Abstract
Ardisia bullata G. H. Huang & G. Hao is a small shrubs of Primulaceae. It is only distributed in Hainan provinces of China. It is a plant medicinal value. There is no study on the genome of A. bullata so far. Here, we report and characterize the complete plastid genome sequence of A. bullata in an order to provide genomic resources useful for promoting its conservation. The complete chloroplast genome of A. bullata is 160,176 bp in length with a typical quadripartite structure, consisting of a large single-copy region (LSC, 89,710 bp), a single-copy region (SSC, 18,357 bp), and a pair of inverted repeats (IRs, 26,054 bp). There are 133 genes annotated, including 83 unique protein-coding genes, eight unique ribosomal RNA genes, and 37 transfer RNA genes. The overall G/C content in the plastome of A. bullata is 36.0%. The complete plastome sequence of A. bullata will provide a useful resource for the conservation genetics of this species as well as for phylogenetic studies in Apocynaceae.
Ardisia bullata G. H. Huang & G. Hao is a shrub of the family Primulaceae. Its distribution range is extremely narrow, only in Hainan provinces of China. It is a plant combines medicinal and dyestuff value (Chen and Pipoly Citation1996). The chloroplast genome sequence carries rich information for plant molecular systematics and Barcoding. To date, there have been no studies on the genome of A. bullata. To provide a rich genetic information and improve A. bullata molecular breeding in the future, we report and characterize the complete plastid genome sequence of A. bullata (GenBank accession number: MT505713).
In this study, the fresh leaves of C. sappan were collected from Diaoluo Mountain in Hainan province (109.91° E, 18.67° N). Voucher specimens (HUTB 187225) were deposited in the Herbarium of the Institute of Tropical Agriculture and Forestry (code of herbarium: HUTB), Hainan University, Haikou, China.
The experiment procedure was as reported in Wang et al. (Citation2019). The total DNA of A. bullata was sequenced with second-generation sequencing technology (Illumina HiSeq 2000, San Diego, CA). The chloroplast genome sequence reads were assembled with bioinformatic pipeline including SOAP2 software (Li et al. Citation2009) and several runs of manual corrections of sequence reads. Genes encoded by this genome were annotated by import the fasta format sequence to the DOGMA (Wyman et al. Citation2004) and recorrected by manual. The results showed that plastome of A. bullata possesses a total length of 160,176 bp with the typical quadripartite structure of angiosperms, containing two Inverted Repeats (IRs) of 26,054 bp, a Large Single-Copy (LSC) region of 89,710 bp, and a Small Single-Copy (SSC) region of 18,357 bp. The plastome contains 133 genes, consisting of 88 unique protein-coding genes, 37 unique tRNA genes, and eight unique rRNA genes. The overall G/C content in the plastome of A. bullata is 36.0%, in which the corresponding values of the LSC, SSC, and IR region were 33.40%, 29.90%, and 42.60%, respectively.
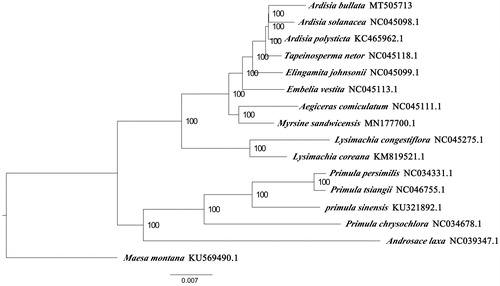
We used RAxML (Stamatakis Citation2006) with 1000 bootstraps under the GTRGAMMAI substitution model to reconstruct a maximum-likelihood (ML) phylogeny of 16 published complete plastomes of Primulaceae, using Maesa montana (Primulaceae) as outgroups. According to the phylogenetic topologies, A. bullata was closely related to A. solanacea. Most nodes in the plastome ML trees were strongly supported (). The complete plastome sequence of A. bullata will provide a useful resource for the conservation genetics of this species as well as for the phylogenetic studies for Primulaceae.
Figure 1. Maximum-likelihood phylogenetic tree based on 16 complete chloroplast genomes. Ardisia bullata (this study); Ardisia solanacea NC045098.1; Ardisia polysticta KC465962; Tapeinosperma netor NC045118.1; Elingamita johnsonii NC045099.1; Embelia vestita NC045113.1; Aegiceras comiculatum NC045111.1; Myrsine sndwicensis MN177700.1; Lysimachia congestiflora NC045275.1; Lysimachia coreana KM819521.1; Primula persimilis NC034331.1; Primula tsiangii NC046755.1; Primula sinensis KU321892.1; Primula chrysochlora NC034678.1; Androsace laxa NC039347.1; outgroup: Maesa Montana KU569490.1. The number on each node indicates the bootstrap value.
Disclosure statement
The authors report that they have no conflicts of interest. The authors alone are responsible for the content and writing of the paper.
Data availability statement
The data that support the findings of this study are openly available in GenBank of NCBI at http://www.ncbi.nlm.nih.gov, reference number MT505713.
Additional information
Funding
References
- Chen J, Pipoly JJ. 1996. Myrsinaceae. In: Wu ZY, Raven PH, editors. Flora of China, Vol. 15. Beijing: Science Press. p. 1–38.
- Li RQ, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, Wang J. 2009. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics. 25(15):1966–1967.
- Stamatakis A. 2006. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 22(21):2688–2690.
- Wang HX, Chen L, Cheng XL, Chen WS, Li LM. 2019. Complete plastome sequence of Callicarpa nudiflora Vahl (Verbenaceae): a medicinal plant. Mitochondrial DNA Part B. 4(2):2090–2091.
- Wyman SK, Jansen RK, Boore JL. 2004. Automatic annotation of organellar genomes with DOGMA. Bioinformatics. 20(17):3252–3255.
