Abstract
The complete mitochondrial genome of Fusarium tricinctum was sequenced. The circular molecule is a total length of 46,314 bp, and the base composition of the mitogenome is as follows: A (33.6%), T (33.1%), C (15.1%), and G (18.2%). The mitogenome contains 19 protein-coding genes, 2 ribosomal RNA (rRNA), and 27 transfer RNA (tRNA) genes. The mitogenome analysis of F. tricinctum provides a molecular basis for further studies on molecular systematics and evolutionary dynamics.
Fusarium is a large genus of filamentous fungi, which is widely distributed in soil and associated with plants (Nelson et al. Citation1994). Most species in this genus are harmless saprobes and also include a number of economically important plant pathogenic species (Moretti Citation2009). Fusarium tricinctum strain KGS17 (GenBank accession number MN365076) was isolated from potato root that was rotted in Gansu, northwest China (35°26.346′N, 104°50.991′E) and identified based on translation elongation factor 1a gene and internal transcribed spacer (White et al. Citation1990; Samson et al. Citation2004). The voucher specimen (No. KGS17) was deposited at the Gansu Provincial Key Lab of Aridland Crop Science, Lanzhou, Gansu, China. Fusarium tricinctum was stored in Gansu Academy of Agricultural Sciences. The total genomic DNA mycelia obtained was extracted using Fungal DNA Kit D3390-02 (Omega Bio-Tek, Norcross, GA) according to the manufacturer’s instructions and was stored in the sequencing company (Xuan Chen Biological Technology Co., Ltd. Shaanxi, China). Purified DNA was used to construct the sequencing libraries following the instructions of NEBNext® UltraTM II DNA Library Prep Kit (NEB, Beijing, China). Whole genome sequencing was performed using the Illumina novaseq6000 Platform (Illumina, San Diego, CA). Multiple steps were used for quality control and de novo assembly of the mitogenome (Bi Citation2017). Adapters and low-quality reads were removed using the NGS QC Toolkit (Patel and Jain Citation2012).
The obtained clean reads were screened out by Bowtie 2 (Langmead and Salzberg Citation2012) with the help of homologous reference sequence (Fusarium commune: LT906348), and then assembled as implemented by NOVOPlasty v3.8.3 (Dierckxsens et al. Citation2017). Genome annotation is mainly carried out by comparing the mitochondrial genome (see phylogenetic tree for details) of the same genus and relative species of GenBank. Geneious R11 software (https://www.genetic.com) was used to make mitochondrial genome map. The length of F. tricinctum mitogenome is 46,314 bp, and its size, structure, and gene content are similar to those of Fusarium. This mitogenome was submitted to the GenBank database under accession no. MT269798. The circular mitogenome contains 19 protein-coding genes, 2 ribosomal RNA (rRNA), and 27 transfer RNA(tRNA) genes (). The base composition of the genome is as follows: A (33.6%), T (33.1%), C (15.1%), and G (18.2%).
Figure 1. Phylogenetic relationships among 13 Fusarium mitogenomes. This tree was drawn with potato Fusarium tricinctum as an outgroup. The length of branch represents the divergence distance.
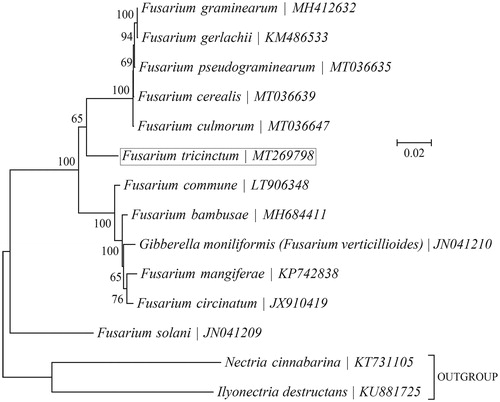
To validate the phylogenetic position of F. tricinctum, the genome-wide alignment of Fusarium mitogenomes was constructed by Geneious R11 software (https://www.genetic.com). Genes with good comparison effect were selected and then connected into a single super gene. It was exported to topali v2.5 software (Milne et al. Citation2009) to build the phylogenetic tree ().
As shown in , Fusarium cerealis (MT036639) and Fusarium culmorum (MT036647) are determined as sisters of Fusarium tricinctum with strong support. High bootstrap and posterior probability values show that presented relations are stable. The mitochondrial genome of Fusarium tricinctum will contribute to the understanding of phylogeny.
Disclosure statement
No potential conflict of interest was reported by the author(s).
Data availability statement
The data that support the findings of this study are openly available in NCBI GenBank database at (https://www.ncbi.nlm.nih.gov/nuccore/) with the accession number is MT269798, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Additional information
Funding
References
- Bi GQ. 2017. The complete mitochondrial genome of northern grasshopper mouse (Onychomys leucogaster). Mitochondrial DNA Part B. 2(2):393–394.
- Dierckxsens N, Mardulyn P, Smits G. 2017. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 45(4):e18.
- Langmead B, Salzberg SL. 2012. Fast gapped-read alignment with Bowtie 2. Nat Methods. 9(4):357–359.
- Milne I, Lindner D, Bayer M, Husmeier D, McGuire G, Marshall DF, Wright F. 2009. TOPALi v2: a rich graphical interface for evolutionary analyses of multiple alignments on HPC clusters and multi-core desktops. Bioinformatics. 25(1):126–127.
- Moretti A. 2009. Taxonomy of Fusarium genus: a continuous fight between lumpers and splitters. Zb Mat Srp Prir Nauk. 117(117):7–13.
- Nelson PE, Dignani MC, Anaissie EJ. 1994. Taxonomy, biology, and clinical aspects of Fusarium species. Clin Microbiol Rev. 7(4):479–504.
- Patel RK, Jain M. 2012. NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS One. 7(2):e30619.
- Samson RA, Hoekstra ES, Frisvad JC. 2004. Introduction to food-and airborne fungi. In: Tjamos EC, Papavizas GC, Cook RJ, editors. 7th ed. Boston, MA: Centraal Bureau voor Schimmelcultures (CBS).7th ed. Boston, MA.
- White TJ, Sninsky JJ, Gelfand DH, Innis MA. 1990. PCR protocols: a guide to methods and applications. San Diego: Academic Press. p. 315.
