Abstract
Next-generation sequencing was used to obtain the complete mitogenome of Henricia pachyderma (Hayashi, Citation1940). The mitogenome form was found to be a circular molecule 16,192 bp long with a 57.1% AT bias. Gene arrangement and composition of H. pachyderma were exactly the same as those of previously reported mitogenomes from other species of the family Echinasteridae, including H. leviuscula and Echinaster (Othilia) brasiliensis. Phylogenetic analysis with maximum likelihood and Bayesian inference revealed that H. pachyderma formed a monophyletic clade with H. leviuscula and E. (O.) brasiliensis belonging the same taxon, family Echinasteridae.
The genus Henricia is a major taxon of sea stars along the coast of the North Pacific and has high species diversity (Wakita et al. Citation2019). Therefore, some studies have attempted analysis of species identification and phylogenetic relationships using molecular phylogenetic methods (Chichvarkhin Citation2017; Knott et al. Citation2018; Wakita et al. Citation2019). However, it is difficult to define species boundaries in the genus Henricia, and this remains an issue in the phylogeny of the genus (Clark and Downey Citation1992; Mah and Blake Citation2012; Chichvarkhin Citation2017). For the above-mentioned reasons, mitogenomic studies of the genus Henricia have not addressed numerous cases. In this study, we carefully identified Henricia specimens based on morphological comparisons with previous morphological studies showing clearly elucidated results (Hayashi Citation1940; Djakonov Citation1961). The complete mitochondrial genome sequence of H. pachyderma was characterized and a phylogenetic analysis within the class Asteroidea with data from GenBank was performed.
A specimen was collected from waters adjacent to Pohang in the East Sea, Korea (36°11′58″N, 129°24′47″E) at a depth of 15 m, by scuba diving. Voucher specimens and mitochondrial DNA samples were deposited in the Marine Echinoderm Resources Bank of Korea (Seoul, Korea) and granted a voucher number: MERBK-A-1261. Mitochondrial DNA analyses conformed to the method described by Lee and Shin (Citation2018). Phylogenetic analysis was performed with 32 complete mitogenomes of echinoderms (16 asteroids, including H. pachyderma, four crinoids, four ophiuroids, four echinoids, and four holothuroids) and two hemichordates (Balanoglossus carnosus and B. clavigerus), which were used as outgroups for this analysis. Phylogenetic analysis of the mitogenome nucleotide sequence dataset was performed using the maximum likelihood (ML) method with RAxML 8.2 (Stamatakis Citation2014) and Bayesian inference (BI) with MrBayes 3.2 (Ronquist et al. Citation2012).
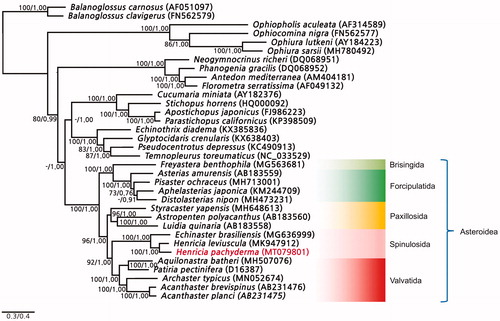
The mitogenome sequence of H. pachyderma was 16,192 bp long and was submitted to GenBank under the accession number MK947912. The gene arrangement of H. pachyderma was exactly the same as that of previously reported mitogenomes for other species of the family Echinasteridae, such as H. leviuscula and Echinaster (Othilia) brasiliensis. The overall nucleotide composition of H. pachyderma was 29.2% A, 24.8% C, 18.1% G, and 27.9% T with a 57.1% AT bias. The 11 protein-coding gene sequences (PCGs) were initiated with the start codon (methionine), but ND3 and ND4L initiated with the isoleucine, ATT and ATC, respectively. The 12 PCGs terminated with a termination codon (TAA or TAG); however, CytB did not have a termination codon, and the 3′ end terminated with phenylalanine (TTC)+T. To reveal the phylogenetic analysis of H. pachyderma within the class Asteroidea, a phylogenetic tree was constructed based on the concatenated sequences of 13 PCGs identified using the ML and BI methods. Henricia pachyderma was closely clustered with H. leviuscula and Echinaster (Othilia) brasiliensis belonging to the same taxon, family Echinasteridae (). The mitogenome newly obtained in this study expanded the genomic resources available for further evolutionary studies of the class Asteroidea among others and will play a significant role in conservation genetics.
Figure 1. Phylogenetic analysis of Henricia pachyderma and an additional 31 echinoderms performed with the maximum likelihood and Bayesian inference methods based on the nucleotide sequences of 13 protein-coding gene sequences. Two hemichordates (Balanoglossus carnosus and B. clavigerus) were used as outgroups. The bootstrap support and posterior properties values are indicated on each node as >70 and 0.7.
Disclosure statement
The authors report no conflicts of interest. The authors alone are responsible for the content and writing of this article.
Additional information
Funding
References
- Chichvarkhin AY. 2017. Sea star Henricia spiculifera (Clark, 1901) in the northwestern Pacific: one species or three? PeerJ. 5:e3489.
- Clark AM, Downey ME. 1992. Starfishes of the Atlantic. London: Chapman and Hall. p. 794.
- Djakonov AM. 1961. Review of sea stars of the genus Henricia Gray from the northwestern parts of Pacific Ocean. Issledovaniya Delnevostochnykh Morei SSSR. 7:1–39. (in Russian).
- Hayashi R. 1940. Contributions to the classification of the sea-stars of Japan. I. Spinulosa J Fac Sci Hokkaido Univ Ser 6 Zool. 7:107–204.
- Knott KE, Ringvold H, Blicher ME. 2018. Morphological and molecular analysis of Henricia Gray, 1840 (Asteroidea: Echinodermata) from the Northern Atlantic Ocean. Zool J Linn Soc. 182(4):791–807.
- Lee T, Shin S. 2018. Complete mitochondrial genome of the taxonomically notorious sea star, Henricia leviuscula (Asteroidea, Spinulosida, Echinasteridae), from South Korea. Mitochondrial DNA Part B Resour. 3(2):1290–1291.
- Mah CL, Blake DB. 2012. Global diversity and phylogeny of the Asteroidea (Echinodermata). PLoS One. 7(4):e35644.
- Ronquist K, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchar-D MA, Huelsenbeck JP. 2012. MrBayes 3.2: efficient Bayesian phylogenetic Inference and model choice across a large model space. Syst Biol. 61(3):539–542.
- Stamatakis A. 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 30(9):1312–1313.
- Wakita D, Fujita T, Kajihara H. 2019. Molecular systematics and morphological analyses of the subgenus Setihenricia (Echinodermata: Asteroidea: Henricia) from Japan. SpecDiv. 24(2):119–135.
