Abstract
Halocynthia hilgendorfi ritteri is an ascidian distributed on the coast of Geoje Island in Korea and found on rocks. The mitochondrial genome of Halocynthia hilgendorfi ritteri consists of 15,181 bp with 13 protein-coding genes, 2 ribosomal RNAs, 23 transfer RNA genes. The overall base composition of the complete genome is 22.94% A, 43.32% T, 25.72% G, and 8.02% C, with a high A + T content of 66.26%.
Halocynthia hilgendorfi ritteri is a subspecies of Halocynthia hilgendorfi that belongs to the family Pyuridae, order Pleurogona. It is native to Asia and occurs off the coast of Jeju Island, Korea (Oka Citation1906; Tokioka Citation1959; Choi, Lee, et al. Citation2004). Halocynthia hilgendorfi ritteri is found mainly on rock surfaces and its shape and color are similar to those of the surrounding rocks (Choi, Kim, et al. Citation2004). Halocynthia hilgendorfi ritteri, an important fishery resource in Korea, is widely consumed and popular. The species in the genus Halocynthia are distinct when alive, but are difficult to distinguish when processed. Therefore, we would like to report the complete mitogenome sequence of H. hilgendorfi ritteri for developing a marker to identify this subspecies.
Halocynthia hilgendorfi ritteri were collected from Geoje Island (34°53′N, 128°37′E) and DNA was extracted using a DNeasy Blood and Tissue Kit (QIAGEN, Hilden, Germany). The genomic DNA is preserved at Soonchunhyang University, Republic of Korea. A complete mitochondrial genome was obtained according to the MGISEQ-2000 protocol (MGI, China) using an MGI Easy DNA Library Prep Kit (MGI, Wuhan, China). The H. hilgendorfi ritteri sequence has been registered at NCBI GenBank (Accession no. MT811760).
The complete mitochondrial genome of H. hilgendorfi ritteri consists of 15,181 bp, with 13 protein-coding genes (PCGs), 2 ribosomal RNA genes (rRNAs), and 23 transfer RNA genes (tRNAs). The CO1, Cytb, ND2, ND4L, and ATP6 genes include the ATG start codon; the remaining eight PCGs start with GTG. Six PCGs (CO3, Cytb, ND2, ND4L, ND5, and ND6) terminate with TAG, a typical stop codon and seven PCGs (CO1, CO2, ND1, ND3, ND4, ATP6, and ATP8) end with a complete TAA. All the genes are encoded on the heavy strand, except for the two tRNA genes (Arg and Ser1) and 12 PCGs (ATP6, ATP8, CO1, CO3, Cytb, ND1, ND2, ND3, ND4, ND4L, ND5, and ND6). The overall base composition of the H. hilgendorfi ritteri genome is 22.94% A, 43.32% T, 25.72% G, and 8.02% C, with a high A + T content of 66.26%. rRNA consists of small rRNA (672 bp) and large rRNA (1126 bp).
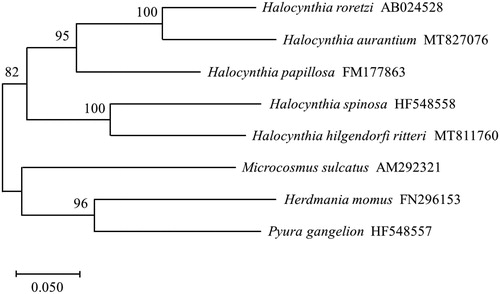
We used the maximum likelihood method to construct a phylogenetic tree based on the complete CO1 of Halocynthia species and included other Pyuridae family sequences as an outgroup (). This method identified H. hilgendorfi ritteri as being closely related to Halocynthia spinosa.
Disclosure statement
No conflict of interest was reported by the author(s).
Data availability statement
The data that support the findings of this study are openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/nuccore/MT811760, reference number MT811760.
Additional information
Funding
References
- Choi YJ, Lee CH, Rho S, Lee YD. 2004. Reproductive cycle and spawning rhythm of the ascidian, Halocynthia hilgendorfi ritteri. Korean J Biol Sci. 8(1):33–40.
- Choi YJ, Kim SY, Lee CH, Rho S, Lee YD. 2004. Early development of the ascidian (Halocynthia hilgendorfi ritteti). Korean J Fish Aquat Sci. 37(2):98–104.
- Oka A. 1906. Notizen uber japanische Ascidien I. Annot Zool Japon. 6:37–52.
- Tokioka T. 1959. Contributions to Japanese ascidian fauna – XIII. Sporadic memoranda (4). Publ Smbl. 7(2):237–239.