of the immune response. J Allergy Clin Immunol. 2010;125(2):S3–S23. doi:https://doi.org/10.1016/j.jaci.2009.12.980.
- Petrosillo N, Viceconte G, Ergonul O, Ippolito G, Petersen E. COVID-19, SARS and MERS: are they closely related? Clin Microbiol Infect. 2020;S1198–743X:30171–73.
- Wang L, Wang Y, Ye D, Liu Q. Review of the 2019 novel coronavirus (SARS-CoV-2) based on current evidence. Int J Antimicrob Agents. 2020;55(6):105948. doi:https://doi.org/10.1016/j.ijantimicag.2020.105948.
- Aoshi T, Koyama S, Kobiyama K, Akira S, Ishii KJ. Innate and adaptive immune responses to viral infection and vaccination. Curr Opin Virol. 2011;1(4):226–32. doi:https://doi.org/10.1016/j.coviro.2011.07.002.
- Rouse BT, Sehrawat S. Immunity and immunopathology to viruses: what decides the outcome? Nat Rev Immunol. 2010;10(7):514–26. doi:https://doi.org/10.1038/nri2802.
- Ju B, Zhang Q, Ge X, Wang R, Yu J, Shan S, Zhou B, Song S, Tang X, Yu J, et al. Potent human neutralizing antibodies elicited by SARS-CoV-2 infection. bioRxiv. 2020;990770.
- Viroj JBW. Hemorrhagic problem among the patients with COVID-19: clinical summary of 41 thai infected patients. J Sage: Clin Appl Thrombosis/Hemostasis. 2020;26:1076029620918308. doi:https://doi.org/10.1177/1076029620918308.
- Peng J, Wang M, Zhang G, Lu E. Seven discharged patients turning positive again for SARS-CoV-2 on quantitative RT-PCR. Am J Infect Control. 2020;S0196–6553:30198–X.
- Tang X, Zhao S, He D, Yang L, Wang MH, Li Y, Mei S, Zou X. Positive RT-PCR tests among discharged COVID-19 patients in Shenzhen, China. Infect Control Hosp Epidemiol. 2020;1–2. doi:https://doi.org/10.1017/ice.2020.1232
- Chen G, Wu D, Guo W, Cao Y, Huang D, Wang H, Wang T, Zhang X, Chen H, Yu H, et al. Clinical and immunological features of severe and moderate coronavirus disease 2019. J Clin Invest. 2020;130(5):2620–29. doi:https://doi.org/10.1172/JCI137244.
- Li Y, Hu Y, Yu Y, Zhang X, Li B, Wu J, Li J, Wu Y, Xia X, Tang H, et al. Positive result of Sars-Cov-2 in faeces and sputum from discharged patient with COVID-19 in Yiwu, China. J Med Virol. 2020;92:1938-1947. doi:https://doi.org/10.1002/jmv.259050.
- Zheng H-Y, Zhang M, Yang C-X, Zhang N, Wang X-C, Yang X-P, Dong X-Q, Zheng Y-T. Elevated exhaustion levels and reduced functional diversity of T cells in peripheral blood may predict severe progression in COVID-19 patients. Cell Mol Immunol. 2020;17(5):541–43. doi:https://doi.org/10.1038/s41423-020-0401-3.
- Tan L, Wang Q, Zhang D, Ding J, Huang Q, Tang Y-Q, Wang Q, Miao H. Lymphopenia predicts disease severity of COVID-19: a descriptive and predictive study. Signal Transduction Targeted Ther. 2020;5(1):33. doi:https://doi.org/10.1038/s41392-020-0148-4.
- Chen X, Zhao B, Qu Y, Chen Y, Xiong J, Feng Y, Men D, Huang Q, Liu Y, Yang B, et al. Detectable serum SARS-CoV-2 viral load (RNAaemia) is closely correlated with drastically elevated interleukin 6 (IL-6) level in critically ill COVID-19 patients. Clin Infect Dis. 2020. doi:https://doi.org/10.1093/cid/ciaa449.
- Diao B, Wang C, Tan Y, Chen X, Liu Y, Ning L, Chen L, Li M, Liu Y, Wang G, et al. Reduction and functional exhaustion of T cells in patients with coronavirus disease 2019 (COVID-19). medRxiv. 2020;20024364. doi:https://doi.org/10.1101/2020.02.18.20024364.
- Wan S, Yi Q, Fan S, Lv J, Zhang X, Guo L, Lang C, Xiao Q, Xiao K, Yi Z, et al. Characteristics of lymphocyte subsets and cytokines in peripheral blood of 123 hospitalized patients with 2019 novel coronavirus pneumonia (NCP). medRxiv. 2020 Feb 10. 20021832. doi:https://doi.org/10.1101/2020.02.10.20021832.
- Yang M. Cell pyroptosis, a potential pathogenic mechanism of 2019-nCoV infection. SSRN. 2020. doi:https://doi.org/10.2139/ssrn.3527420.
- Biggs JR, Zhang D-E. Chapter 16 - molecular basis of lymphoid and myeloid diseases. In: Coleman WB, Tsongalis GJ, editors. Molecular pathology. Seconded. Academic Press;2018. p.299–328.
- Liao M, Liu Y, Yuan J, Wen Y, Xu G, Zhao J, Cheng L, Li J, Wang X, Wang F, et al. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat Med. 2020;26(6):842–44. doi:https://doi.org/10.1038/s41591-020-0901-9.
- Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, Liu S, Zhao P, Liu H, Zhu L, et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med. 2020;8(4):420–22. doi:https://doi.org/10.1016/S2213-2600(20)30076-X.
- Newton K, Dixit VM. Signaling in innate immunity and inflammation. Cold Spring Harb Perspect Biol. 2012;4(3):a006049. doi:https://doi.org/10.1101/cshperspect.a006049.
- Amarante-Mendes GP, Adjemian S, Branco LM, Zanetti LC, Weinlich R, Bortoluci KR. Pattern recognition receptors and the host cell death molecular machinery. Front Immunol. 2018;9:2379. doi:https://doi.org/10.3389/fimmu.2018.02379.
- Tang D, Kang R, Coyne CB, Zeh HJ, Lotze MT. PAMPs and DAMPs: signal 0s that spur autophagy and immunity. Immunol Rev. 2012;249:158–75.
- Vardhana SA, Wolchok JD. The many faces of the anti-COVID immune response. J Exp Med. 2020;217(6):e20200678. doi:https://doi.org/10.1084/jem.20200678.
- Wherry EJ. T cell exhaustion. Nat Immunol. 2011;12(6):492–99. doi:https://doi.org/10.1038/ni.2035.
- Saeidi A, Zandi K, Cheok YY, Saeidi H, Wong WF, Lee CYQ, Cheong HC, Yong YK, Larsson M, Shankar EM. T-cell exhaustion in chronic infections: reversing the state of exhaustion and reinvigorating optimal protective immune responses. Front Immunol. 2018;9:2569. doi:https://doi.org/10.3389/fimmu.2018.02569.
- Callow KA, Parry HF, Sergeant M, Tyrrell DA. The time course of the immune response to experimental coronavirus infection of man. Epidemiol Infect. 1990;105(2):435–46. doi:https://doi.org/10.1017/S0950268800048019.
- Wu L-P, Wang N-C, Chang Y-H, Tian X-Y, Na D-Y, Zhang L-Y, Zheng L, Lan T, Wang L-F, Liang G-D, et al. Duration of antibody responses after severe acute respiratory syndrome. Emerg Infect Dis. 2007;13(10):1562–64. doi:https://doi.org/10.3201/eid1310.070576.
- Li CK-F, Wu H, Yan H, Ma S, Wang L, Zhang M, Tang X, Temperton NJ, Weiss RA, Brenchley JM, et al. T cell responses to whole SARS coronavirus in humans. J Immunol. 2008;181(8):5490–500. doi:https://doi.org/10.4049/jimmunol.181.8.5490.
- Park WB, Perera RAPM, Choe PG, Lau EHY, Choi SJ, Chun JY, Oh HS, Song K-H, Bang JH, Kim ES, et al. Kinetics of serologic responses to MERS coronavirus infection in humans, South Korea. Emerg Infect Dis. 2015;21(12):2186–89. doi:https://doi.org/10.3201/eid2112.151421.
- Ko J-H, Müller MA, Seok H, Park GE, Lee JY, Cho SY, Ha YE, Baek JY, Kim SH, Kang J-M, et al. Serologic responses of 42 MERS-coronavirus-infected patients according to the disease severity. Diagn Microbiol Infect Dis. 2017;89(2):106–11. doi:https://doi.org/10.1016/j.diagmicrobio.2017.07.006.
- Huang AT, Garcia-Carreras B, Hitchings MDT, Yang B, Katzelnick L, Rattigan SM, Borgert B, Moreno C, Solomon BD, Rodriguez-Barraquer I, et al. A systematic review of antibody mediated immunity to coronaviruses: antibody kinetics, correlates of protection, and association of antibody responses with severity of disease. medRxiv. 2020;20065771. doi:https://doi.org/10.1101/2020.04.14.20065771.
- Cheng MP, Papenburg J, Desjardins M, Kanjilal S, Quach C, Libman M, Dittrich S, Yansouni CP. Diagnostic testing for severe acute respiratory syndrome–related coronavirus-2. Ann Intern Med. 2020;M20-1301. doi:https://doi.org/10.7326/M20-1301.
- Mohammadi A, Esmaeilzadeh E, Li Y, Bosch RJ, Li JZ. SARS-CoV-2 detection in different respiratory sites: a systematic review and meta-analysis. EBioMedicine. 2020;59:102903. doi:https://doi.org/10.1016/j.ebiom.2020.102903.
- Tang Y-W, Schmitz JE, Persing DH, Stratton CW, McAdam AJ. Laboratory diagnosis of COVID-19: current issues and challenges. J Clin Microbiol. 2020;58(6):e00512–20. doi:https://doi.org/10.1128/JCM.00512-20.
- Barr JN, Fearns R. How RNA viruses maintain their genome integrity. J Gen Virol. 2010;91(6):1373–87. doi:https://doi.org/10.1099/vir.0.020818-0.
- Khailany RA, Safdar M, Ozaslan M. Genomic characterization of a novel SARS-CoV-2. Gene Rep. 2020;19:100682. doi:https://doi.org/10.1016/j.genrep.2020.100682.
- Sanjuán R, Domingo-Calap P. Mechanisms of viral mutation. Cell Mol Life Sci. 2016;73(23):4433–48. doi:https://doi.org/10.1007/s00018-016-2299-6.
- Kumar S, Nyodu R, Maurya VK, Saxena SK. Host immune response and immunobiology of human SARS-CoV-2 infection. Coronavirus Dis 2019 (COVID-19). 2020;43–53.
- Ortega-Prieto AM, Dorner M. Immune evasion strategies during chronic hepatitis B and C virus infection. Vaccines (Basel). 2017;5:24.
- Guha D, Ayyavoo V. Innate immune evasion strategies by human immunodeficiency virus type 1. Isrn Aids. 2013;2013:954806. doi:https://doi.org/10.1155/2013/954806.
- Guerra-Assunção JA, Houben RMGJ, Crampin AC, Mzembe T, Mallard K, Coll F, Khan P, Banda L, Chiwaya A, Pereira RPA, et al. Recurrence due to relapse or reinfection with Mycobacterium tuberculosis: a whole-genome sequencing approach in a large, population-based cohort with a high HIV infection prevalence and active follow-up. J Infect Dis. 2015;211:1154–63. doi:https://doi.org/10.1093/infdis/jiu574.
- Brown JR, Roy S, Tutill H, Williams R, Breuer J. Super-infections and relapses occur in chronic norovirus infections. J Clin Virol. 2017;96:44–48. doi:https://doi.org/10.1016/j.jcv.2017.09.009.
- Bao L, Deng W, Gao H, Xiao C, Liu J, Xue J, Lv Q, Liu J, Yu P, Xu Y, et al. Reinfection could not occur in SARS-CoV-2 infected rhesus macaques. BioRxiv. 2020. doi:https://doi.org/10.1101/2020.03.13.990226.
- Chandrashekar A, Liu J, Martinot AJ, McMahan K, Mercado NB, Peter L, Tostanoski LH, Yu J, Maliga Z, Nekorchuk M, et al. SARS-CoV-2 infection protects against rechallenge in rhesus macaques. Science. 2020;369(6505):812-817. doi:https://doi.org/10.1126/science.abc4776.
- Saif LJ. Bovine respiratory coronavirus. Vet Clin North Am Food Anim Pract. 2010;26(2):349–64. doi:https://doi.org/10.1016/j.cvfa.2010.04.005.
- Monto AS, Lim SK. The tecumseh study of respiratory illness. VI. Frequency of and relationship between outbreaks of coronavirus infection. J Infect Dis. 1974;129(3):271–76. doi:https://doi.org/10.1093/infdis/129.3.271.
- Gaspar HB, Goldblatt D. Immunodeficiency syndromes and recurrent infection. Br J Hosp Med. 1997;58:565–68.
- Plotkin SA. Vaccines: past, present and future. Nat Med. 2005;11(S4):S5–11. doi:https://doi.org/10.1038/nm1209.
- WHO. Draft landscape of COVID-19 candidate vaccines. Geneva: World Health Organization; 2020.
- Enjuanes L, Zuniga S, Castano-Rodriguez C, Gutierrez-Alvarez J, Canton J, Sola I. Molecular basis of coronavirus virulence and vaccine development. Adv Virus Res. 2016;96:245–86.
- Blanco-Melo D, Nilsson-Payant BE, Liu WC, Uhl S, Hoagland D, Moller R, Jordan TX, Oishi K, Panis M, Sachs D, et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell. 2020;181(5):1036–45e9. doi:https://doi.org/10.1016/j.cell.2020.04.026.
- Janeway CA Jr., Medzhitov R. Innate immune recognition. Annu Rev Immunol. 2002;20(1):197–216. doi:https://doi.org/10.1146/annurev.immunol.20.083001.084359.
- Crampton SP, Voynova E, Bolland S. Innate pathways to B-cell activation and tolerance. Ann N Y Acad Sci. 2010;1183(1):58–68. doi:https://doi.org/10.1111/j.1749-6632.2009.05123.x.
- Cyster JG, Allen CDC. B cell responses: cell interaction dynamics and decisions. Cell. 2019;177(3):524–40. doi:https://doi.org/10.1016/j.cell.2019.03.016.
- Khodadadi L, Cheng Q, Radbruch A, Hiepe F. The maintenance of memory plasma cells. Front Immunol. 2019;10:721.
- Munz C, Steinman RM, Fujii S. Dendritic cell maturation by innate lymphocytes: coordinated stimulation of innate and adaptive immunity. J Exp Med. 2005;202(2):203–07. doi:https://doi.org/10.1084/jem.20050810.
- Shin KS, Jeon I, Kim BS, Kim IK, Park YJ, Koh CH, Song B, Lee J-M, Lim J, Bae E-A, et al. Monocyte-derived dendritic cells dictate the memory differentiation of CD8(+) T cells during acute infection. Front Immunol. 2019;10:1887. doi:https://doi.org/10.3389/fimmu.2019.01887.
- Iwasaki A, Medzhitov R. Regulation of adaptive immunity by the innate immune system. Science. 2010;327(5963):291–95. doi:https://doi.org/10.1126/science.1183021.
- Gellin BG, Greenberg RN, Hart RH, Bertino, Jr. JS Jr., Stein DH, Deloria MA, Clements‐Mann ML. Immunogenicity of two doses of yeast recombinant hepatitis B vaccine in healthy older adults. J Infect Dis. 1997;175(6):1494–97. doi:https://doi.org/10.1086/516485.
- Nisini R, Poerio N, Mariotti S, De Santis F, Fraziano M. The multirole of liposomes in therapy and prevention of infectious diseases. Front Immunol. 2018;9:155.
- van Doremalen N, Lambe T, Spencer A, Belij-Rammerstorfer S, Purushotham JN, Port JR, Avanzato V, Bushmaker T, Flaxman A, Ulaszewska M, et al. ChAdOx1 nCoV-19 vaccination prevents SARS-CoV-2 pneumonia in rhesus macaques. bioRxiv. 2020 May 13. 093195. doi:https://doi.org/10.1101/2020.05.13.093195.
- Aldhamen YA, Seregin SS, Amalfitano A. Immune recognition of gene transfer vectors: focus on adenovirus as a paradigm. Front Immunol. 2011;2:40. doi:https://doi.org/10.3389/fimmu.2011.00040.
- Steinmetz NF, Lim S, Sainsbury F. Protein cages and virus-like particles: from fundamental insight to biomimetic therapeutics. Biomater Sci. 2020;8(10):2771–77. doi:https://doi.org/10.1039/D0BM00159G.
- Zha L, Zhao H, Mohsen MO, Hong L, Zhou Y, Yao C, Guo L, Li Z, Chen H, Liu X, et al. Development of a COVID-19 vaccine based on the receptor binding domain displayed on virus-like particles. bioRxiv. 2020 May 06. 079830. doi:https://doi.org/10.1101/2020.05.06.079830.
- Bachmann MF, Rohrer UH, Kundig TM, Burki K, Hengartner H, Zinkernagel RM. The influence of antigen organization on B cell responsiveness. Science. 1993;262(5138):1448–51. doi:https://doi.org/10.1126/science.8248784.
- Raghunandan R. Virus-like particles: innate immune stimulators. Expert Rev Vaccines. 2011;10(4):409–11. doi:https://doi.org/10.1586/erv.11.37.
- Fu C, Jiang A. Dendritic cells and CD8 T cell immunity in tumor microenvironment. Front Immunol. 2018;9:3059. doi:https://doi.org/10.3389/fimmu.2018.03059.
- Keller SA, Bauer M, Manolova V, Muntwiler S, Saudan P, Bachmann MF. Cutting edge: limited specialization of dendritic cell subsets for MHC class II-associated presentation of viral particles. J Immunol. 2010;184(1):26–29. doi:https://doi.org/10.4049/jimmunol.0901540.
- Rynda-Apple A, Patterson DP, Douglas T. Virus-like particles as antigenic nanomaterials for inducing protective immune responses in the lung. Nanomedicine. 2014;9(12):1857–68. doi:https://doi.org/10.2217/nnm.14.107.
- Wiley JA, Richert LE, Swain SD, Harmsen A, Barnard DL, Randall TD, Jutila M, Douglas T, Broomell C, Young M, et al. Inducible bronchus-associated lymphoid tissue elicited by a protein cage nanoparticle enhances protection in mice against diverse respiratory viruses. PLoS One. 2009;4(9):e7142. doi:https://doi.org/10.1371/journal.pone.0007142.
- Rynda-Apple A, Dobrinen E, McAlpine M, Read A, Harmsen A, Richert LE, Calverley M, Pallister K, Voyich J, Wiley JA, et al. Virus-like particle-induced protection against MRSA pneumonia is dependent on IL-13 and enhancement of phagocyte function. Am J Pathol. 2012;181(1):196–210. doi:https://doi.org/10.1016/j.ajpath.2012.03.018.
- Patel C, Brotherton JML, Pillsbury A, Jayasinghe S, Donovan B, Macartney K, Marshall H. The impact of 10 years of human papillomavirus (HPV) vaccination in Australia: what additional disease burden will a nonavalent vaccine prevent? Eurosurveillance. 2018;23(41):1700737. doi:https://doi.org/10.2807/1560-7917.ES.2018.23.41.1700737.
- Schwarz TF, Huang LM, Valencia A, Panzer F, Chiu CH, Decreux A, Poncelet S, Karkada N, Folschweiller N, Lin L, et al. A ten-year study of immunogenicity and safety of the AS04-HPV-16/18 vaccine in adolescent girls aged 10–14 years. Hum Vaccin Immunother. 2019;15(7–8):1970–79. doi:https://doi.org/10.1080/21645515.2019.1625644.
- Slifka MK, Amanna IJ. Role of multivalency and antigenic threshold in generating protective antibody responses. Front Immunol. 2019;10:956.
- Zur Hausen H. Papillomaviruses and cancer: from basic studies to clinical application. Nat Rev Cancer. 2002;2(5):342–50. doi:https://doi.org/10.1038/nrc798.
- Hung CF, Ma B, Monie A, Tsen SW, Wu TC. Therapeutic human papillomavirus vaccines: current clinical trials and future directions. Expert Opin Biol Ther. 2008;8:421–39.
- Dashti NH, Abidin RS, Sainsbury F. Programmable in vitro coencapsidation of guest proteins for intracellular delivery by virus-like particles. ACS Nano. 2018;12(5):4615–23. doi:https://doi.org/10.1021/acsnano.8b01059.
- Myoung J, Lee SA, Lee HR. Beyond viral interferon regulatory factors: immune evasion strategies. J Microbiol Biotechnol. 2019;29:1873–81.
- Chen S, Yang C, Zhang W, Mahalingam S, Wang M, Cheng A. Flaviviridae virus nonstructural proteins 5 and 5A mediate viral immune evasion and are promising targets in drug development. Pharmacol Ther. 2018;190:1–14. doi:https://doi.org/10.1016/j.pharmthera.2018.05.004.
- Park A, Iwasaki A. Type I and Type III interferons - induction, signaling, evasion, and application to combat COVID-19. Cell Host Microbe. 2020;27(6):870–78. doi:https://doi.org/10.1016/j.chom.2020.05.008.
- Menachery VD, Yount BL, Debbink K, Agnihothram S, Gralinski LE, Plante JA, Graham RL, Scobey T, Ge X-Y, Donaldson EF, et al. A SARS-like cluster of circulating bat coronaviruses shows potential for human emergence. Nat Med. 2015;21:1508–13. doi:https://doi.org/10.1038/nm.3985.
- Siu KL, Kok KH, Ng MH, Poon VK, Yuen KY, Zheng BJ, Jin D-Y. Severe acute respiratory syndrome coronavirus M protein inhibits type I interferon production by impeding the formation of TRAF3.TANK.TBK1/IKKepsilon complex. J Biol Chem. 2009;284(24):16202–09. doi:https://doi.org/10.1074/jbc.M109.008227.
- Yang Y, Zhang L, Geng H, Deng Y, Huang B, Guo Y, Zhao Z, Tan W. The structural and accessory proteins M, ORF 4a, ORF 4b, and ORF 5 of Middle East respiratory syndrome coronavirus (MERS-CoV) are potent interferon antagonists. Protein Cell. 2013;4(12):951–61. doi:https://doi.org/10.1007/s13238-013-3096-8.
- Kopecky-Bromberg SA, Martinez-Sobrido L, Frieman M, Baric RA, Palese P. Severe acute respiratory syndrome coronavirus open reading frame (ORF) 3b, ORF 6, and nucleocapsid proteins function as interferon antagonists. J Virol. 2007;81(2):548–57. doi:https://doi.org/10.1128/JVI.01782-06.
- Lokugamage KG, Hage A, Schindewolf C, Rajsbaum R, Menachery VD. SARS-CoV-2 is sensitive to type I interferon pretreatment. BioRxiv. 2020 March 07.982264. doi:https://doi.org/10.1101/2020.03.07.982264.
- Thoms M, Buschauer R, Ameismeier M, Koepke L, Denk T, Hirschenberger M, Kratzat H, Hayn M, Mackens-Kiani T, Cheng J, et al. Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2. Science. 2020;369(6508):1249–55. doi:https://doi.org/10.1126/science.abc8665.
- Konno Y, Kimura I, Uriu K, Fukushi M, Irie T, Koyanagi Y, Nakagawa S, Sato K. SARS-CoV-2 ORF3b is a potent interferon antagonist whose activity is further increased by a naturally occurring elongation variant. bioRxiv. 2020 May 11.088179. doi: https://doi.org/10.1101/2020.05.11.088179.
- Antia A, Ahmed H, Handel A, Carlson NE, Amanna IJ, Antia R, Slifka M. Heterogeneity and longevity of antibody memory to viruses and vaccines. PLoS Biol. 2018;16(8):e2006601. doi:https://doi.org/10.1371/journal.pbio.2006601.
- Mayer A, Zhang Y, Perelson AS, Wingreen NS. Regulation of T cell expansion by antigen presentation dynamics. Proc Natl Acad Sci U S A. 2019;116(13):5914–19. doi:https://doi.org/10.1073/pnas.1812800116.
- Zinkernagel RM. Immunological memory not equal protective immunity. Cell Mol Life Sci: CMLS. 2012;69(10):1635–40. doi:https://doi.org/10.1007/s00018-012-0972-y.
- Hoffmann M, Kleine-Weber H, Schroeder S, Kruger N, Herrler T, Erichsen S, Schiergens TS, Herrler G, Wu N-H, Nitsche A, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 2020;181(2):271–80e8. doi:https://doi.org/10.1016/j.cell.2020.02.052.
- Letko M, Marzi A, Munster V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat Microbiol. 2020;5(4):562–69. doi:https://doi.org/10.1038/s41564-020-0688-y.
- Pinto D, Park YJ, Beltramello M, Walls AC, Tortorici MA, Bianchi S, Jaconi S, Culap K, Zatta F, De Marco A, et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature. 2020;583(7815):290–95. doi:https://doi.org/10.1038/s41586-020-2349-y.
- Wang X, Xu W, Hu G, Xia S, Sun Z, Liu Z, Xie Y, Zhang R, Jiang S, Lu L, et al. Retraction Note to: SARS-CoV-2 infects T lymphocytes through its spike protein-mediated membrane fusion. Cell Mol Immunol. 2020;17(8):894–894. doi:https://doi.org/10.1038/s41423-020-0498-4.
- Zhou B, She J, Wang Y, Ma X. The duration of viral shedding of discharged patients with severe COVID-19. Clin Infect Dis. 2020. doi:https://doi.org/10.1093/cid/ciaa451.
- Grifoni A, Weiskopf D, Ramirez SI, Mateus J, Dan JM, Moderbacher CR, Rawlings SA, Sutherland A, Premkumar L, Jadi RS, et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell. 2020;181(7):1489–1501.e15.
- Braun J, Loyal L, Frentsch M, Wendisch D, Georg P, Kurth F, Hippenstiel S, Dingeldey M, Kruse B, Fauchere F, et al. Presence of SARS-CoV-2 reactive T cells in COVID-19 patients and healthy donors. medRxiv. 2020 Apr 17. 20061440. doi:https://doi.org/10.1101/2020.04.17.20061440.
- Alexander J, Bilsel P, Del Guercio MF, Marinkovic-Petrovic A, Southwood S, Stewart S, Ishioka G, Kotturi MF, Botten J, Sidney J, et al. Identification of broad binding class I HLA supertype epitopes to provide universal coverage of influenza A virus. Hum Immunol. 2010;71(5):468–74. doi:https://doi.org/10.1016/j.humimm.2010.02.014.
- Masters PS. The molecular biology of coronaviruses. Adv Virus Res. 2006;66:193–292.
- Zhao Z, Li H, Wu X, Zhong Y, Zhang K, Zhang YP, Boerwinkle E, Fu YX. Moderate mutation rate in the SARS coronavirus genome and its implications. BMC Evol Biol. 2004; 28(4):21. doi:https://doi.org/10.1186/1471-2148-4-21.
- Menachery VD, Yount BL Jr., Josset L, Gralinski LE, Scobey T, Agnihothram S, Katze MG, Baric RS. Attenuation and restoration of severe acute respiratory syndrome coronavirus mutant lacking 2ʹ-o-methyltransferase activity. J Virol. 2014;88(8):4251–64. doi:https://doi.org/10.1128/JVI.03571-13.
- Elbahesh H, Saletti G, Gerlach T, Rimmelzwaan GF. Broadly protective influenza vaccines: design and production platforms. Curr Opin Virol. 2019;34:1–9. doi:https://doi.org/10.1016/j.coviro.2018.11.005.
- Vemula SV, Sayedahmed EE, Sambhara S, Mittal SK. Vaccine approaches conferring cross-protection against influenza viruses. Expert Rev Vaccines. 2017;16(11):1141–54. doi:https://doi.org/10.1080/14760584.2017.1379396.
- Milian E, Kamen AA. Current and emerging cell culture manufacturing technologies for influenza vaccines. Biomed Res Int. 2015;2015:504831. doi:https://doi.org/10.1155/2015/504831.
- Soema PC, Kompier R, Amorij JP, Kersten GF. Current and next generation influenza vaccines: formulation and production strategies. Eur J Pharm Biopharm. 2015;94:251–63. doi:https://doi.org/10.1016/j.ejpb.2015.05.023.
- Brown LE, Kelso A. Prospects for an influenza vaccine that induces cross‐protective cytotoxic T lymphocytes. Immunol Cell Biol. 2009;87(4):300–08. doi:https://doi.org/10.1038/icb.2009.16.
- Grifoni A, Weiskopf D, Ramirez SI, Mateus J, Dan JM, Moderbacher CR, Rawlings SA, Sutherland A, Premkumar L, Jadi RS, et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell. 2020;181(7):1489–501e15.
- Gilbert SC. T-cell-inducing vaccines - what’s the future. Immunology. 2012;135(1):19–26. doi:https://doi.org/10.1111/j.1365-2567.2011.03517.x.
- Braun J, Loyal L, Fnotesrentsch M, Wendisch D, Georg P, Kurth F, Hippenstiel S, Dingeldey M, Kruse B, Fauchere F, et al. SARS-CoV-2-reactive T cells in healthy donors and patients with COVID-19. Nature. 2020. doi:https://doi.org/10.1038/s41586-020-2598-9.
- Yuan J, Kou S, Liang Y, Zeng J, Pan Y, Liu L. PCR assays turned positive in 25 discharged COVID-19 patients. Clin Infect Dis. 2020. doi:https://doi.org/10.1093/cid/ciaa398.
- Nakayama T. Causal relationship between immunological responses and adverse reactions following vaccination. Vaccine. 2019;37(2):366–71. doi:https://doi.org/10.1016/j.vaccine.2018.11.045.
- Chen J. Pathogenicity and transmissibility of 2019-nCoV-A quick overview and comparison with other emerging viruses. Microbes Infect. 2020;22(2):69–71. doi:https://doi.org/10.1016/j.micinf.2020.01.004.
- Metcalf CJE, Ferrari M, Graham AL, Grenfell BT. Understanding herd immunity. Trends Immunol. 2015;36(12):753–55. doi:https://doi.org/10.1016/j.it.2015.10.004.
- Rodpothong P, Auewarakul P. Viral evolution and transmission effectiveness. World J Virol. 2012;1(5):131–34. doi:https://doi.org/10.5501/wjv.v1.i5.131.
