ABSTRACT
The Andean tree genus Polylepis (Rosaceae) has recently been recognized to include polyploid species, but their occurrence within the genus is still incompletely known, especially in light of a forthcoming taxonomic treatment based on a narrow species concept including morphological, climatic and biogeographic distinctness that recognizes 45 species. We obtained guard cell measurements as proxies of ploidy level from 114 individuals of 33 species of Polylepis, including all species for which no previous measurements were available. In combination with previously published data, also on nucleus mass and chromosome counts, we infer that on current knowledge 19 (42%) species are probably purely diploid, 15 (33%) purely tetraploid, and one (2%) purely octoploid. The remaining eight (18%) species have mixed ploidy levels, with three (7%) being di- and tetraploid, two (4%) di- and hexaploid, and one each tetra- and hexaploid, tetra- and octoploid, and di-, tri-, tetra- and hexaploid. Based on our understanding of the evolutionary relationships in Polylepis, it would appear that polyploidy has originated at least about eight times independently in the genus, sometimes as autopolyploidy, sometimes as a result of interspecific hybridization, and sometimes in relation to cultivation. The taxonomic implications of the ploidy levels are complex, in some cases supporting species-level distinction and in others posing the question whether different ploidy levels within a species should better be treated as distinct species. Ploidy level needs to be taken into account for the conservation of the genus, as for example if different populations of a species have different ploidy levels, mixing these origins in reforestation schemes may lead to the formation of sterile hybrids. Guard cell measurement is a low cost and simple technique that can be readily used on both live and dried plant material for such applications, but it has limitations and further data on chromosome counts and nucleus mass are also needed to fully understand the evolution of ploidy levels in Polylepis and its implications.
KEYWORDS:
Introduction
Changes in ploidy level are an important process in plant evolution, impacting both the diversification of major clades as well as microevolutionary processes at the species level [Citation1,Citation2]. In many plant groups, one can find different ploidy levels, resulting from either auto- or allopolyploidization, the latter linked to hybridization, where it allows for the stabilization of genomes of mixed origin [Citation3–5]. Besides its evolutionary implications [Citation6], polyploidy is also of taxonomic relevance, since populations of different ploidy levels are at least partly reproductively isolated, allowing for divergent evolutionary trajectories that can be treated at species level [Citation7].
The genus Polylepis (Rosaceae) is a taxonomically complex genus that has variously been treated as comprising 33 species [Citation8], 15 species [Citation9], 26 species [Citation10], or 45 species [Citation11]. This variability already hints at the difficulty of delimiting species in this genus, which is due to overall morphological similarity between species coupled with high intraspecific variability and plasticity, which themselves are linked to hybridization and polyploidization [Citation12]. The family Rosaceae, to which Polylepis belongs, is renowned for the occurrence of polyploid complexes, e.g., in Crataegus [Citation13] and Sorbus [Citation14,Citation15].
In Polylepis, polyploidization has long been hypothesized to occur [Citation9,Citation12,Citation16], but direct chromosome counts in Polylepis have been hampered by their small size and the difficulty of preparation [Citation9,Citation17–19]. Different ploidy level in Polylepis were first documented by Schmidt-Lebuhn et al. [Citation19] who made flow cytometry measurements of 11 species, finding that the majority of species have nucleus masses of around 1.5–1.65 pg, which they interpreted as diploids, whereas some species had values around 2.9–3.1 (tetraploids), with a single species having a mean value of 5.7 (octoploid). Because flow cytometry in Polylepis requires live material (all efforts to obtain measures from dried material have failed so far, even when working in collaboration with experts such as J. Suda) and not all species were available in cultivation, they additionally used guard cell measurements of herbarium specimens to estimate ploidy levels of additional species. Guard cell size is well known to be correlated to ploidy level in angiosperms in general [Citation5,Citation20,Citation21] and also Rosaceae in [Citation22], although in some genera such as Crataegus guard cell sizes overlap between ploidy levels, so that ploidy inference is challenging [Citation23]. In Polylepis, by sampling the same individuals as used for flow cytometry, Schmidt-Lebuhn et al. [Citation19] confirmed the applicability of guard cell length for separating at diploid and polyploid individuals, even though the differentiation between tetra- and octoploids was not possible with their limited sample size. Based on their combined flow cytometry and guard cell size data, they arrived at a phylogenetic hypothesis for the development of ploidy level in Polylepis, stating that the ancestral diploid condition is found in the phylogenetically older sections Sericea and Reticulata (as defined by [Citation11]), whereas the more derived section Incana is largely polyploid. The latter section on average occurs at higher elevations and in more arid environments, which corresponds well with the polyploid condition since polyploids are well known to be overrepresented at high latitudes and elevations [Citation5,Citation24], possibly because the different paralogs offer higher adaptive potential [Citation25]. A major limitation of this study was that each species was only sampled with 2–4 individuals, so that intraspecific variability may have gone undetected. The only species for which several ploidy levels were detected were P. australis and P. pauta.
In a later study, Kessler et al. [Citation26] conducted more detailed flow cytometry measurements of three Argentinian species of Polylepis, finding constant ploidy levels in P. hieronymi (diploid, N = 52 individuals from three sites) and P. tomentella (tetraploid, N = 43, three sites) but marked variation in P. australis. In this species, among 361 individuals from 27 populations, 75 (21%) were interpreted as diploids, 24 (7%) as triploids, 261 (72%) as tetraploids, and one (0.003%) as a hexaploid. These ploidy levels showed a clear geographical pattern, with populations from the northern Argentinean Andes being purely diploid and those from the central Andes tetraploid, whereas in the isolated Sierra de Córdoba, all four ploidy levels co-occurred in mixed populations. This suggests that the triploid plants may be hybrids between the di- and tetraploid ones, but whether they are sterile primary hybrids or can reproduce by themselves is unresolved. Also, the degree of reproductive isolation and hence evolutionary independence between the diploid and tetraploid populations remains unknown.
The next studies examining ploidy levels in Polylepis were conducted by Segovia-Salgado and colleagues on Ecuadorean species [Citation27–33]. Using flow cytometric measures of nucleus mass, chromosome numbers, and guard cell lengths, of dozens of individuals of each species, they documented that many species have variable chromosome numbers and nucleus mass, suggesting reductions in chromosome numbers. These reductions in chromosome numbers and DNA content could be the result of aneuploidy (loss of DNA and reduction of chromosome size) and dysploidy (chromosome fusion) [Citation34–36].
Currently, a new taxonomic treatment of Polylepis is being prepared by TB and MK which applies a narrower species concept than previous treatments, resulting in 45 species recognized [Citation11,Citation37–39]. This species concept combined information on morphology, distribution, and ecology to infer evolutionarily independent units that are treated as species. For example, applying this concept to the previously very widespread and morphologically variable P. sericea as defined by Simpson [Citation9], has resulted in the recognition of seven species, which partly occur in sympatry without hybridization, clearly supporting their treatment as independent species [38, 40. Because ploidy level can also yield important taxonomic information, in the present study we set out to estimate ploidy levels of all species based on guard cell measurements. We used this methodological approach because it can be applied to herbarium material and because accessing live plants of so many species would have been impossible. Also, guard cell measurement is the only possible approach to assess ploidy level in type specimens.
Our study also aims to provide a framework to understand the implications of polyploidy of Polylepis, and in particular regarding intraspecific ploidy variation, in strategies of conservation and restoration for the genus. This is particularly important in species reintroduction or ecological restoration, because there are potentially negative consequences when ploidy variants are unintentionally mixed within populations, such as the formation of sterile triploid individuals that may reduce overall population fitness.
Methods
Sampling
To measure guard cell length, we selected 1–24 herbarium specimens of 33 species from the herbaria F, GOET, MO, NY, QCA, and Z/ZH, depending on the availability of previous measurements (previously well-studied species were not sampled). Species level taxonomy follows Boza & Kessler [Citation11].
Guard cell measurements
Guard cell size measurements followed the approach of Kessler et al. [Citation26], with some modifications. Three leaflets per specimen were used for measurements. On each leaflet, we selected a central part halfway between the midvein and the margin to measure the guard cells. In this area, we first carefully removed the hair and wax cover with a scalpel or brush. In species with a wax layer on the leaflet surface, we soaked the leaflets for 1–4 h in acetone or isopropanol, then rinsed them in clear water, and finally dried samples between tissue paper. Once all covering substances had been eliminated from the leaves, we applied clear nail varnish that was allowed to dry for 4–12 h before removal. Application of the varnish was complicated by the fact that the guard cells of many Polylepis species are small and deeply sunken in stomatal pits, so that we used different dilution levels of the varnish with acetone. Often, several layers of varnish were applied to obtain a cover that would not change shape when pulling it off. In species in which the nail varnish level did not work, we used a second method: we mixed ethanol (80%) and hydrogen peroxide (30%) in a relation 1 to 2. Leaflets were left in this solution for up to 3 days until bleached and bloated, with the epidermis separating from the parenchyma. The samples were then rinsed with clear water and stained with safranin. Once the samples were considered adequate, we measured the guard cell length on 10–30 stomata from each sample at 400X magnification under a light microscope and averaged these measurements. For some species, obtaining measurements was difficult, especially when the leaves were thick, covered by a dense hair layer, and when the stomata were deeply sunken in pits. In such cases, repeated attempts with different methods were necessary. Nevertheless, by combining different preparation methods, we obtained at least one measurement for each species.
Ploidy level assignment
Inferring ploidy levels based on guard cell measurements faces several challenges. First there is the issue of assigning the correct base chromosome number. Segovia-Salcedo [Citation31] used the base haploid chromosome number of n = 7 in the family Rosaceae [Citation40] as reference, thus interpreting a chromosome count of 42 as hexaploid (x = 6). Schmidt-Lebuhn et al. [Citation19] and Kessler et al. [Citation26] instead used the lowest number in the genus (2 n = 42) as baseline, interpreting this as a diploid (x = 2). Polylepis has been hypothesized to have originated from the homoploid hybridization between two species of Acaena with n = 21 each [Citation31,Citation41,Citation42], which might be taken as indication of a chromosome base number of n = 21 in the genus. However, we consider that ultimately, within a plant group, the crucial factor is the meiotic pairing behavior of the chromosomes, i.e., whether they behave as bivalents so that during chromosome pairing each chromosome has a single counterpart, or as polyvalents where they can pair with several other chromosomes. This behavior is unknown for Polylepis or related genera. For simplicity, we here use a chromosome number of 2 n = 42 as baseline for defining diploids, while acknowledging that a baseline of x = 7 may ultimately prove to be more appropriate.
The second issue in inferring ploidy levels concerns variation in measurements and conflicts between different data sources. For inferring ploidy from flow cytometry measurements of nucleus mass, we considered that the diploid condition is related to values around 1.4–1.7 pg, triploidy to 2.0–2.3, tetraploidy to 2.6–3.4 pg, hexaploidy with 4.6–4.9 pg, and octoploidy to 5.7–5.8 pg [Citation19,Citation26]. Following the assignement of a base chromosome number of x = 21, chromosome counts of around 42 correspond to diploids, around 84 to tetraploids, and around 126 to hexaploids. However, numerous published counts differ notably from these values. For example, Caiza et al. [Citation33] and Segovia-Salcedo & Quijia [Citation32] reported chromosome counts of 59–77 for nine individuals of P. ochreata (as P. sericea) from Yanacocha, Ecuador. In this situation, it is unclear if these numbers reflect the difficulty of fully counting the tiny chromosomes, or whether they correspond to real values with would suggest triploidy and other intermediate chromosome levels as a result of aneuploidy and dysploidy.
Regarding guard cell measurements, there is notable variation within species, even when only a single ploidy level is believed to occur in the species [Citation19]. This variation may be due to anatomical plasticity of a species depending on growth conditions as also found in Crataegus [Citation23], differences in measurement methods, or aneuploidy. Thus, to assess the suitability of guard cell measurements in inferring ploidy levels, we plotted the frequencies of measurements in size classes of 1 µm (), and indicated those measurements that come from species for which available flow cytometry or chromosome counts have so far only indicated a single ploidy level. Overall, we found that the guard cell measurements showed a bimodal distribution, with one peak at around 13 µm, and the other at around 17 µm, with perhaps a third minor peak at 22 µm. All measurements that can be linked with known diploid species show guard cell measurements below 15 µm, whereas the majority of measurements that can be linked with known tetraploid species correspond to guard cell measurements of 15–20 µm. However, three measurements associated with tetraploid species had measurements of 13–14 µm. Two of these measurements come from P. triacontandra, which has a single chromosome count of around 80 (tetraploid) [Citation19] and three guard cell measurements of 18.0–20.4, which is consistent with tetraploidy, but where we also have two measurements of 13.5 µm and 14.1 µm. Polylepis triacontandra is morphologically variable and occurs in a region with a long history of human impact, where species of Polylepis have been transplanted for long times [Citation10], and where hybridization with other species cannot be ruled out, so that we cannot say with any certainty whether these low measurements truly correspond to tetraploid individuals or whether they might not actually be diploid or triploid plants. The other tetraploid species with a low guard cell measurement is P. nana, with two flow cytometry measurements of 2.93–2.96 pg (tetraploid) and five corresponding guard cells measurements between 18.6 µm and 20.3 µm (tetraploid), but also two measurements of 13.1 µm and 15.4 µm. Here, again, it is clear that the species has tetraploid individuals, but it is unclear whether the low measurements are from individuals that are tetraploid or diploid. In addition, we have two measurements of a species known to be octoploid based on flow cytometry (undescribed species P. sacra), where the same individuals used for flow cytometry had guard cells sizes of 16.7 µm and 20.2 µm, respectively, thus overlapping with measurements of known tetraploid plants of other species.
Figure 1. Number of Polylepis guard cell measurements in size classes (μm, with 9 indicating measurements from 9.0 to 9.9 μm), including all previously published and newly generated measurements. Total height of each bar shows the total number of measurements in this size class, with white indicating measurements that cannot be directly linked with chromosome counts or flow cytometry measurements, light gray the number of samples from species with only diploid chromosome counts or flow cytometry measurements between 1.4 and1.7 pg, dark gray species with tetraploid chromosome counts or flow cytometry measurements between 2.6 and 3.4 pg, and black species with octoploid flow cytometry measurements
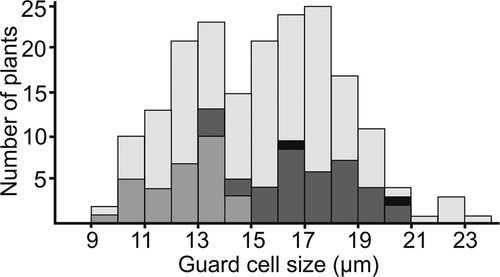
From these data we can conclude several things. First, the bimodal distribution of guard cell measurements and the strong association of the two peaks with known diploid and tetraploid ploidy levels confirmed by flow cytometry or chromosome counts support the notion that guard cell measurements reflect ploidy levels in Polylepis. Second, however, the overlap between the two peaks and the overlap of the individuals that can be associated with known ploidy levels shows that unambiguous assignment of ploidy levels is not possible for each single guard cell measurement. We conclude that guard cell measurements below 13 µm are very likely to correspond to diploid plants and that measurements above 15 µm are equally likely to be tetraploids or higher ploidy level plants. Values between 13.0 µm and 15.0 µm may correspond to both diploid or tetraploid individuals. Finally, plants with guard cell measurements above 21 may well correspond to hexa- or octoploids, but our data is insufficient to confirm this.
Based on these considerations and in combining all available data sources, our inferences of ploidy levels in species of Polylepis were based on the majority of the data available. For instance, in the case of P. nana, 5 of 6 guard cell measurements are above 15 µm, indicating tetraploidy, and flow cytometry also points to tetraploidy, so that we interpret this species as being tetraploid. The single guard cell measurement of 13.1 is intriguing in that it might indicate that lower ploidy levels also occur in this species, but not given much weight until further data are available. In , we indicate our assessment of the confidence we have in the ploidy inferences based on the amount and quality of data, and concordance between different data sources, in three steps, ranging from “low confidence” (*) to “high confidence” (***).
Table 1. Overview of the available data on DNA content, chromosome numbers, and guard cell sizes in species of the genus Polylepis. Species taxonomy follows Boza et al. [Citation14] and Boza & Kessler [Citation11]. Where possible, literature records were assigned to this taxonomy, but a few data points (especially from the hybrid zone at Mojanda, Ecuador) had to be excluded because they could not be unambiguously assigned to a species. Species arrangement in sections and subsections follows Boza & Kessler [Citation11]. Data from the literature are in italics, newly generated data in bold. Depending on data source, we report the mean ± standard deviation, only the mean, or a range. Our confidence in the inferences of ploidy levels, based on the amount and quality of data, and concordance of data from different sources, is indicated by asterisks ranging from low (*) to high (***). Data sources: 1: Schmidt-Lebuhn et al [Citation19].; 2: Quijia et al. [Citation26]; 3: Montalvo [Citation28]; 4: Zurita et al. [Citation29]; 5: Segovia-Salcedo & Quijia [Citation31]; 6: Segovia-Salcedo [Citation30]; 7: Kessler et al. [Citation25]; 8: Caiza et al. [Citation32]; 9: this study. Abbreviations: Ar = Argentina, Ec = Ecuador, cult. = cultivated (in botanical garden), ind. = individuals, pl = planted
Results and discussion
We obtained guard cell measurements from 114 individuals of 33 species of Polylepis, including all species for which no previous measurements were available ().
For those species for which previous data were available, our new measurements were largely congruent. For example, for P. argentea Kessler et al. [Citation26] reported sizes from two living plants as 12.4 µm and 13.0 µm, which is in the diploid range, and our measurements from herbarium specimens amounted to 13.4–13.9 µm which is within the mixed diploid/tetraploid range. In this case, we interpreted the species to be diploid, with an intermediate level of confidence. In the case of P. tarapacana, we measured the same collection Kessler 3599 that was also measured by Schmidt-Lebuhn et al. [Citation19]. Although the measurements were not identical, the standard deviations overlapped broadly (17.4 ± 2.2 and 19.7 ± 2.0), confirming the replicability of the measurements while also showing some study-dependent variation which may be due to the preparation and measurement methods.
Where all three types of data are available for a species, they are often congruent and point to a single ploidy level, as in P. lanuginosa, undescribed species P. simpsonii, and P. tarapacana (). In other cases, the data are complementary, as in P. ochreata, where the two flow cytometry measurements point to tetra- and hexaploidy, the chromosome counts to di- and tetraploidy (and perhaps intermediate values as discussed under ploidy level assignment), and the guard cell measurements to diploidy. In this situation, we infer that the species includes di-, tetra-, and hexaploid individuals, possibly with aneuploids or hybrids. None of the three data sources would have by itself provided this picture, pointing to the importance of obtaining complementary data and highlighting the limitations of single types of data. At present, for the 45 species of Polylepis, we have combined data on guard cell length, chromosome number, and genome size for 9 (20%) species, on guard cell length and genome size for another 9 (20%) species, and on guard cell length and chromosome number for 3 (7%) species, whereas for 24 (53%) species only guard cell measurements are available.
Bearing in mind the potential limitations of incomplete data and some uncertainty in the interpretation of the data, we infer that at present knowledge, 19 (42%) species appear to be purely diploid, 15 (33%) purely tetraploid, and one (2%) purely octoploid. The remaining 8 (18%) species have mixed ploidy levels, with 3 (7%) being di- and tetraploid, 2 (4%) di- and hexaploid, 1 (2%) tetra- and hexaploid, 1 (2%) tetra- and octoploid, and 1 (2%) di-, tri-, tetra- and hexaploid. While it is likely that further studies will reveal more cases of mixed ploidy, at least some well-studied species appear to consistently show diploid (P. hieronymi, P. lanuginosa, P. simpsonii) or tetraploid (P. tarapacana, P. tomentella) conditions, suggesting that not all species have mixed ploidy levels.
From an evolutionary point of view, Schmidt-Lebuhn et al. [Citation19] proposed that the a chromosome number of 2 n = 42 is ancestral in Polylepis, and that higher ploidy levels evolved once at the base of the Incana complex as defined by Simpson [Citation9] and twice within species outside of this section (P. australis, P. pauta). The data that has since then been compiled in this and other studies points to a more complex picture (). In section Sericea, which has been considered to include a grade of the phylogenetically basal members of the genus [Citation9,Citation12,Citation16], most species are diploid (taking n = 21 as the base number), but polyploidy is present in P. ochreata and P. pauta. These two species overlap in northern Ecuador where they hybridize extensively [Citation11,Citation38]. It is conceivable that the polyploid condition in these two species stems from this hybridization, as polyploidy is often correlated with hybridization [Citation3,Citation4]. In the presumably monophyletic section Reticulata, again most species are diploid, but polyploidy occurs in two species that appear not to be closely related (P. microphylla, P. reticulata). Interestingly, at least in P. reticulata, polyploidy is so far only known from cultivated plants [Citation31], suggesting that this condition may be related to domestication. Polyploidization as a result of domestication is a common phenomenon in plants [Citation43]. In section Australis, P. australis has been well studied and includes di-, tri-, tetra- and hexaploids, most likely due to autopolyploidization [Citation26]. Section Subsericans includes three purely tetraploid species. Finally, section Incana mainly includes tetraploid species, but diploidy occurs in P. incana and the closely related undescribed species P. fjeldsaai of subsection Incana. Polylepis incana has been considered to be one of the most derived species in this section [Citation9,Citation12,Citation16], so that a diploid condition is surprising if one assumes that P. incana is nested within a tetraploid clade. This suggests that the assumption that P. incana is a derived member of this section is wrong, and that the evolution of this section is more complex than previously assumed. In any case, assuming that the species sections recognized here based on morphological and ecological similarity are evolutionary units, we now can deduce that polyploidy evolved at least eight times in the genus, possibly more often. However, considering that the evolution of Polylepis is probably reticulate and that the phylogeny of the genus remains poorly understood [Citation12,Citation42], inferring the origins of polyploidy in Polylepis is very difficult, and the conclusions drawn here should be viewed as hypotheses to be tested by future studies.
Figure 2. Simplified phylogenetic hypothesis of Polylepis based on Schmidt-Lebuhn et al. [Citation19] indicating the ploidy levels as inferred in this study (diploid = light gray, tetraploid = dark gray, hexa- to octoploid = black) and the presumed origins of polyploidy (*). Triangles are proportional to species numbers in each section or subsection; see for species. Names of subsections (left) and sections (right) are indicated next to the triangles; Austr. = Australis, Lan. = Lanuginosa, Subser. = Subsericans. Photos show representative species of the sections and subsections. From top to bottom: Polylepis rugulosa (E. Urquiaga), P. racemosa (T. Boza), P. incana (E. Urquiaga), P. subsericans (T. Boza), P. australis (A. Bernhard), P. hieronymi (A. Bernhard), P. rodolfo-vasquezii (T. Boza), P. humboldtii (T. Boza), P. pauta (T. Boza), and P. lanuginosa (T. Boza)
![Figure 2. Simplified phylogenetic hypothesis of Polylepis based on Schmidt-Lebuhn et al. [Citation19] indicating the ploidy levels as inferred in this study (diploid = light gray, tetraploid = dark gray, hexa- to octoploid = black) and the presumed origins of polyploidy (*). Triangles are proportional to species numbers in each section or subsection; see Table 1 for species. Names of subsections (left) and sections (right) are indicated next to the triangles; Austr. = Australis, Lan. = Lanuginosa, Subser. = Subsericans. Photos show representative species of the sections and subsections. From top to bottom: Polylepis rugulosa (E. Urquiaga), P. racemosa (T. Boza), P. incana (E. Urquiaga), P. subsericans (T. Boza), P. australis (A. Bernhard), P. hieronymi (A. Bernhard), P. rodolfo-vasquezii (T. Boza), P. humboldtii (T. Boza), P. pauta (T. Boza), and P. lanuginosa (T. Boza)](/cms/asset/930b3363-f6ed-4212-b25a-cb78bdd35ce4/tneo_a_1844992_f0002_oc.jpg)
Placing Polylepis in the broader context of the family Rosaceae and particularly the tribe Sanguisorbae to which it belongs, shows that it higher chromosome numbers than many other genera of the tribe (e.g., 14–56 in Sanguisorba, 28 in Bencomia, and 28–56 in Agrimonia [Citation40]). On the other hand, the values in Polylepis are comparable to those of the most closely related genus Acaena, which has reported chromosome counts between 42 and 126. This is consistent with the interpretation of Polylepis having evolved from the homoploid hybridization between two species of Acaena with n = 21 each [Citation31,Citation41]. Whether 2 n = 42 is a functionally polyploid or diploid conditions in these genera remains unknown.
Finally, focusing on the taxonomic implications of ploidy levels in the genus, we found that in some cases, closely related species have different ploidy levels, supporting their treatment as distinct species. For example, P. fjeldsaai has previously been identified as part of P. tomentella [Citation44], but whereas the first species is diploid based on our data, the latter is consistently tetraploid. On the other hand, at least eight species include individuals of different ploidy levels. At least in P. australis, this is clearly a natural condition [Citation27], which raises the question as to how to treat the different ploidy levels taxonomically. It has been suggested that different ploidy levels within a “species” should be treated at species level if there is morphological, ecological, or biogeographical evidence that they are evolutionarily largely independent units [Citation7]. This approach has been taken in polyploid-apomict species complexes of other genera of Rosaceae such as Crataegus [Citation45] and Sorbus [Citation14], but more information is needed before this approach can be applied to Polylepis. On the other hand, in several species polyploidization is apparently linked to cultivation, as in P. incana, P. racemosa, and P. reticulata. Polyploidization of cultivated plants is a common phenomenon either via auto- or allopolyploidization where higher ploidy levels are often associated with higher plant vigor and adaptive potential [Citation46–48]. Polylepis has long been planted by Andean inhabitants as a source of building material, firewood, and as fences [Citation17], and it is conceivable that natural or artificial hybrids have been favored.
Conclusions
The novel data on guard cell length presented in this study, in combination with previous data, provide a more comprehensive view of the distribution of ploidy levels within the genus Polylepis. This reveals a complex evolutionary history, with repeated polyploidization events partly linked to hybridization and cultivation. Understanding variation in ploidy has important practical implications for the management and conservation of Polylepis, many species of which are threatened with extinction [Citation49]. For instance, if populations of a species have different ploidy levels, mixing these origins in reforestation schemes may lead to the formation of sterile hybrids, reducing the reproductive potential of the entire population. On the other hand, if different ploidy levels are reproductively isolated, then they represent different evolutionary units that deserve independent conservation attention. Clearly, ploidy level needs to be taken into account for the conservation of the genus. While the assessment of ploidy level using flow cytometry requires expensive technology, and direct chromosome counts are quite challenging [Citation27], the measurement of guard cells is a low cost and simple technique that can be readily applied to both live and dried plant material. However, guard cell measurements are also the least precise of the three methods, and our study only represents a further step in our understanding of ploidy level distribution and evolution in Polylepis.
More detailed studies are clearly needed to clarify many important questions related to polyploidy and its consequences within Polylepis. Some that we consider to be particularly important include:
What is the functional chromosome base number in Polylepis? In species with 42 chromosomes here considered as diploids, do they behave as 21 pairs of bivalents or as six groups of seven chromosomes that can pair in different combinations? Detailed studies of meiotic pairing behavior are needed, as done, e.g., for Crataegus [Citation50].
Are individuals of naturally occurring different ploidy levels with a “species” evolutionarily independent? This might be addressed in species of known multiple ploidy levels by combining ploidy measurements with population genetic approaches. Particularly intriguing target species for this kind of study include P. australis, P. incana, P. racemosa, and P. reticulata.
Are triploid individuals, which are known to occur at least in P. australis and which may occur in other species having both diploid and tetraploid individuals, sterile or can they reproduce via apomixis? In P. australis, seed viability is very low in many individuals [Citation51–53] but it is not known if this related to ploidy.
More generally, does apomixis occur in Polylepis? This is commonly found in other genera of Rosaceae, where it is frequently linked with polyploidization [Citation40]. This can be dome via pollination experiments and flow cytometry of embryonic tissue, as conducted, e.g., for Lachemilla [Citation54].
What is the prevalence of aneuploidy and dysploidy in the genus? Studies in Ecuador [Citation27–33] suggest that these occur in the genus, but data are inconclusive.
How does hybridization between ploidy levels induced by translocation and reforestation activities affect the population viability of species? This is a situation where guard cell measurements may be very helpful, since their variability is likely to be lower within individual species and among individuals growing under similar environmental conditions.
Author contribution
This work is part of the Ph.D. thesis research of T. E. Boza E. and was developed under the supervision of M. Kessler. Both authors consolidated the ploidy database and contributed to any taxonomic decision adopted in this paper. The labwork was done by V. Popp. All authors provided comments on drafts of this manuscript.
Acknowledgments
We thank the curator of the herbaria (AAU, COL, CUZ, F, GOET, HUA, LOJA, MEDEL, MERF, MO, NY, QCA, US, USM, VEN, and Z/ZH) for loans of specimens, M.C. Segovia S. for allowing us to include unpublished data from her PhD thesis, Mario Coiro for sharing the chemical preparation method. For support of the present study, we thank FONDECYTCONCYTEC (No227-2014-FONDECYT) for providing funding to TEBE, and to the Servicio Nacional Forestal y de Fauna Silvestre-Peru, for providing research authorizations under the R.D.G. No 233-2015, No 237-2015-SERFOR/DGGSPFFS, and Ministerio del Ambiente-Ecuador for providing research authorization under MAE-DNB-CM-2018-0082. T. Dickinson and an anonymous reviewer provided critical comments that greatly improved the manuscript.
Disclosure statement
No potential conflict of interest was reported by the authors.
Additional information
Funding
References
- Adams KL, Wendel JF. Polyploidy and genome evolution in plants. Curr Opin Plant Biol. 2005;8(2):135–141. Available from: https://doi.org/https://doi.org/10.1016/j.pbi.2005.01.001
- Jiao Y, Wickett NJ, Ayyampalayam S, et al. Ancestral polyploidy in seed plants and angiosperms. Nature. 2011;473(7345):97. Available from: https://doi.org/https://doi.org/10.1038/nature09916
- de Wet JMJ. Polyploidy and evolution in plants. Taxon. 1971;20(1):29–35. Available from: www.jstor.org/stable/1218531
- Tate JA, Soltis DE, Soltis PS. Polyploidy in plants. In T. Ryan Gregory (Ed.), The evolution of the genome. Academic Press; 2005. p. 371–426. Department of Entomology, The Natural History Museum, London, UK. Available from: https://doi.org/https://doi.org/10.1016/B978-012301463-4/50009-7
- Masterson J. Stomatal size in fossil plants:</L> evidence for polyploidy in majority of angiosperms. Science. 1994;264(5157):421–424. Available from: https://science.sciencemag.org/content/264/5157/421
- Weiss-Schneeweiss H, Emadzade K, Jang TS, et al. Evolutionary consequences constraints and potential of polyploidy in plants. Cytogenet Genome Res. 2013;140(2–4):137–150. Available from: https://doi.org/https://doi.org/10.1159/000351727
- Soltis DE, Soltis PS, Schemske DW, et al. Autopolyploidy in angiosperms: have we grossly underestimated the number of species? Taxon. 2007;56(1):13–30. Available from: https://www.jstor.org/stable/25065732
- Bitter G. Revision der Gattung Polylepis. Bot Jahb Syst. 1911;565–656.
- Simpson BB. A revision of the genus Polylepis (Rosaceae: sanguisorbae). Smithsonian Contrib Bot. 1979;43:1–62. .
- Kessler M, Schmidt-Lebuhn AN. Taxonomical and distributional notes on Polylepis (Rosaceae). Org Divers Evol. 2006;6:67–70. Available from: https://doi.org/https://doi.org/10.1016/j.ode.2005.04.001
- Boza TE, Kessler M. A monograph of the genus Polylepis (Rosaceae). Phytokeys. in prep.
- Schmidt-Lebuhn AN, Kessler M, Kumar M. Promiscuity in the Andes: species relationships <J>in polylepis (Rosaceae, Sanguisorbeae) based on AFLP and morphology. Syst Bot. 2006;31:547–559. Available from: https://doi.org/https://doi.org/10.1043/05-25.1
- EYY L, Stefanovic S, Dickinson TA. Reconstructing reticulation history in a phylogenetic framework and the potential of allopatric speciation in an agamic complex <J>in Crataegus (Rosaceae). Evolution. 2010;64:3593–3608. Available from: https://doi.org/https://doi.org/10.1111/j.1558-5646.2010.01063.x
- Robertson A, Rich TIMOTHYCG, Allen AM, et al. Hybridization and polyploidy as drivers of continuing evolution and speciation in Sorbus. Mol Ecol. 2010;19(8):1675–1690. Available from: https://doi.org/https://doi.org/10.1111/j.1365-294X.2010.04585.x
- Pellecier J, Clermont S, Houston L, et al. Cytotype diversity in the Sorbus complex (Rosaceae) in Britain: sorting out the puzzle. Ann Bot. 2012;110:1185–1193. Available from: https://doi.org/https://doi.org/10.1093/aob/mcs185
- Simpson BB. Speciation and specialization of Polylepis in the Andes. In: Vuilleumier F, Monasterio M, editors. High altitude tropical biogeography. Oxford: Oxford University Press; 1986. p. 304–316.
- Kessler M. The genus Polylepis (Rosaceae) in Bolivia. Candollea. 1995;50:131–171.
- Schmidt-Lebuhn AN, Seltmann P, Kessler M. Consequences of the pollination system on genetic structure and patterns of species distribution in the Andean genus Polylepis (Rosaceae): a comparative study. Plant Syst Evol. 2007;266:91–103. Available from: https://doi.org/https://doi.org/10.1007/s00606-007-0543-0
- Schmidt-Lebuhn AN, Fuchs J, Hertel D, et al. An Andean radiation: polyploidy in the tree genus <J>Polylepis (Rosaceae, Sanguisorbeae). Plant Biol. 2010;12:917–926. Available from: https://doi.org/https://doi.org/10.1111/j.1438-8677.2009.00297.x
- Sugimoto-Shirasu K, Roberts K. “Big it up”: endoreduplication and cell-size control in plants. Curr Opin Plant Biol. 2003;6(6):544–553. Available from: https://doi.org/https://doi.org/10.1016/j.pbi.2003.09.009
- Beaulieu JM, Leitch IJ, Patel S. Genome size is a strong predictor of cell size and stomatal density in angiosperms. New Phytol. 2008;179(4):975–986. Available from: https://doi.org/https://doi.org/10.1111/j.1469-8137.2008.02528.x
- Joly S, Bruneau A. Delimiting species boundaries in Rosa sect. Cinnamomeae (Rosaceae) in eastern North America. Syst Bot. 2007;32(4):819–836. Available from: https://doi.org/https://doi.org/10.1043/06-69.1
- McGoey BV, Chau K, Dickinson TA. Stomata size in relation to ploidy level in North American hawthorns (Crataegus, Rosaceae). Madroño. 2014;61(2):177–193.
- Brochmann C, Brysting AK, Alsos IG, et al. Polyploidy in arctic plants. Biol J Linn Soc. 2004;82:521–536. Available from: https://doi.org/https://doi.org/10.1111/j.1095-8312.2004.00337.x
- Chung S, Elisens W, Skvarla J. Pollen Morphology and its phylogenetic significance in the tribe Sanguisorbeae (Rosaceae). Plant Syst Evol. 2010;285::135–148. Available from: https://doi.org/https://doi.org/10.1007/s00606-009-0262-9
- Kessler M, Kühn A, Solís VG, et al. Complex geographical distribution of ploidy levels in Polylepis australis (Rosaceae), an endemic tree line species in Argentina. Int J Plant Sci. 2014;175:955–961. Available from:
- Quija-Lamina P, Segovia-Salcedo MC, Jadán M, et al. Estandarización de la metodología para el conteo cromosómico en especies del género Polylepis en el Ecuador. Rev Ecuatoriana de Med y Cienc Biológicas. 2010;31:33–49.
- Quijia-Lamina P, Jadán M, Proaño K, et al. Estudio Citogenetico de las especies del Genero Polylepis (P. incana y P. racemosa) en el Ecuador. Conference Ciencia y Tecnología; Ecuador; 2010.
- Montalvo JL Determinación del número cromosómico de P. pauta y P. serícea presentes en la Provincia de Pichincha [Thesis]. Sangolquí. Escuela Politécnoca del Ejército; 2013.
- Zurita CF, Segovia-Salcedo MC, Jadán M, et al. Análisis Cromosómico de la especie P. incana en tres poblaciones de la zona centro norte del Ecuador (El Inga, Papallacta, El Angel) a partir de meristemos radicales. Conference Ciencia y Tecnología; Ecuador; 2013.
- Segovia-Salcedo MC New insight into the evolutionary history of the complex Andean genus Polylepis (Rosaceae: sanguisorbeae) and its implications for conservation and management [dissertation]. Florida (FI): University of Florida; 2014.
- Segovia-Salcedo MC, Quijia-Lamina P. Citogeografía de cuatro especies de Polylepis (Rosaceae) en el Ecuador: información relevante para el manejo y conservación de los bosques andinos. In: Cuesta F, Sevink FJ, Llambi LDeditors. Avances en investigación para la conservación de los páramos andinos. Quito, Pichincha, Ecuador: CONDESAN; 2014. p. 467–485.
- Caiza JC, Vargas D, Olmedo C, et al. Morfometría y morfología de estomas y de polen como indicadores indirectos de poliploidía en especies del género Polylepis (Rosaceae) en Ecuador. Ecología Austral. 2018;28:175–187. Available from: https://doi.org/https://doi.org/10.25260/EA.18.28.1.1.528
- Stebbins GL. Chromosomal evolution in higher plants. London: Edward Arnold (Publishers) Ltd.; 1971. p. viii-+ 216.
- Morgan D, Soltis D, Robertson A. Systematic and Evolutionary Implications of rbcL sequence variation in Rosaceae. Am J Bot. 1994;81:890–903. Available from: www.jstor.org/stable/2445770
- Mishima M, Ohmido N, Fukui M, et al. Trends in site-number change of rDNA loci during polyploid evolution in Sanguisorba (Rosaceae). Chromosoma. 2002;110:550–558. Available from: https://doi.org/https://doi.org/10.1007/s00412-001-0175-z
- Boza TE, Quispe-Melgar H, Kessler M. Taxonomic reevaluation of the Polylepis sericea Complex (Rosaceae), with the description of a new species. Syst Bot. 2019;44(2):324–334. Available from: https://doi.org/https://doi.org/10.1600/036364419X15562052252225
- Romoleroux K. Rosaceae 79. In: Harling G, Andersson L, editors. Flora of Ecuador. Vol. 56. Göteborg/Stockholm/Quito: University of Gothenburg/Riksmuseum/Pontificia Universidad Católica del Ecuador; 1996. p. 1–159.
- Boza TE, Romoleroux K, Kessler M. Taxonomic revaluation of the Polylepis pauta and P. sericea (Rosaceae) from Ecuador. Phytotaxa. 2020454 (2): 111–126. https://doi.org/https://doi.org/10.11646/phytotaxa.454.2.3
- Dickinson TA, Lo E, Talent N. Polyploidy, reproductive biology, and Rosaceae: understanding evolution and making classifications. Plant Syst Evol. 2007;266(1–2):59–78. Available from: https://doi.org/https://doi.org/10.1007/s00606-007-0541-2
- Eriksson T, Hibbs MS, Yoder AD, et al. The phylogeny of Rosoideae (Rosaceae) based on sequences of the internal transcribed spacers (ITS) of nuclear ribosomal DNA and the trnL/F region of chloroplast DNA. Int J Plant Sci. 2003;164:197–211. Available from: https://doi.org/https://doi.org/10.1086/346163
- Kerr MS A phylogenetic and biogeographic analysis of Sanguisorbae (Rosaceae), with emphasis on the pleistocene radiation of the high andean genus Polylepis [dissertation]. Maryland (MD) University of Maryland; 2004.
- Soltis DE, Soltis PS, Tate JA. Advances in the study of polyploidy since plant speciation. New Phytol. 2004;161(1):173–191. Available from: https://doi.org/https://doi.org/10.1046/j.1469-8137.2003.00948.x
- Mendoza W, Cano A. El género Polylepis en el Perú Taxonomía, Morfología y Distribución. Editorial Académica Española; AV Akademikerverlag GmbH & Co. KG. Saarbrücken, Deutschland / Alemania. 2012.
- Talent N, Dickinson TA. Endosperm formation in aposporous Crataegus (Rosaceae Spiraeoideae tribe Pyreae): parallels to Ranunculaceae and Poaceae. New Phytol. 2007;173(2):231–249. Available from: https://doi.org/https://doi.org/10.1111/j.1469-8137.2006.01918.x
- Paterson AH. Polyploidy evolutionary opportunity and crop adaptation. Genética. 2005;123:191–196. Available from: https://doi.org/https://doi.org/10.1007/s10709-003-2742-0
- Matsuoka Y. Evolution of polyploid Triticum wheats under cultivation: the role of domestication natural hybridization and allopolyploid speciation in their diversification. Plant Cell Physiol. 2011;52(5):750–764. Available from: https://doi.org/https://doi.org/10.1093/pcp/pcr018
- Sattler MC, Carvalho CR, Clarindo WR. The polyploidy and its key role in plant breeding. Planta. 2016;243(2):281–296. Available from: https://doi.org/https://doi.org/10.1007/s00425-015-2450-x
- Oldfield S, Lusty C, MacKinven A, eds. The world list of threatened trees. Cambridge: World Conservation Press; 1998.
- Evans RC, Dickinson TA. North American black‐fruited hawthorns. II. Floral development of 10‐and 20‐stamen morphotypes in Crataegus section Douglasii (Rosaceae: maloideae). Am J Bot. 1996;83(8):961–978. Available from: https://doi.org/https://doi.org/10.1002/j.1537-2197.1996.tb12793.x
- Enrico L, Funes G, Cabido M. Regeneration of Polylepis australis Bitt. in the mountains of central Argentina. For Ecol Manage. 2004;190(2–3):301–309. Available from: https://doi.org/https://doi.org/10.1016/j.foreco.2003.10.020
- Renison D, Hensen I, Cingolani AM. Anthropogenic soil degradation affects seed viability in Polylepis australis mountain forests of central Argentina. For Ecol Manage. 2004;196(2–3):327–333. Available from: https://doi.org/https://doi.org/10.1016/j.foreco.2004.03.025
- Menoyo E, Renison D, Becerra AG. Arbuscular mycorrhizas and performance of Polylepis australis trees in relation to livestock density. For Ecol Manage. 2009;258(12):2676–2682. Available from: https://doi.org/https://doi.org/10.1016/j.foreco.2009.09.031
- Samaniego F, Kolár F, Urfus T, et al. Determination of apomixis by flow cytometry in two species of Lachemilla (Rosaceae) in Ecuador. Neotrop Biodivers. 2018;4(1):152–163. Available from: https://doi.org/https://doi.org/10.1080/23766808.2018.1542785
