Abstract
Introduction
Circular RNAs (circRNAs) function as efficient microRNA sponges with gene-regulatory potential and are promising cancer biomarkers. In this study, we used the Arraystar Human circRNA Array to construct a genome-wide circRNA profile of esophageal squamous cell cancer (ESCC) and breast cancer (BC).
Patients and methods
Expression levels between cancer lesions and adjacent normal-appearing tissues were compared. We observed 469 upregulated circRNAs and 275 downregulated circRNAs in ESCC. Hsa_circRNA_103670 was upregulated 20.3-fold, while hsa_circRNA_030162 was downregulated 12.1-fold. For BC, 715 circRNAs were upregulated, and 440 circRNAs were downregulated. Hsa_circRNA_005230 was upregulated 12.2-fold, while hsa_circRNA_406225 was downregulated 12.4-fold.
Results
When we set the criteria as fold change in expression ≥2 between cancer and adjacent normal-appearing tissue with a P-value <0.01, there were 22 common circRNAs (11 upregulated and 11 downregulated) in relation to both ESCC and BC. Gene ontology and the Kyoto encyclopedia of genes and genomes analyses showed that these circRNAs were involved in the tumorigenesis of human cancers.
Conclusion
Our study revealed that circRNAs are promising candidates as valuable biomarkers for ESCC and BC, although relevant research is still in its infancy and the functional role of specific circRNAs in tumorigenesis is just starting to be elucidated.
Introduction
Cancer is the leading cause of death in developed countries, and its prevalence is increasing in developing countries as well, which imposes a heavy burden to society at large. According to a GLOBOCAN report, there were an estimated 14.1 million new cancer cases and 8.2 million cancer-related deaths in 2012 worldwide.Citation1,Citation2 Esophageal cancer ranks as the eighth most common cancer worldwide and ranks sixth in cancer-causing deaths. Histologically, the majority of esophageal cancers are divided into squamous cell carcinoma and adenocarcinoma. The high-risk areas include northern Iran, southern Russia, central Asian countries, and northern China, where esophageal squamous cell cancer (ESCC) accounts for 90% of all esophageal cancer cases.Citation3 The prognosis of ESCC is poor, and patients have only 15%–25% of 5-year survival rates after diagnosis.Citation4–Citation6
The poor prognosis and rising incidence of esophageal cancer have highlighted the need for improved detection and prediction methods that are essential prior to treatment.Citation7 Current clinically approved surveillance practices highly depend on expensive, invasive, and sampling-error-prone endoscopic procedures.Citation8 Therefore, there is a great demand to establish reliable biomarkers that could identify patients at higher risks of neoplastic progression who would hence greatly benefit from further monitoring and/or intervention. Emerging molecular tools promise to extend the diagnostic research of the endoscopist and open doors to population screening for ESCC.Citation9 An increasing evidence has shown that noncoding RNAs, such as microRNAs (miRNAs), long noncoding RNAs (lncRNAs), and circular RNAs (circRNAs), play an important role in the development and progression of multiple human cancers and could be used as prognostic factors and therapeutic targets for esophageal cancer.Citation10
CircRNAs are a class of noncoding RNA molecules that lack 5′3′ ends and a poly A tail, covalently forming closed continuous loops.Citation11,Citation12 In general, circRNAs are stable molecules, and some have functioned as efficient miRNA sponges with gene-regulatory ability.Citation13 CircRNAs, with their distinctive characteristics, have superior potential to serve as novel markers for human diseases. The list of endogenous circRNAs involved in cancer continues to grow; however, the functional relevance of the majority of endogenous circRNAs has yet to be discovered.Citation13
In one of our previous studies, we have provided evidence that circRNAs are differentially expressed in breast cancer (BC) and play an important role in carcinogenesis because they participate in cancer-related pathways and sequester miRNAs.Citation14 However, whether circRNAs are sensitive and specific biomarkers of ESCC and BC remains largely unknown. In this study, we used an Arraystar human circRNA array (Arraystar Inc., Rockvile, MD, USA) to construct a genome-wide circRNA profile of ESCC compared with altered expression in BC, with the aim of exploring the potential functions of these circRNAs as diagnostic biomarkers.
Patients and methods
Ethics statement
This study was approved by the Institutional Review Board of Nanjing Medical University, China. Written informed consent was obtained from all participants included in the study.
Patients
Patients with ESCC were enrolled from the First People’s Hospital of Yancheng in October 2016. Tissues in the cancer lesion and adjacent normal-appearing esophagus were collected from patients who underwent surgical resection and met the following criteria: 1) a pathologic diagnosis of ESCC; 2) no previous history of cancer; 3) HIV negative; and 4) no history of radiotherapy or chemotherapy before specimen collection. The recruitment process of patients with BC was detailed in a previously published paper.Citation14 Tissue samples were placed in RNA storage solution (Shanghai Biotechnology Corporation, Shanghai, China) and stored at −80°C until use. Tumor stages were determined according to the tumor-node-metastasis (TNM) staging criteria.Citation15
CircRNA microarray
The RNA was isolated with an RNeasy mini kit (Qiagen, Hilden, Germany) and analyzed using an 8*15K Arraystar human circRNA microarray V2 (Catalog No: AS-CR-H-V2.0). Sample preparation and array hybridization followed the manufacturer’s protocol.
Bioinformatics and data analysis
We performed gene ontology (GO) analysis (http://www.geneontology.org)Citation16 to construct meaningful annotation of genes in any organism covering domains of biological processes, cellular components, and molecular functions. The log10(P-value) denotes the significance of GO term enrichment correlated to the genes producing differentially expressed circRNAs. Kyoto encyclopedia of genes and genomes (KEGG) analysis (http://www.genome.jp/kegg)Citation17 was carried out to confirm the pathway clusters covering molecular interaction and reaction networks in genes producing differentially expressed circRNAs.
Statistical analysis
The fold change in circRNA expression was calculated by comparing expression levels between cancer lesions and control tissues. Student’s t-test was used to estimate the significance of the difference between the two groups. CircRNAs with fold change ≥2 and P<0.05 were considered to be statistically significant. We used the filter criteria of fold change ≥2 and P<0.01 to screen for common differentially expressed circRNAs shared by both ESCC and BC. The false discovery rate was used to adjust for P-values in microarray analysis. Agilent feature extraction software (version 11.0.1.1, Agilent, Santa Clara, CA, USA) was used to analyze the acquired array images. R software version 3.3.1 (https://www.r-project.org/)Citation18 was used to perform quantile normalization and for GO and KEGG analysis.
Results
Differentially expressed circRNAs
Seven paired ESCC samples and four paired BC samples were collected for the Arraystar human circRNA array. The characteristics of the patients are shown in and . When we set the criteria as fold change ≥2.0 and P<0.05 between cancer lesions and adjacent normal-appearing tissues, there were 744 differentially expressed circRNAs in ESCC and 1155 differentially expressed circRNAs in BC. For ESCC, 469 circRNAs were upregulated, and 275 circRNAs were downregulated. The top 10 upregulated and top 10 downregulated circRNAs for ESCC are listed in . Hsa_circRNA_103670 was upregulated 20.3-fold, while hsa_circRNA_030162 was downregulated 12.1-fold. For BC, 715 circRNAs were upregulated, and 440 circRNAs were downregulated. The top 10 upregulated and top 10 downregulated circRNAs of BC are listed in . Hsa_circRNA_005230 was upregulated 12.2-fold, while hsa_circRNA_406225 was downregulated 12.4-fold.
Table 1 The top 10 upregulated and top 10 downregulated circRNAs for ESCC
Table 2 The top 10 upregulated and top 10 downregulated circRNAs for BC
illustrates the hierarchical clustering analysis showing the distinct circRNA expression profiling in ESCC () and BC (). shows the scatter plots demonstrating the heterogeneity of circRNA expression of ESCC cancer lesions () and BC cancer lesions () with their adjacent normal-appearing tissues. The expression of the circRNAs above the top reference line and below the bottom reference line changed by >2-fold. Volcano plots were used to visualize the significantly differentially expressed circRNAs for ESCC () and BC ().
Figure 1 Heat map showing the expression profiles of circRNAs in ESCC (A) and BC (B).
Notes: The expression values are represented by the color scale. The intensity increases from green (relatively lower expression) to red (relatively higher expression). Each column represents one tissue sample, and each row represents a single circRNA. C1–C7 represent seven ESCC specimens collected from ESCC patients. N1–N7 represent paired adjacent normal-appearing tissues. C8–C11 represent four BC specimens collected from BC patients. N8–N11 represent paired adjacent normal-appearing tissues.
Abbreviations: ESCC, esophageal squamous cell cancer; BC, breast cancer.
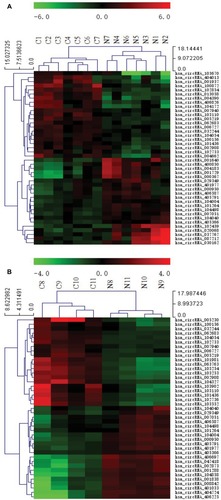
Figure 2 Scatter plots demonstrating the heterogeneity of ESCC lesions (A) and BC lesions (B) with their adjacent normal-appearing tissues.
Notes: The values of the X and Y axes represent the averaged normalized signal values of the group (log2-scaled). The green line stands for 2-fold changes. The expression of the circRNAs above the top green line and below the bottom green line indicate changes by >2-fold between the two groups of samples.
Abbreviations: ESCC, esophageal squamous cell cancer; BC, breast cancer.
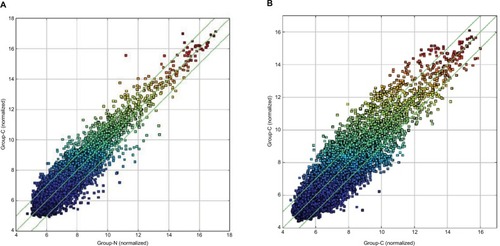
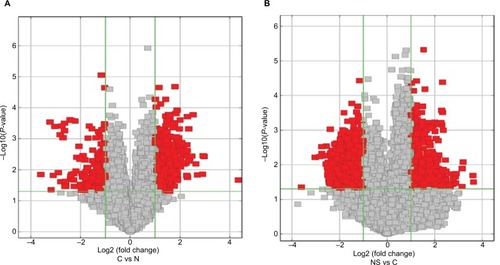
Figure 3 Volcano plots visualizing differential expression of ESCC lesions (A) and BC lesions (B) with adjacent normal tissues.
Notes: The vertical lines correspond to 2.0-fold up- and downregulation (log2 ratio), and the horizontal line represents a P-value of 0.05. The red points in the plot represent the differentially expressed circRNAs with statistical significance (fold change >2 and P<0.05), the gray points represent the remaining circRNAs (fold change <2 or P>0.05).
Abbreviations: ESCC, esophageal squamous cell cancer; BC, breast cancer.
When we set the criteria as the fold change in expression ≥2 between cancer and adjacent normal-appearing tissue and P<0.01, there were 22 common circRNAs in relation to both ESCC and BC. Among them, 11 were upregulated and 11 were downregulated ().
Table 3 Common circRNAs in relation to both ESCC and BC
GO enrichment and KEGG analysis
We further used GO analysis to explore the roles of these dysregulated circRNAs. The top 10 upregulated circRNAs in ESCC were related to positive regulation of cellular processes, biological processes, and macromolecule metabolic processes (Figure S1A), whereas the top 10 downregulated circRNAs in ESCC were related to positive regulation of cellular metabolic processes, RNA metabolic processes, and cellular processes (Figure S1B). In BC, the top 10 upregulated circRNAs were associated with negative regulation of cellular processes, biological processes, and macromolecule metabolic processes (Figure S1C), while the top 10 downregulated circRNAs were associated with the regulation of macromolecule metabolic processes, metabolic processes, and primary metabolic processes (Figure S1D). We further performed GO analysis on 22 dysregulated circRNAs shared in both ESCC and BC. Our data revealed that 11 shared upregulated circRNAs were involved in the regulation of signaling, cellular processes, and biological processes (Figure S1E), while 11 shared downregulated circRNAs were involved in the regulation of signaling and developmental processes (Figure S1F).
In KEGG pathway analysis, for the top 10 upregulated circRNAs in ESCC, HTLV-I infection, cancer, and cGMP-PKG signaling were the top three pathways (Figure S2A). For the top 10 downregulated circRNAs in ESCC, Wnt signaling, ubiquitin-mediated proteolysis, and HTLV-I infection were the top three pathways (Figure S2B). Moreover, the top three KEGG pathways for the top 10 upregulated circRNAs in BC were MAPK signaling, neurotrophin signaling, and ubiquitin-mediated proteolysis (Figure S2C), and the top three KEGG pathways for the top 10 downregulated circRNAs in BC were MAPK signaling, synaptic vesicle cycle, and hippo signaling pathway-multiple species (Figure S2D). For the 11 upregulated circRNAs shared in ESCC and BC, the three most enriched pathways in the KEGG analysis were endocytosis, thyroid hormone signaling pathway, and proteoglycans in cancer (Figure S2E). The three most enriched pathways for the 11 downregulated circRNAs shared in ESCC and BC were proteoglycans in cancer, endocytosis, and pathways in cancer (Figure S2F).
Discussion
CircRNAs are RNA molecules in which a covalent linkage typically contains the exon sequences and a splice between an upstream 3′ splice site and a downstream 5′ splice site.Citation19,Citation20 CircRNAs have been hypothesized to function as miRNA sponges to offset the impact of miRNAs. In recent years, more roles of circRNAs such as sequestering proteins or regulating transcription have been discovered.Citation21 Studies have shown that abnormal circRNAs are involved in tumorigenesis and disease progression.Citation22–Citation25 However, whether they are general cancer biomarkers or cancer specific biomarkers is not clear. To distinguish the differentially expressed circRNAs between ESCC and BC, we performed a comparative study.
For ESCC, hsa_circRNA_103670 was the most upregulated circRNA and aligned with CNOT6L (CCR4-NOT transcription complex subunit 6 like, Gene ID: 246175), which is a main deadenylase complex regulating gene expression in eukaryotes. The most downregulated was hsa_circRNA_030162. This circRNA derives from the gene TPT1 (tumor protein translationally controlled 1, Gene ID: 7178), also called translationally controlled tumor protein (TCTP),Citation26,Citation27 which encodes a cell growth-associated protein and plays an important role in the development of various organisms. TPT1 has recently been identified as related to human skin squamous cell carcinoma and is targeted by miRNA-216b-5p in pancreatic cancer.Citation28,Citation29
For BC, the top upregulated circRNA was hsa_circRNA_005230. It was spliced from the gene DNM3OS (DNM3 opposite strand/antisense RNA, Gene ID: 10062831), which produces a noncoding RNA (ncRNA) that is involved in the formation of miR-199a2 and miR-214.Citation30 These miRNAs have been reported to be downregulated in hepatocellular cancer.Citation31 The top downregulated circRNA was hsa_circRNA_406225. It aligned with the gene TAMM41 (TAM41 mitochondrial translocator assembly and maintenance homolog, Gene ID: 132001), which is a homolog of mitochondrial translocator assembly and maintenance protein.Citation32,Citation33
We observed 22 dysregulated circRNAs shared by both ESCC and BC. Among them, hsa_circRNA_002908 (hsa_ circ_0002908) was found to be upregulated in peripheral blood mononuclear cells of tuberculosis patients compared with those of paired healthy controls and may be an alternative biomarker for pulmonary tuberculosis.Citation34 Hsa_circRNA_101436 (hsa_circ_0000567) can inhibit colorectal tumor growth and was upregulated in gefitinib-acquired resistant non-small-cell lung cancer cells.Citation35,Citation36 Hsa_circRNA_103110 (hsa_circ_0004771) has been reported to be significantly expressed in BC14 and papillary thyroid carcinoma.Citation37 Hsa_circRNA_000950 (hsa_circ_0001525) and hsa_circRNA_104040 (hsa_circ_0075410) were listed in the top 10 downregulated circRNAs from a microarray on cutaneous squamous cell carcinoma.Citation38 The expression of hsa_circRNA_401977 (hsa_circ_0000567) was found to be downregulated in early stage lung adenocarcinoma tissues and cell lines.Citation39 These findings provided evidence of the role of circRNAs in tumorigenesis.
KEGG analysis indicated that differentially expressed circRNAs were associated with multiple cancers. Many pathways are related to cancer-related functions. For example, the Wnt pathway has vital roles in the prognosis of non-small-cell lung cancer and may have future clinical value.Citation40 The Hippo signaling pathway, consisting of the critical downstream effectors YAP/TAZ, contributes to the development of cancer by resulting in an overgrowth phenotype.Citation41 The MAPK signaling pathway is an essential process for CD97 to promote gastric cancer cell proliferation and invasion.Citation42
Our study has several limitations. First, the sample size is too small to make any reasonable conclusion. These results might not even be educational for further studies by other research groups. Differentially expressed circRNAs discovered in this study need further validation. Second, the paired t-test used in this study to compare circRNA expression between the groups may not have been appropriate for the microarray experiment. Other robust hyperparameter estimations need to be used to protect against hypervariable genes and improve the power to detect differential expression.Citation43,Citation44 Third, the study on the role of circRNAs in human cancers is still in its infancy, and a common standard for reporting and naming circRNAs is lacking. Fourth, we used tissue samples to detect circRNA. More easily acquired and noninvasive clinical samples, such as blood, urine, or saliva, should be explored as sources of biomarkers in future research.
Conclusion
In conclusion, circRNAs are promising candidates as valuable biomarkers for ESCC and BC, although circRNA research is still in its infancy, and the functional role of circRNAs in tumorigenesis is just starting to be elucidated.
Acknowledgments
The present study was supported by the National Natural Science Foundation of China (81673249), the Social Development Project in Jiangsu Province (BE2015694), the Scientific Research Innovation Project for Graduate Students in Jiangsu Province (KYCX17_1293), and the Priority Academic Program Development of Jiangsu Higher Education Institutions. The funding agencies had no role in the study design, data collection, analysis, decision to publish, or preparation of the manuscript.
Supplementary materials
Figure S1 GO enrichment analysis for differentially expressed genes.
Notes: (A) GO annotation of the top 10 upregulated circRNAs in ESCC; (B) GO annotation of the top 10 downregulated circRNAs in ESCC; (C) GO annotation of the top 10 upregulated circRNAs in BC; (D) GO annotation of the top 10 downregulated circRNAs in BC; (E) GO annotation of 11 shared upregulated circRNAs in ESCC and BC; (F) GO annotation of 11 shared downregulated circRNAs in ESCC and BC.
Abbreviations: circRNA, circular RNA; ESCC, esophageal squamous cell cancer; BC, breast cancer.
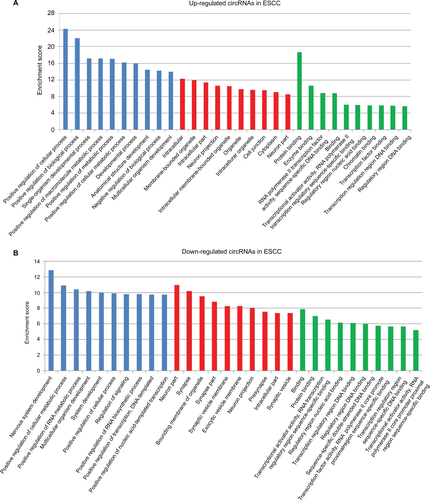
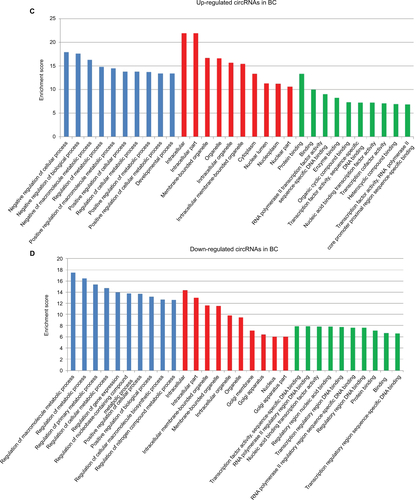
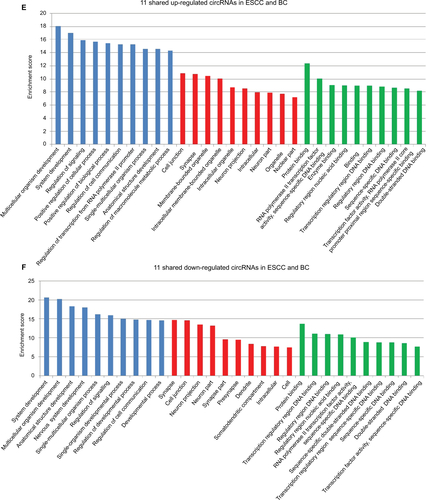
Figure S2 Kyoto encyclopedia of genes and genomes (KEGG) enrichment analysis of differentially expressed genes.
Notes: (A) Pathways corresponding to the top 10 upregulated circRNAs in ESCC; (B) Pathways corresponding to the top 10 downregulated circRNAs in ESCC; (C) Pathways corresponding to the top 10 upregulated circRNAs in BC; (D) Pathways corresponding to the top 10 downregulated circRNAs in BC; (E) Pathways corresponding to 11 shared upregulated circRNAs in ESCC and BC; (F) Pathways corresponding to 11 shared downregulated circRNAs in ESCC and BC.
Abbreviations: circRNA, circular RNA; ESCC, esophageal squamous cell cancer; BC, breast cancer.
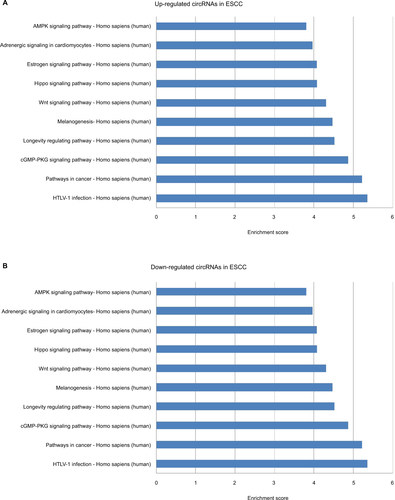
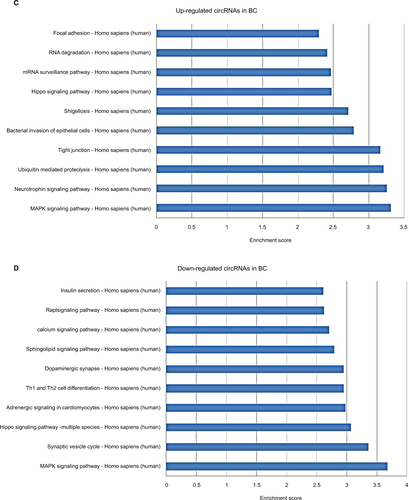
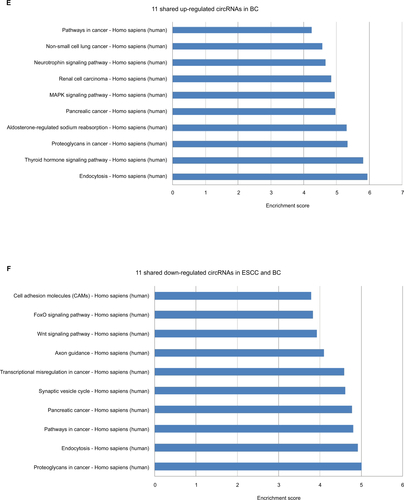
Table S1 General information of ESCC patients for microarray test
Table S2 General information of BC patients for microarray test
Author contributions
PS and JMW conceived, initiated, and led the study. PS, JS, JMW, and HX analyzed the data with input from all the authors. PS and JMW prepared the manuscript. All authors contributed to data analysis, drafted and revised the paper, and agreed to be accountable for all aspects of the work. All authors reviewed and approved the manuscript.
Disclosure
The authors report no conflicts of interest in this work.
References
- BrownJCWinters-StoneKLeeASchmitzKHCancer, physical activity, and exerciseCompr Physiol2012242775280923720265
- TorreLABrayFSiegelRLFerlayJLortet-TieulentJJemalAGlobal cancer statistics, 2012CA Cancer J Clin20156528710825651787
- Recio-BoilesABabikerHMCancer, esophagealStatpearlsTreasure Island, FL2018
- Lao-SirieixPFitzgeraldRCScreening for oesophageal cancerNat Rev Clin Oncol20129527828722430857
- OhashiSMiyamotoSKikuchiOGotoTAmanumaYMutoMRecent advances from basic and clinical studies of esophageal squamous cell carcinomaGastroenterology201514971700171526376349
- WangBSongHJiangHFuYDingXZhouCEarly diagnostic potential of APC hypermethylation in esophageal cancerCancer Manag Res20181018119829440928
- HuangFLYuSJEsophageal cancer: risk factors, genetic association, and treatmentAsian J Surg201841321021527986415
- PusungMZekiSFitzgeraldRGenomics of esophageal cancer and biomarkers for early detectionAdv Exp Med Biol201690823726327573775
- AndersonBWAhlquistDAMolecular detection of gastrointestinal neoplasia: innovations in early detection and screeningGastroenterol Clin North Am201645352954227546847
- HouXWenJRenZZhangGNon-coding RNAs: new biomarkers and therapeutic targets for esophageal cancerOncotarget2017826435714357828388588
- ZhangYLiangWZhangPCircular RNAs: emerging cancer biomarkers and targetsJ Exp Clin Cancer Res201736115229096676
- QuSYangXLiXCircular RNA: a new star of noncoding RNAsCancer Lett2015365214114826052092
- KristensenLSHansenTBVenoMTKjemsJCircular RNAs in cancer: opportunities and challenges in the fieldOncogene201837555556528991235
- LuLSunJShiPIdentification of circular RNAs as a promising new class of diagnostic biomarkers for human breast cancerOncotarget2017827440964410728484086
- EdgeSBComptonCCThe American Joint Committee on Cancer: the 7th edition of the AJCC cancer staging manual and the future of TNMAnn Surg Oncol20101761471147420180029
- Gene Ontology Consortium Available from: http://www.geneontology.org/Accessed June 21, 2018
- Kanehisa Laboratories Available from: http://www.genome.jp/keggAccessed June 21, 2018
- R Foundation for Statistical Computing Available from: https://www.r-project.org/Accessed June 21, 2018
- SalzmanJCircular RNA expression: its potential regulation and functionTrends Genet201632530931627050930
- SzaboLSalzmanJDetecting circular RNAs: bioinformatic and experimental challengesNat Rev Genet2016171167969227739534
- Cortes-LopezMMiuraPEmerging functions of circular RNAsYale J Biol Med201689452753728018143
- HansenTBKjemsJDamgaardCKCircular RNA and miR-7 in cancerCancer Res201373185609561224014594
- HeJXieQXuHLiJLiYCircular RNAs and cancerCancer Lett201739613814428342987
- XiaWQiuMChenRCircular RNA has_circ_0067934 is upregulated in esophageal squamous cell carcinoma and promoted proliferationSci Rep201663557627752108
- ZhouJZhangWWPengFSunJYHeZYWuSGDownregulation of hsa_circ_0011946 suppresses the migration and invasion of the breast cancer cell line MCF-7 by targeting RFC3Cancer Manag Res20181053554429593432
- AmsonRPeceSMarineJCDi FiorePPTelermanATPT1/TCTP-regulated pathways in phenotypic reprogrammingTrends Cell Biol2013231374623122550
- ChenWWangHTaoSTumor protein translationally controlled 1 is a p53 target gene that promotes cell survivalCell Cycle201312142321232824067374
- WuDGuoZMinWUpregulation of TCTP expression in human skin squamous cell carcinoma increases tumor cell viability through anti-apoptotic action of the proteinExp Ther Med20123343744222969908
- YouYTanJGongYMicroRNA-216b-5p Functions as a tumor-suppressive RNA by targeting TPT1 in pancreatic cancer cellsJ Cancer20178142854286528928875
- LiCZhouYLobergATaharaSMMalikPKalraVKActivated transcription factor 3 in association with histone deacetylase 6 negatively regulates microRNA 199a2 transcription by chromatin remodeling and reduces endothelin-1 expressionMol Cell Biol201636222838285427573019
- DuanQWangXGongWER stress negatively modulates the expression of the miR-199a/214 cluster to regulates tumor survival and progression in human hepatocellular cancerPLoS One201272e3151822359598
- HicksCSitthi-AmornJDouglasJMolecular analysis of central nervous system disease spectrum in childhood acute lymphoblastic leukemiaClin Med Insights Oncol20161051526997880
- KariminejadANafissiSNilipoorYIntellectual disability, muscle weakness and characteristic face in three siblings: a newly described recessive syndrome mapping to 3p24.3-p25.3Am J Med Genet A2015167A112508251526192890
- QianZLiuHLiMPotential diagnostic power of blood circular RNA expression in active pulmonary tuberculosisEBioMedicine201827182629248507
- WangJLiXLuLHeLHuHXuZCircular RNA hsa_circ_0000567 can be used as a promising diagnostic biomarker for human colorectal cancerJ Clin Lab Anal Epub2018115
- DaiYHuangYHuangJWenCZhouHWuLDifferential expression of circular RNAs in gefitinib-acquired resistant non-small cell lung cancer cellsTUMOR2017371111281135
- PengNShiLZhangQHuYWangNYeHMicroarray profiling of circular RNAs in human papillary thyroid carcinomaPLoS One2017123e017028728288173
- SandMBecharaFGGambichlerTCircular RNA expression in cutaneous squamous cell carcinomaJ Dermatol Sci201683321021827298156
- ZhaoJLiLWangQHanHZhanQXuMCircRNA Expression profile in early-stage lung adenocarcinoma patientsCell Physiol Biochem20174462138214629241190
- StewartDJWnt signaling pathway in non-small cell lung cancerJ Natl Cancer Inst20141061djt35624309006
- MoJSParkHWGuanKLThe Hippo signaling pathway in stem cell biology and cancerEMBO Rep201415664265624825474
- LiCLiuDRLiGGCD97 promotes gastric cancer cell proliferation and invasion through exosome-mediated MAPK signaling pathwayWorld J Gastroenterol201521206215622826034356
- RitchieMEPhipsonBWuDLimma powers differential expression analyses for RNA-sequencing and microarray studiesNucleic Acids Res2015437e4725605792
- PhipsonBLeeSMajewskiIJAlexanderWSSmythGKRobust hyperparameter estimation protects against hypervariable genes and improves power to detect differential expressionAnn Appl Stat201610294696328367255
