Abstract
The genus Coelastrum Nägeli (Sphaeropleales; Scenedesmaceae) is a diverse genus of green algae with potential biotechnical applications. A sound understanding of its phylogeny will be a useful tool for predicting the distribution of traits that may enhance its utility, and may lead to a better understanding of its evolution and ecology. Here we present the plastome of Coelastrum microporum. Our exemplar was isolated from Gull Lake, Michigan and the complete plastome as assembled was 169,961 bp in length. The plastome contained 104 genes of which 68 were protein-coding genes (CDSs), 27 tRNA genes and three rRNA genes. The GC content of the plastome was 31.2%. The maximum likelihood phylogeny suggested that C. microporum was the sister group to a clade of single exemplars of three other genera in the Scenedesmaceae (Tetradesmus, Pectinodesmus and Coelastrella).
Introduction
Coelastrum microporum Nägeli, 1855 is a broadly distributed freshwater green alga with three-dimensional, spherical coenobia. This genus has recently been investigated for biotechnological applications such as bioremediation of wastewater, production of carotenoids, fatty acids, and biodiesel (Bhuyar et al. Citation2021; Liu et al. Citation2020; Maltsev et al. Citation2021). Understanding the plastome is known to be important to understanding synthesis and regulation of unsaturated fatty acids (He et al. Citation2020). Phylogenetic reconstructions have been attempted using marker genes such as ITS, and 18s rDNA regions (Goecke et al. Citation2020; Hegewald et al. Citation2010), with Coelastrum Nägeli itself recovered as monophyletic or not. The plastome of C. microporum has not been published previously. To help better understand evolutionary relationships of Coelastrum, we fully sequenced the plastome of C. microporum and compared it to available plastomes of other Sphaeropleales.
Materials and methods
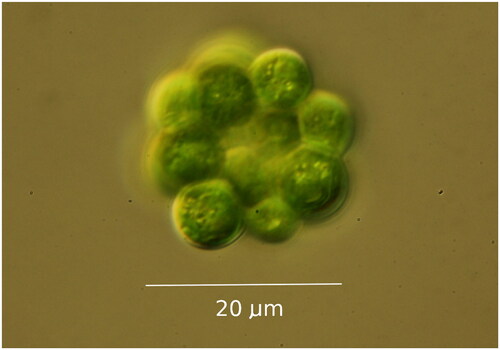
Coelastrum microporum was collected from Gull Lake in southwest Michigan (42.403061 N; 85.414341 W) in July 2019. The isolated strain was cultured in WC artificial freshwater medium (Guillard Citation1975) at 14 °C (a temperature similar to lakes when samples were collected, which should be conducive for the maximum growth and survival rate of most species (Thomas et al. Citation2016)). The strain used is available from the UTEX Culture Collection of Algae (https://utex.org, Dr. David Nobles, Curator and Director: [email protected]) as Coelastrum microporum UTEX LB 3178. Light microscopy using morphological criteria were used for initial species identification, with the isolate () corresponding to morphological descriptions of C. microporum (Komárek and Fott Citation1983). DNA was extracted using a DNeasy® Plant Mini Kit (Qiagen, Hilden, Germany) following the kit’s protocol. Monoclonal cultures were sampled in late exponential phase after visual inspection in the light microscope of the culture for fungal hyphae and bacterial abundance. Approximately 30 million 150 bp paired-end reads were sequenced using Illumina HiSeq 4000 platform (Illumina, San Diego, CA) at the Genome Sequencing and Analysis Facility (GSAF) at the University of Texas at Austin. The raw reads were trimmed using BBDuk from the BBTools software package (https://jgi.doe.gov/data-and-tools/bbtools/), and assembled with NOVOPlasty v. 4.2.1 (Dierckxsens et al. Citation2017). The assembled contig was imported into Geneious 2020.2.4 (Biomatters Ltd., http://www.geneious.com (Kearse et al. Citation2012)), and compared to vouchered sequences from the NCBI database using BLAST (Altschul et al. Citation1990) to check for any possible contaminants. The contig was annotated with available complete plastomes of closely related species in Geneious. The plastome was mapped using Bowtie2 v.2 (Langmead and Salzberg Citation2012) with trimmed reads to determine sequence coverage and potential mis-assemblies. Verification of protein-coding genes was manually performed in Geneious, and tRNA genes were verified using tRNAscan-SE v.2.0 (Chan and Lowe Citation2019). We extracted 58 CDS sequences shared by C. microporum and 22 publicly accessible plastomes from other Sphaeropleales, aligning them using MAFFT v.7.450 (Katoh and Standley Citation2013) presented in Table S1. ModelFinder (Kalyaanamoorthy et al. Citation2017) was used to find the best fitting model for maximum likelihood analysis. A maximum likelihood phylogeny was constructed using IQTree2 v. 1.6.12 (Minh et al. Citation2020) with 1000 bootstrap replicates.
Results
The complete plastome of Coelastrum microporum (GenBank accession number: NC_068582) was 169,961 bp in length. We annotated 104 genes which included 68 protein-coding genes (CDSs), 27 tRNA genes and three rRNA genes. The average coverage was 645.5 (ranging from 359 to 1337) mapped from 745,406 trimmed raw reads (Figure S1). The circularized plastome had a typical quadripartite construction (), with a large single copy region (length 85,914 bp; GC content 30.2%), and a small single copy region (length 66,611 bp; GC content 29.2%), interrupted by the two inverted repeats, A and B (IRA and IRB, length of each 8,718 bp; GC content 43.7%). The overall plastome GC content was 31.2%. The best model for maximum likelihood analysis was GTR + F+R5. Our maximum likelihood tree () recovered C. microporum as sister to a clade containing Pectinodesmus pectinatus (Meyen) Hegewald et al. Citation2010, Tetradesmus obliquus (Turpin) M.J. Wynne, 2016, and Coelastrella saipanensis N. Hanagata, 2001.
Figure 2. Gene map of Coelastrum microporum plastome. Inverted repeats (IRA and IRB) and two single-copy regions are indicated on the inner circle with G/C content (dark grey) and A/T content (light grey). colored gene boxes are indicated by functional group as shown in the key.
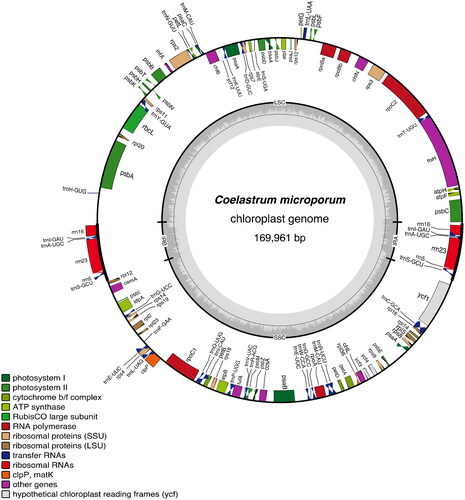
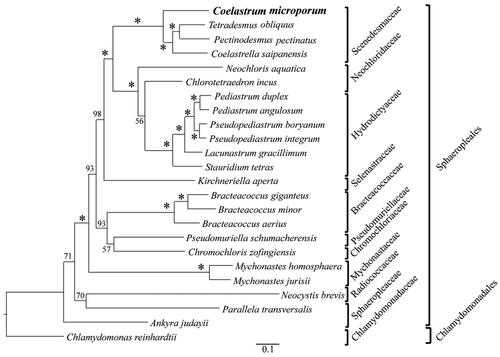
Figure 3. Maximum likelihood phylogeny of Coelastrum microporum, 22 species in Sphaeropleales and Chlamydomonas reinhardtii (outgroup) based on 58 CDSs shared by C. microporum and 23 publicly accessible plastomes in Sphaeropleales and chlamydomonadales. Numbers above branches are bootstrap values from 1000 bootstrap replicates in which asterisks represent bootstrap values of 100. The following sequences were used: Tetradesmus obliquus NC_008101 (de Cambiaire et al. Citation2006), Pectinodesmus pectinatus NC_036668 (unpublished), Coelastrella saipanensis NC_042181 (unpublished), neochloris aquatica NC_029670 (Fučíková et al. Citation2016), chlorotetraedron incus NC_029673 (Fučíková et al. Citation2016), Pediastrum duplex NC_034654 (McManus et al. Citation2017), Pediastrum angulosum NC_037919 (McManus et al. Citation2018), Pseudopediastrum boryanum NC_037920 (McManus et al. Citation2018), Pseudopediastrum integrum NC_037921 (McManus et al. Citation2018), lacunastrum gracillimum NC_037918 (McManus et al. Citation2018), stauridium tetras NC_037923 (McManus et al. Citation2018), kirchneriella aperta NC_029676 (Fučíková et al. Citation2016), Bracteacoccus giganteus NC_028586 (Lemieux et al. Citation2015), Bracteacoccus minor NC_029674 (Fučíková et al. Citation2016), Bracteacoccus aerius NC_029675 (Fučíková et al. Citation2016), pseudomuriella schumacherensis NC_029669 (Fučíková et al. Citation2016), chromochloris zofingiensis NC_029672 (Fučíková et al. Citation2016), Mychonastes homosphaera NC_029671 (Fučíková et al. Citation2016), Mychonastes jurisii NC_028579 (Lemieux et al. Citation2015), neocystis brevis NC_025535 (Lemieux et al. Citation2014), parallela transversalis NC_042241 (unpublished), ankyra judayi NC_029735 (Fučíková et al. Citation2016), Chlamydomonas reinhardtii NC_005353 (Maul et al. Citation2002).
Discussion and conclusions
Goecke et al. (Citation2020), using ITS and 18s rDNA sequences, but much denser taxon sampling than our study, recovered similar relationships among Sphaeropleales genera, as did Hegewald et al. (Citation2010), using only ITS2 sequences. We hesitate to argue for any one of these hypotheses of relationships without further analysis, because different taxon sampling strategies can change inferred phylogenetic relationships even when the same gene(s) and same optimality criterion are applied (Theriot et al. Citation2009). This description of the complete plastome of C. microporum provides additional support for its inclusion in the Sphaeropleales and valuable information for further research on phylogenetic relationships in this order, which in turn can help guide research on practical applications of these algae. We intend to expand our studies to better understand the evolution and classification of the genus Coelastrum.
Authors’ contributions
Edward Theriot, Joshua Cooper, Chanhee Lee, and Trisha Spanbauer conceived and designed the experiments. Edward Theriot acquired and isolated the alga. Chanhee Lee, and Ana Salim grew the algae and extracted DNA; Chanhee Lee, Ana Salim, Chaehee Lee, and Francesca Moroni made substantial contributions to the acquisition and analysis of data; Chanhee Lee and Edward Theriot contributed substantially to authorship; all authors contributed to revision of the manuscript, and all approved the final version to be published. All authors agree to be accountable for all aspects of the work.
Geolocation
Gull Lake, Kalamazoo County, Michigan. 42.403061 N; 85.414341 W.
Research statement
Phytoplankton sampling and cell isolation was done following all local, state, and US Federal regulations, in the publicly accessible Gull Lake. To our knowledge there is no convention on ethical treatment of microalgae.
Supplemental Material
Download MS Word (87.4 KB)Disclosure statement
The authors report that there is no conflict of interest involved.
Data availability statement
The genome sequence data that supported the findings of this study are openly available in GenBank of NCBI at (https://www.ncbi.nlm.nih.gov/) under the accession no. NC_068582. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA909268, PRJNA909268, and SAMN32062636, respectively. The strain used is available from the UTEX Culture Collection of Algae (https://utex.org, Dr. David Nobles, Curator and Director: [email protected]) as Coelastrum microporum UTEX LB 3178.
Additional information
Funding
References
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. 1990. Basic local alignment search tool. J Mol Biol. 215(3):403–410. doi:10.1016/S0022-2836(05)80360-2.
- Bhuyar P, Sundararaju S, Rahim MHA, Maniam GP, Govindan N. 2021. Enhanced productivity of lipid extraction by urea stress conditions on marine microalgae Coelastrum sp. for improved biodiesel production. Bioresour Technol Rep. 15:100696. doi:10.1016/j.biteb.2021.100696.
- Chan PP, Lowe TM. 2019. tRNAscan-SE: searching for tRNA Genes in Genomic Sequences. In: kollmar, M. (Ed.), Gene prediction: methods and Protocols. Springer New York, New York, NY, p. 1–14.
- de Cambiaire JC, Otis C, Lemieux C, Turmel M. 2006. The complete chloroplast genome sequence of the chlorophycean green alga Scenedesmus obliquus reveals a compact gene organization and a biased distribution of genes on the two DNA strands. BMC Evol Biol. 6(1):37. doi:10.1186/1471-2148-6-37.
- Dierckxsens N, Mardulyn P, Smits G. 2017. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucl Acids Res. 45(4):e18-e18. doi:10.1093/nar/gkw955.
- Fučíková K, Lewis PO, Lewis LA. 2016. Chloroplast phylogenomic data from the green algal order Sphaeropleales (Chlorophyceae, Chlorophyta) reveal complex patterns of sequence evolution. Mol Phylogenet Evol. 98:176–183. doi:10.1016/j.ympev.2016.01.022.
- Goecke F, Noda J, Paliocha M, Gislerød HR. 2020. Revision of Coelastrella (Scenedesmaceae, Chlorophyta) and first register of this green coccoid microalga for continental Norway. World J Microbiol Biotechnol. 36(10):149. doi:10.1007/s11274-020-02897-0.
- Guillard RRL. 1975. Culture of phytoplankton for feeding marine invertebrates. In: smith, W.L., Chanley, M.H. (Eds.), Culture of marine invertebrate animals. Plenum Press, New York, p. 26–60.
- He M, Qin CX, Wang X, Ding NZ. 2020. Plant unsaturated fatty acids: biosynthesis and regulation. Front Plant Sci. 11:390. doi:10.3389/fpls.2020.00390.
- Hegewald E, Wolf M, Keller A, Friedl T, Krienitz L. 2010. ITS2 sequence-structure phylogeny in the Scenedesmaceae with special reference to Coelastrum (Chlorophyta, Chlorophyceae), including the new genera Comasiella and Pectinodesmus. Phycologia. 49(4):325–335. doi:10.2216/09-61.1.
- Kalyaanamoorthy S, Minh BQ, Wong TK, Von Haeseler A, Jermiin LS. 2017. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods. 14(6):587–589. doi:10.1038/nmeth.4285.
- Katoh K, Standley DM. 2013. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 30(4):772–780. doi:10.1093/molbev/mst010.
- Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, et al. 2012. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 28(12):1647–1649. doi:10.1093/bioinformatics/bts199.
- Komárek J, Fott B. 1983. Chlorophyceae (Grünalgen) Ordnung: chlorococcales. E. Schweizerbart’sche Verlagsbuchhandlung (Nägele u. Obermiller), Stuttgart.
- Langmead B, Salzberg SL. 2012. Fast gapped-read alignment with Bowtie 2. Nat Methods. 9(4):357–359. doi:10.1038/nmeth.1923.
- Lemieux C, Otis C, Turmel M. 2014. Chloroplast phylogenomic analysis resolves deep-level relationships within the green algal class Trebouxiophyceae. BMC Evol Biol. 14(1):211. doi:10.1186/s12862-014-0211-2.
- Lemieux C, Vincent AT, Labarre A, Otis C, Turmel M. 2015. Chloroplast phylogenomic analysis of chlorophyte green algae identifies a novel lineage sister to the Sphaeropleales (Chlorophyceae). BMC Evol Biol. 15(1):264. doi:10.1186/s12862-015-0544-5.
- Liu Z, Hou Y, He C, Wang X, Chen S, Huang Z, Chen F. 2020. Enhancement of linoleic acid content stimulates astaxanthin esterification in Coelastrum sp. Bioresour Technol. 300:122649. doi:10.1016/j.biortech.2019.122649.
- Maltsev Y, Krivova Z, Maltseva S, Maltseva K, Gorshkova E, Kulikovskiy M. 2021. Lipid accumulation by Coelastrella multistriata (Scenedesmaceae, Sphaeropleales) during nitrogen and phosphorus starvation. Sci Rep. 11(1):19818. doi:10.1038/s41598-021-99376-9.
- Maul JE, Lilly JW, Cui L, DePamphilis CW, Miller W, Harris EH, Stern DB. 2002. The Chlamydomonas reinhardtii plastid chromosome: islands of genes in a sea of repeats. Plant Cell. 14(11):2659–2679. doi:10.1105/tpc.006155.
- McManus HA, Fučíková K, Lewis PO, Lewis LA, Karol KG. 2018. Organellar phylogenomics inform systematics in the green algal family Hydrodictyaceae (Chlorophyceae) and provide clues to the complex evolutionary history of plastid genomes in the green algal tree of life. Am J Bot. 105(3):315–329. doi:10.1002/ajb2.1066.
- McManus HA, Sanchez DJ, Karol KG. 2017. Plastomes of the green algae Hydrodictyon reticulatum and Pediastrum duplex (Sphaeropleales, Chlorophyceae). PeerJ. 5:e3325. doi:10.7717/peerj.3325.
- Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, Lanfear R. 2020. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 37(5):1530–1534. doi:10.1093/molbev/msaa015.
- Theriot EC, Cannone JJ, Gutell RR, Alverson AJ. 2009. The limits of nuclear-encoded SSU rDNA for resolving the diatom phylogeny. Eur J Phycol. 44(3):277–290. doi:10.1080/09670260902749159.
- Thomas MK, Kremer CT, Litchman E. 2016. Environment and evolutionary history determine the global biogeography of phytoplankton temperature traits. Glob Ecol Biogeogr. 25(1):75–86. doi:10.1111/geb.12387.