Abstract
The role that metabolic phenotyping can increasingly play in patient stratification and personalised medicine is discussed. The background to the general approach, comprehensive and simultaneous analysis of small-molecule metabolites in biofluids, tissues and tissue extracts combined with suitable multivariate statistical models, is summarised. The main techniques used (NMR and mass spectrometry) are cited, and the implementation of dedicated phenome centres is explained. Finally, the advantages and limitations, opportunities and drawbacks of the approach are discussed.
1. Metabolic phenotyping background
Many new approaches have been developed recently to provide increased mechanistic understanding within molecular biology. This expanding systems biology approach is a way of integrating information over many aspects and levels of molecular biology utilising multivariate data from various ‘omics’ technologies and employing advances in computation, statistics and bioinformatics. The main levels of organisation considered in systems biology comprise genomics, epigenetics, transcriptomics, proteomics and metabonomics (also called metabolomics and now increasingly metabolic phenotyping), that is, multivariate analyses at the genome, gene transcription, protein and metabolite levels, respectively. Many other ‘omics’ terms have been invented, but only a few of these, such as lipidomics and metallomics, have gained general acceptance.
Metabolic phenotyping comprises the comprehensive, simultaneous profiling of multiple metabolites in a biological system, as well as their temporal changes caused by, for example, genetic, environmental and pharmaceutically induced effects. This is usually carried out by investigating the metabolic content of biological fluids, tissues and their extracts. This approach to understanding metabolic responses of complex systems to some sort of stimulus has been carefully defined with respect to measurement of biological effects Citation[1,2].
Metabonomics is particularly important in systems biology because it provides a ‘top-down’ approach that gives an integrated view of the biochemistry as opposed to the ‘bottom-up’ approach where the effects of an individual gene, or groups of genes, are investigated.
In humans and other complex systems, at the gene, protein and metabolite levels, changes are highly interdependent and can occur on very different time scales. The situation is even more complex because species with an intestinal system have an internal ecosystem of many species of gut microbial organisms that interact with the host to also co-regulate many host biochemical pathways Citation[3].
Urine and plasma are the chief fluids used for metabolic phenotyping because they are easily obtained and these fluids are widely used in clinical chemistry and chemical pathology. However, many different biofluids have been studied including seminal fluids, amniotic fluid, cerebrospinal fluid, synovial fluid, cyst fluids, lung aspirates and dialysis fluids. Analysis of intact tissue biopsy samples and their lipid and aqueous extracts has also been widely employed, as well as investigation of in vitro cell systems Citation[4]. Metabolic phenotypes generated using in vivo magnetic resonance spectroscopy approaches can also be considered as metabonomic data sets.
2. Enabling technologies
The main analytical techniques that are employed are based on NMR spectroscopy and mass spectrometry (MS) Citation[4]. This is because both of these technologies can provide identification and quantitation information on a wide range of compounds simultaneously and with an acceptable analytical precision. MS analysis generally requires a pre-separation of the metabolic components using either gas chromatography or ultra-high-performance liquid chromatography. Other separation modalities such as capillary electrophoresis (CE) are finding increased usage and methods such as electrochemical detection have been employed in a few cases.
The relative strengths and weaknesses of both NMR and MS for metabolic profiling have been well documented Citation[4,5], but the techniques are considered complementary and both are often used in a comprehensive study. NMR spectroscopy has remained a stable platform with incremental improvements in sensitivity, stability and reproducibility. MS continues to develop rapidly and the i-knife concept Citation[6] for real-time diagnosis during surgery by analysing the smoke from a surgical diathermy knife has pushed the whole idea of metabolic phenotyping into real-time analyses.
3. Development of the phenome centre model
Metabolic phenotyping has expanded rapidly into clinical studies but, to date, most studies can only be regarded as pilot-scale and hypothesis generation investigations. Nevertheless, a number of initiatives have been made to generate regional or national centres of excellence. In the USA, the NIH has taken the initiative to fund several regional ‘metabolomics core facilities’ that allow access to analytical and bioinformatics expertise Citation[7]. Although, there is no direct mandate for unification of technologies between centres, there are plans for database sharing and educational integration.
In the UK, the Medical Research Council (along with substantial support from the analytical instrument manufacturers and the UK National Institute for health Research [NIHR]) has awarded funding to build a National phenome centre based on both NMR and MS technologies for large-scale epidemiological urine and plasma/serum cohorts Citation[8]. This is now operational and is sited on the Hammersmith Hospital campus of Imperial College London Citation[9]. Similarly, the NIHR through its funding of Biomedical Research Centres (BRCs) has funded a clinical metabolic phenome centre, sited on the St Mary’s Hospital campus of Imperial College London, for following patient journeys in hospital in order to provide improved diagnosis, prediction of outcome and stratification (i.e., dynamic patient stratification). As the remit of the BRCs is translational medicine, the output of this centre will have to benefit patients in a cost-effective manner in 3 years.
4. Examples of metabolic phenotyping in patient stratification
It is widely held that patient stratification is dominated by genetic variants and similar stratification criteria, and indeed, there are good examples as to how genetics can improve the selection of therapies for particular patient classes, for example, the well-known drug targeting in breast cancer. However, this genomic approach is relatively static as genomic composition does not change much during therapy, whereas metabolism and physiology can change significantly due to the efficacious effects of drugs or the toxic side effects of some therapies.
Examples of static stratification from the cardiovascular field include the prediction of exercise-induced ischemia in patients with suspected coronary artery disease using NMR spectroscopy of serum Citation[10] and prediction of adverse effects after coronary artery bypass surgery using a targeted MS approach based on 69 metabolites in 478 patients Citation[11]. In the oncology field, there has been success in using the approach based on NMR spectra of serum for prediction of toxicity in 54 patients with inoperable colorectal cancer who were treated with capecitabine Citation[12] and using MAS-NMR of tumor tissue samples, it was possible to identify profiles that enabled colorectal cancer staging and prognosis Citation[13].
There is, therefore, scope for the development of novel dynamic approaches to stratification based on physiological responses that can be modelled in the patient journey Citation[14], and these would revolve around the use of metabonomics. A number of recent studies have shown the potential of metabolic phenotyping for patient stratification and prognosis and the studies that use pre-treatment metabolic information to predict post-treatment outcomes (pharmacometabonomics) have been reviewed recently Citation[15].
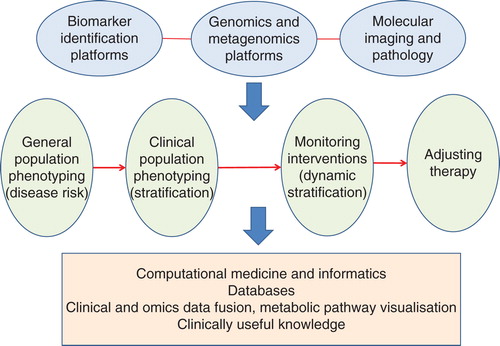
The spread of stratification methods and approaches is illustrated in . The top row indicates, in general, the types of technology platforms used as further explained in the section on enabling technologies and in the figure caption, the second row shows the trajectory from population screening through static patient stratification to the concept of dynamic stratification and the ability to tailor therapies and monitor the patient’s response and the bottom row indicates some of the deliverables from such an approach.
Figure 1. A schematic showing how metabolic phenotyping fits with other biomarker technologies such as genomics and imaging, leading to various types of patient stratification strategies from disease risk in populations, via static and dynamic stratification through to the ability to monitor and change therapies. The outputs of such a process include large databases for retrospective studies, and for improving future predictions, and a better understanding of human metabolism, leading to improved clinical practice.
5. Expert opinion
Over the past decade, the applications of metabolic phenotyping based on biochemical analysis of body fluids such as urine and serum have burgeoned from mainly animal model studies of drug effects to real clinical applications for diagnosis and prediction of treatment effects. This trend will surely continue now that the required analytical facilities are being installed directly into hospitals for real-time studies in the form of clinical phenome centres. Such activities are complemented by large-scale metabolic phenotyping on biofluid samples taken from epidemiological cohorts that provide information on disease risk factors in populations Citation[16].
One requirement still to be achieved is to transform the huge data sets that can be acquired and the complex statistical and other computational processes necessary to interpret the data into formats that are easily understood by clinicians to aid their decision processes and the wider lay population so that they are better informed about their personalised treatment. This becomes even more true when different ‘omics’ data sets have to be combined, as for example, in distilling out the required knowledge from complex combined genomic, transcriptomic, epigenetic and metabolic data sets in order to unravel the effects of genes the environment and life style on an individual’s disease diagnosis, progression, treatment and outcome.
Nevertheless, some targets are already being met, such as in personalising surgical procedures based on metabolic phenotyping when the patient enters hospital and during their stay there. Surgery is a particularly attractive area as a test bed because currently the discipline is largely molecule-free, decisions have to be taken very quickly and the consequences of such decisions can be life-threatening Citation[14]. Moreover, such an approach leads to the concept of the metabolic phenotypically augmented clinical trial as a means of testing new drugs for efficacy and adverse effects, allowing improved patient/subject selection and earlier prediction of post-treatment effects.
Turning such biochemical information into real clinical tests is still some way off, but efforts are currently in place in translational research centres to provide real patient benefits within 3 years. Such tests have to be faster, cheaper, more accurate or more robust that the current technologies and it is clearly necessary to build in cost-benefit calculations to any such translational medicine studies. Moreover, the unique ability of metabolic profiling to follow changes in phenotype before, during and after therapy will lead to a new paradigm of dynamic patient stratification.
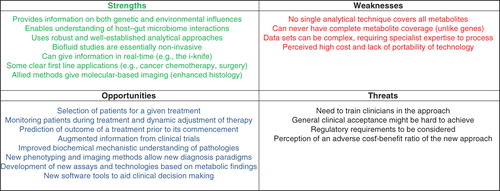
These views are summarised in , which shows the perceived strengths, weaknesses, opportunities and threats relating to the emerging use of metabolic phenotpying in patient stratification.
Figure 2. An analysis of the strengths, weaknesses, opportunities and threats relating to metabolic phenotyping for patient stratification.
In summary, we can expect to see a series of new clinical assays based on NMR and MS discovery (either using these techniques in the clinic or assays based on the metabolic profiles obtained), in particular the use of the i-knife concept in general surgery, and in wider applications such as endoscopy, colonoscopy and bronchoscopy for real-time diagnosis. New methodologies that are emerging in ‘big data’ analyses will enhance the information recovery from metabolic profiles and allow clinicians and other non-specialists to use this knowledge effectively in decision making.
Declaration of interests
The authors are employees of Imperial College London and also Directors and stock-holders in Metabometrix Ltd. This work was produced without specific funding. JK Nicholson also has paid consultancies with analytical instrument companies. The authors have no other relevant affiliations or financial involvement with any organisation or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
Bibliography
- Nicholson JK, Lindon JC, Holmes E. “Metabonomics”: understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica 1999;29:1181-9
- Nicholson JK, Connelly J, Lindon JC, et al. Metabonomics: a platform for studying drug toxicity and gene function. Nat Rev Drug Discov 2002;1:153-61
- Yatsunenko T, Rey FE, Manary MJ, et al. Human gut microbiome viewed across age and geography. Nature 2012;486:222-7
- Lindon JC, Nicholson JK. Spectroscopic and statistical techniques for information recovery in metabonomics and metabolomics. Annu Rev Anal Chem 2008;1:45-69
- Bouatra S, Aziat F, Mandal R, et al. The human urine metabolome. PLoS One 2013;8:e73076
- Balog J, Sasi-Szabó L, Kinross J, et al. Intraoperative tissue identification using rapid evaporative ionization mass spectrometry. Sci Transl Med 2013;5:194ra93
- National Institutes of Health Press Release. NIH announces new program in metabolomics. Available from: http://www.nih.gov/news/health/sep2012/od-19.htm [Last accessed 7 May 2014]
- Imperial College London, MCR-NIHR National Phenome Centre. Available from: http://www1.imperial.ac.uk/phenomecentre/ [Last accessed 7 May 2014]
- NHS National Institute for Health Research. MRC/NIHR Phenome Centre. http://www.nihr.ac.uk/infrastructure/Pages/PhenomeCentre.aspx [Last accessed 7 May 2014]
- Barba I, de León G, Martín E. Nuclear magnetic resonance-based metabolomics predicts exercise-induced ischemia in patients with suspected coronary artery disease. Magn Reson Med 2008;60:27-32
- Shah AA, Craig DM, Sebek JK. Metabolic profiles predict adverse events after coronary artery bypass grafting. J Thorac Cardiovasc Surg 2012;143:873-8
- Backshall A, Sharma R, Clarke SJ, et al. Pharmacometabonomic profiling as a predictor of toxicity in patients with inoperable colorectal cancer treated with capecitabine. Clin Cancer Res 2011;17:3019-28
- Jiménez B, Mirnezami R, Kinross J, et al. 1H HR-MAS NMR spectroscopy of tumor-induced local metabolic “field-effects” enables colorectal cancer staging and prognostication. J Proteome Res 2013;12:959-68
- Nicholson JK, Holmes E, Kinross JM, et al. Metabolic phenotyping in clinical and surgical environments. Nature 2012;491:384-92
- Everett JR, Loo RL, Pullen FS. Pharmacometabonomics and personalized medicine. Ann Clin Biochem 2013;50:523-45
- Yap IK, Brown IJ, Chan Q, et al. Metabolome-wide association study identifies multiple biomarkers that discriminate north and south Chinese populations at differing risks of cardiovascular disease: INTERMAP study. J Proteome Res 2010;9:6647-54
