Abstract
Introduction
Small cell lung cancer (SCLC), an aggressive subtype of lung cancer characterized by the development of neuroendocrine tumors, is prone to distant metastasis, resistant to platinum-based drugs and has a poor prognosis. The development of next-generation sequencing technology (NGS) has led to the identification of many genetic alterations in SCLC. Few druggable targeted molecules can be used in clinical practice. Currently, NGS is widely employed in routine clinical practice of non-small cell lung cancer to assist in therapeutic options and prognosis evaluation. This study aims to investigate genes involved in small cell lung cancer (SCLC), their occurrence and their significance in clinical events.
Methods
Tumor tissue specimens from 18 Chinese SCLC patients were collected through a 520 cancer‐related genes panel for next-generation sequencing. First, the association between sequence results and clinical outcomes was examined. Subsequently, data on clinical pathology and sequencing results were analyzed.
Results
The Kaplan–Meier curve displayed a significant reduction in PFS for SCLC patients with LRP1B or MAP3K13 mutations. Overall survival (OS) of SCLC patients with MSH6 mutation was significantly higher than those with SPEN mutation.
Conclusion
Next-generation sequencing demonstrates that the genetic landscape of SCLC. Mutation status of LRP1B, MAP3K13, MSH6 and SPEN has prognostic significance, which might be potential therapeutic targets. We found possible genes and related signaling pathways that affect metastasis. These results can improve our understanding of the mutation characteristics of SCLC and identify potential biomarkers to guide targeted therapies.
Introduction
Cancer has been one of the leading causes of human death. According to estimates, more than 220 thousand new cases of lung cancer will occur in the United States in 2020.Citation1 In general, lung cancer is subdivided into two categories, of which small cell lung cancer (SCLC) represents about 13–15%. At diagnosis, about 80% of patients with SCLC are in an advanced stage and cannot undergo surgery.Citation2
SCLC is a highly heterogeneous malignant neuroendocrine tumor.Citation3 Small cell transformation has been demonstrated to be one of the ways in which non-small cell lung cancer (NSCLC) develops resistance to epidermal growth factor receptor tyrosine kinase inhibitors (EGFR-TKIs), leading to a poor prognosis.Citation4 Although TP53 and RB1 are tumor suppressor genes with high incidence, few genetic driver events have been reported.Citation5 Recent evidence reveals that immune checkpoint inhibitors (ICIs) and specifically programmed death ligand 1 inhibitors combined with standard platinum/etoposide can enhance progression-free survival with minimal adverse effects in patients with extensive SCLC.Citation6,Citation7 Furthermore, recent evidence shows that co-stimulatory B7-H3 may act as an independent prognostic indicator for SCLC cases, which might provide a theoretical basis for subsequent research targeting B7-H3.Citation8,Citation9
FOXM1 is crucial for SCLC tumorigenesis and is associated with a poor prognosis. After standard chemotherapy, patients with high FOXM1 expression had shorter progression-free survival compared to those with low FOXM1 expression (3.90 vs 8.69 months).Citation10 In vitro experiments and experiments on xenograft (PDX) models derived from SCLC patient-derived xenograft reveal that pharmacological inhibition of DHODH can suppress the cell viability.Citation11 Therefore, in SCLC treatment, DHODH has been a promising target.
SCLC is prone to develop chemoresistance due to its intratumoral heterogeneity.Citation12 Despite recent advances, SCLC remains the most lethal lung cancer with limited therapeutic options. Studies report that patients with SCLC have a poor prognosis, except for a minimal number of early-stage cases.Citation13
Accumulated studies demonstrate that SCLC is a highly heterogeneous tumor, although fewer genetic colonies have been reported comparison with NSCLC. Transcriptomic analyses reveal four different molecular subtypes, including SCLC-A driven by the transcription factor ASCL1, SCLC-N driven by NEUROD1, SCLC-Y by YAP1 and SCLC-P by POU2F3,Citation14 allowing for more targeted therapeutic approaches. Subsequent studies confirmed that a unique YAP1 subtype did not exist, and an inflamed subtype of SCLC (SCLC-I) was recently proposed to replace YAP1 subtype.Citation15 Although SCLC-I subtype experienced the greatest benefit from immune checkpoint inhibitor therapy, the clinical significance of this molecular subtyping in guiding treatment and estimating prognosis remains limited to other standard SCLC treatments.
The development of next-generation sequencing technology (NGS) has led to the identification of many genetic alterations in SCLC, including TP53 and RB1 inactivation and frequent chromosomal abnormalities (deletion 3p).Citation16 In addition to the inactivated Notch pathway, MYC family amplification has a high incidence.Citation17 Few druggable targeted molecules can be used in clinical practice.Citation18 Currently, NGS is widely employed in routine clinical practice of non-small cell lung cancer to assist in therapeutic options and prognosis evaluation. Therefore, it is essential to investigate the genetic characteristics of SCLC and their clinical implications.
In this study, 18 cases of pathologically proven SCLC cases were sequenced using a panel of 520 cancer-related genes. The average median sequencing depth of the samples was 1260x. The average median q30 ratio of the samples was 91%, and all samples are qualified. Analyzing the sequencing results and pathological data revealed some genetic variants with distinctive characteristics.
SCLC presents a distinctive mutation spectrum compared to adenocarcinoma. In our cohorts, results reveal 72% co‐occurring TP53/RB1 mutations. Furthermore, in the samples of small cell lung cancer, the core 8 gene was not detected. In SCLC, the frequency of gene mutations is significantly lower than in adenocarcinoma. The mutation frequency of Rb1, MSH6, KDR, IL7R, ATRX, EPHB1, PDGFRA and KIT in SCLC is markedly higher than those in lung squamous cell carcinoma. Due to the small sample size, it might be affected by some random factors.
Materials and Methods
Patients and Sample Collection
Patients diagnosed with SCLC at Henan Cancer Hospital, Affiliated Cancer Hospital of Zhengzhou University, were enrolled in this study after obtaining informed consent. Biopsy specimens of advanced stage SCLC were obtained from the Department of Pathology Affiliated Cancer Hospital of Zhengzhou University. The study protocols were approved by the institutional ethics review board (Ethics Review Committee, Zhengzhou University). While collecting specimens for analysis, we complied with the declaration of Helsinki (2013 Edition) and relevant regulations. Additionally, the clinical characteristics were recovered in this research.
DNA Isolation and Capture-Based Targeted DNA Sequencing
DNA extraction and targeted sequencing were performed in Burning Rock Biotech as described in the previous protocol.Citation19,Citation20 In brief, formalin-fixed, paraffin-embedded (FFPE) specimens were used for DNA extraction through QIAamp DNA kit (Qiagen, Germany). Subsequently, DNA purification, hybridization and amplification were performed. Target capture was conducted through a commercial 520 genes panel (OncoScreen Plus). The quality of fragments was determined using a Bioanalyzer 2100 (Agilent Technologies, USA). Finally, the samples were sequenced via the platform of nextseq 500 (Illumina, Inc., USA).
Sequence Data Analysis and Tumor Mutation Burden (TMB) Calculation
The reference genome Hg19 was used through Burrows-Wheeler Aligner version 0.7.10.Citation21 Subsequently, sequence alignment and variant calling were performed through programs including varscan version 2.4.3 and the genome analysis tool kit version 3.2.Citation22 The genes variants were annotated with ANNOVAR.Citation23 The structural variations (SVs) were analyzed using Factera version 1.4.3.Citation24
It should count as mutations that non-synonymous single nucleotide variants (SNVs) and indels in the coding sequences and two adjacent base pairs around these regions, while hot mutation, copy number variations (CNVs), structural variations (SVs), and SNPs were excluded.
The size of coding sequences required to estimate TMB is 1.003 MB in the 520 gene panel. The MB per patient was calculated using the following formula.
Follow-Up and Statistical Analysis
lists the clinical and pathological information of SCLC patients. Prospective follow-up was conducted through routine hospital visits or telephone calls. Once every three months, trained medical staff made telephone calls to patients or their family members until death or the last follow-up. The follow-up data of this cohort is included in the Supplementary Materials. Overall survival (OS) was estimated from the day of SCLC diagnosis to the day of death from any cause and was analyzed using Kaplan–Meier estimates and Log rank test. The correlations between categorical variables were calculated using Chi-squared and Fisher’s exact tests. Statistical analyses were performed using SPSS23.0, and P ≤ 0.05 was considered statistically significant.
Table 1 Clinicopathological Characteristics of the 18 SCLC Patients
Results
Next-Generation Sequencing Shows the Genetic Landscape of SCLC
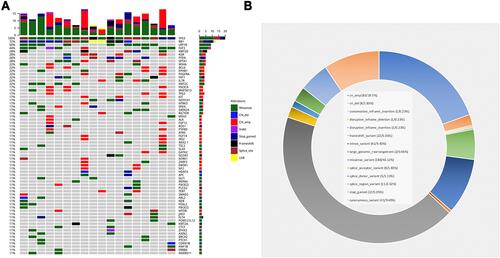
Capture-based targeted sequencing was performed by Burning Rock Biotech, Guangzhou, China. The average median sequencing depth of the samples was 1260x, consistent with expectations. The average median q30 ratio of the samples was 91%, and all samples were qualified (Supplementary Figure 1). The deletion of the tumor suppressor genes TP53 and RB1 has long been recognized as a common mutation in SCLC. Out of 18, 13 patients showed a co-mutation of TP53 and RB1, reaching a mutation rate of 72%, and the remaining five patients were found with wild-type RB1 as displayed in . Other frequent mutant genes are LRP1B, FAT3, KMT2D, KDR, PTEN, SPTA1, MSH6, Bcl6, EPHB1, etc. illustrates the distribution of identified mutations in SCLC.
Genetic Alteration in SCLC Specimen Was Different from That in Adenocarcinoma and Squamous Cell Carcinoma
The genetic alteration observed in our cohort was in accordance with the SCLC database of Burning Rock Biotech (RS_SCLC) (). includes genes detected in at least three samples. The gene mutation frequency in this cohort did not differ significantly from that in RS_SCLC database.
Figure 2 (A) Genetic alteration in our cohort consistent with Burning Rock Biotech SCLC database (RS_SCLC). (B) Genetic differences between SCLC and non-small cell lung cancer (NSCLC) (C). Single nucleotide variation analysis. *in (B and C) represents statistically significant differences (p < 0.05). **in (B and C) represents statistically significant differences (0.01<p < 0.05). *** in (B and C) represents statistically significant differences (p < 0.01). **** in (B and C) represents statistically significant differences (p < 0.001).
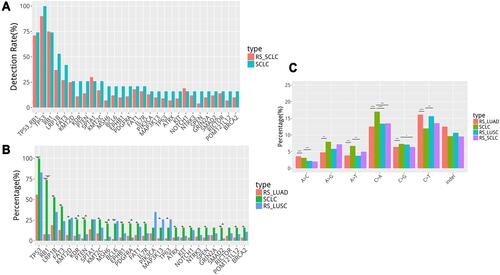
The genetic differences between SCLC and non-small cell lung cancer were further analyzed () to explore the molecular genetic characteristics of SCLC. In comparison with NSCLC (adenocarcinoma and squamous cell carcinoma), the mutation spectrum of SCLC differed significantly from the adenocarcinoma database of Burning Rock Biotech (RS_LUAD). Neither the core eight gene nor the frequent mutations in SCLC were detected in adenocarcinomas. Compared to Burning Rock Biotech’s database of squamous cell carcinoma (RS_LUSC), the mutation frequency of genes RB1, MSH6, KDR, IL7R, ATRX, EPHB1, PDGFRA, and KIT was significantly higher. Considering the small sample size, other factors might affect it.
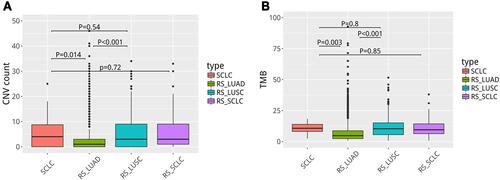
Furthermore, a single nucleotide variation analysis revealed that C > A mutations were more frequent in SCLC, whereas C > T mutations were significantly lower, as illustrated in . Moreover, we analyzed the gene copy number variation (CNV) and tumor mutation burden (TMB) in our cohorts (). There was no significant difference in gene copy number variation (CNV) between the cohorts of SCLC group and RS_LUSC group (P = 0.54). In contrast, there was a significant difference between SCLC and RS_LUAD groups (P = 0.014). In terms of tumor mutation burden (TMB), there was no significant difference between cohorts of SCLC and RS_LUSC groups (P = 0.80), whereas there was a considerable difference between SCLC and RS_LUAD groups (P = 0.003). In general, SCLC showed higher copy number amplification and TMB compared to lung adenocarcinomas.
Part Mutant Genes are Related to Clinical Factors to Some Extent
In the final analysis, we examined the association between gene mutation and clinicopathological data, including gender, smoking, specimen origins, metastasis site, progression-free survival and overall survival (). The criteria for gene inclusion were as follows: a. genes detected in at least three samples, b. clinical factors with at least three statistics, c. baseline sample data were included for analysis. The results revealed a significant correlation between several mutant genes and clinical factors. The Kaplan–Meier curve showed that patients with LRP1B () or MAP3K13 () mutation exhibited significantly shortened PFS.
Figure 4 The correlation between mutant gene and clinicopathological data including gender, smoking, specimen origins, metastasis site, and progression free survival and overall survival. Red indicates statistically significant differences (p < 0.05).
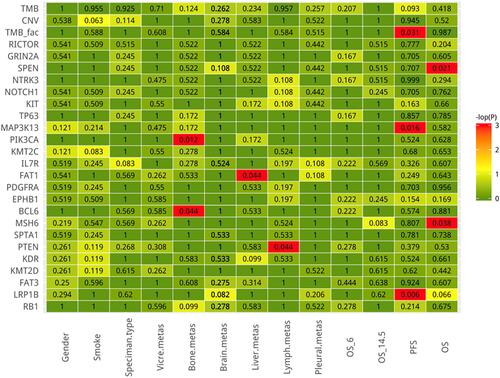
Figure 5 Kaplan–Meier curve indicated that SCLC patients with LRP1B (A) or MAP3K13 (B) mutation had significantly shorter PFS. SCLC patients with MSH6 mutation had significantly longer OS (C) while OS was significantly shorter in patients with SPEN mutation (D).
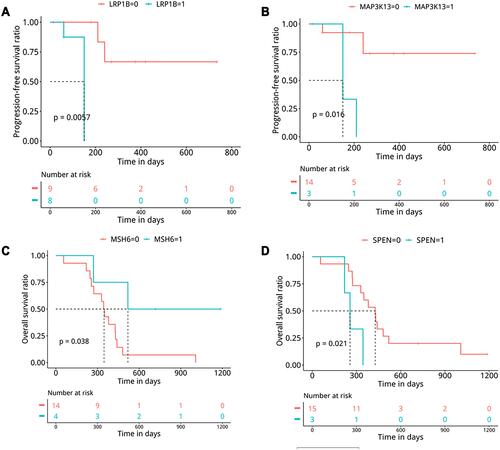
Moreover, SCLC patients with MSH6 mutation had significantly longer OS (), while the OS decreased significantly in patients with SPEN mutation (). In conclusion, MSH6 mutations are associated with a better prognosis than SPEN mutations. The results of this study may help to diagnose and treat Chinese SCLC patients.
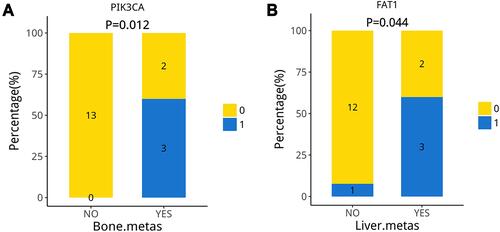
The proportion of mutant-PIK3CA was higher in patients with bone metastases (P = 0.012), and the proportion of mutant-FAT1 was higher in patients with liver metastases (P = 0.044), as displayed in . Of 18, five patients eventually developed bone metastases; three were detected with PIK3CA mutation, while five of the patients with liver metastases were detected with FAT1 mutation. Based on these results, it can be concluded that related signaling pathways could play a role in regulating organ-specific metastasis.
Figure 6 The proportion of mutant-PIK3CA is higher in patients with bone metastases (A), the proportion of mutant-FAT1 is higher in patients with liver metastases (B).
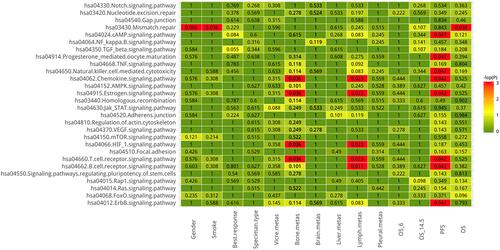
The signaling pathways based on KEGG involved in mutant genes were analyzed (). A pathway is deemed mutant when at least one sample has a mutation in it. Signaling pathways with statistical correlations are listed in , which requires further investigation. The results indicated that HIF1 signaling pathway, estrogen signaling pathway, chemokine signaling pathway and T cell receptor signaling pathway contributed to bone and lymph node metastasis. Additionally, signaling pathways linked to PFS and OS were identified, and these findings may provide genetic explanations for clinicopathological features.
Table 2 The Clinical Factors are Associated with Signaling Pathway Based on KEGG Pathway to Some Extent
Figure 7 The correlation between signaling pathways involved in mutant genes (based on KEGG pathway) and clinicopathological data including gender, smoking, specimen origins, metastasis site, and progression free survival and overall survival. Red indicates statistically significant differences (p < 0.05).
Discussion
SCLC in the early stages is sensitive to radiotherapy and chemotherapy. In most patients, satisfactory treatment results cannot be achieved due to the disease’s early progression, recurrence, and chemotherapy resistance. Despite recent advances in SCLC, even on standard platinum-containing two-drug chemotherapy combined with immunization, survival did not exceed two years.Citation25 Nitin Roper et al disclosed that different transcriptional SCLC subtypes could experience clinical benefit to ICIs, as was Notch signaling activation, which might provide the means for more effective application of ICIs in SCLC.Citation26 Chemo-immunotherapy has become the preferred initial treatment for advanced SCLC, but only a small subset of SCLC patients’ benefits from ICIs, and there is an urgent need for biomarkers with efficient prediction.Citation27
SCLC has long been considered to have higher genomic instability than other types of cancer.Citation28 The inactivation of TP53 and RB1 occurs early in the course of SCLC.Citation29 Inactivation of RB1 allows cells to enter the cell cycle, while TP53 loss can prevent cell cycle arrest and apoptosis. Amplification of Myc family members may enhance cell proliferation, while ASCL1 promotes neuroendocrine fate.Citation30
Furthermore, studies have found that histone modification is highly prevalent in SCLC. Overall, these genetic alterations result in replication stress in SCLC, providing a potential direction for gene intervention therapy. Therefore, the study examined various gene mutations associated with SCLC and the effects of those mutations on the pathogenesis of clinical events and their prognosis. In the cbioportal site, we observed whole-genome sequencing of 120 small cell lung cancer (SCLC) tumour samples and matched normal materials.Citation31 The tumour samples in this study were enriched for earlier stages, while tumour samples were all advanced stages in our study. Surgical treatment is considered for less than 5% of early patients limited to the lung parenchyma for SCLC patients. Then, therapeutic options are completely different. The differences in treatment outcomes were not comparable, but there were no significant differences in the frequency of key gene mutation (P53,RB), diagnosis age and sex ratio, indicating that our cohort is representative to some extent.
It is worth mentioning that Kaplan–Meier curve suggested that SCLC patients with LRP1B or MAP3K13 mutation had shorter PFS in our study. LRP1B (low-density lipoprotein receptor-related protein 1b) is a putative tumor suppressor. Recent evidence suggests that LRP1B might be a genetic marker for immune checkpoint inhibitors (ICI) in multiple types of cancers.Citation32,Citation33 However, nuclear LRP1B, which was released by the intracellular LRP1B domain and transported to the nucleus, increased the invasion activity of breast cancer cells by upregulation of NEAT1.Citation34 Further studies are required to understand the molecular significance of LRP1B in SCLC progression. MAP3K13 (encoding LZK) is an amplified driver gene in head and neck cancer cells and plays a crucial role in maintaining mutant p53 expression.Citation35 Qiang ZhangCitation36 revealed a regulatory pathway that supervised Myc protein stability via MAP3K13-TRIM25-FBXW7α signaling axis, suggesting a potential therapeutic target for cancers that over-express Myc. Considering that MAP3K13 is a targetable oncogenic kinase, its clinical application deserves further investigation in SCLC.
Researchers found that MSH6 mutations have a better prognosis compared to SPEN mutations. DNA mismatch repair genes (MMR) function in maintaining genomic stability. The dysfunction of MMR genes could lead to accumulation during the repair process of DNA and lead to gene instability and overexpression of cancer-related genes, causing tumor initiation and progression.Citation37 Nine MMR genes related to human mismatch repair have been isolated from human bacteria. MMR is a bacterial MUTS homologue and mainly forms a mismatch complex with MSH2 (MSH2-MSH6) to play a role in mismatch repair.Citation38,Citation39
SPEN family transcriptional repressor (SPEN), also known as SMART/HDAC1-related repressor protein, can regulate transcription and is essential for X chromosome inactivation.Citation40,Citation41 Multiple studies have demonstrated that SPEN can perform different functions in tumorigenesis. SPEN mutation alters a protein complex that represses the transcription of chronic lymphocytic leukemia NOTCH1 target genes.Citation42 SPEN induces miR-4652-3p expression by activating PI3K/AKT/c-JUN signaling to target HIPK2.Citation43 High levels of SPEN RNA are associated with early metastasis in two independent cohorts of 77 (HR 2.25, P = 0.03) and 170 (HR = 2.23, P = 0.004) patients with ERα-negative breast cancer.Citation44 In contrast, SPEN is a tumor-suppressor gene that may be clinically useful as a predictive biomarker of tamoxifen response in ERα-positive breast cancers.Citation45 In colon cancer, it acts as an oncogene for its pathogenesis, since it positively regulates Wnt signaling.Citation46 However, the molecular mechanism of SPEN in SCLC remains unknown. In our cohort, the OS of SCLC patients with SPEN mutation decreased significantly, indicating that SPEN could be used as a prognostic biomarker.
PI3K/Akt-mediated pathways have been implicated in many tumors. PIK3CA is a P110 catalytic subunit of phosphatidylinositol-3 kinases (PI3Ks), which encodes the PIK3CA protein, namely PI3KP110A.Citation47 PIK3CA plays a role in multiple signaling pathways and controls cellular functions.Citation48 Among them, PIK3CA encoding PI3KP110A is the only oncogenic gene with somatic mutation among the members of the PI3K family found at present. About 4/5 mutations of PIK3CA occur in two hot spots, the helix region (Exon 9) and the kinase region (Exon 20).Citation49 PIK3CA has been identified as an oncogene based on its function and genetics. In tumor cells, the pathogenic mutation of PIK3CA causes abnormal encoding of the p110α subunit that leads to continuous activation of PI3K enzyme. Furthermore, it can activate the downstream Akt, causing independent cell proliferation and enhancing the ability of cells to metastasize.Citation50,Citation51 A series of PIKK3CA mutations have been reported in various cancer lines, including SCLC (3.4–4.3%), colorectal cancer (30–40%), ovarian cancer, thyroid,Citation52,Citation53 cancer,Citation54 gastric cancer, and breast cancer (7–40%).Citation55 This gene has also been targeted in previous studies of SCLC. Triciribine, a small-molecule Akt inhibitor, was found to act as a pathway inhibitor and was more sensitive to SCLC types with PIK3CA mutations.Citation56 In this study, three samples had PIK3CA mutations, and all of them had bone metastases, resulting in a poor prognosis for the patients. Further studies are needed to understand how this gene could improve the treatment of patients with SCLC.
FAT1 is a member of the cadherin superfamily, which encodes procadherin and is frequently mutated in human cancers, especially squamous cell carcinoma.Citation57,Citation58 Similarly, our results also showed that nonsense mutations of FAT1 are common and can cause loss of gene function. Previous studies have linked FAT1 mutations to poor outcomes in cancer patients. Recurrent somatic mutation of FAT1 in multiple human cancers could result in aberrant Wnt signaling activation, thus promoting the tumor progression.Citation59,Citation60 Studies have demonstrated that FAT1 can encode a protein that binds to β-catenin and antagonizes β-catenin to enter cells to target and activate Wnt that can inhibit cell proliferation and tumor growth.Citation61 The experiments on constructed mouse models of skin squamous cell carcinoma and lung cancer found that loss of FAT1 can inhibit adhesion and promote epithelial to mesenchymal cell transformation (EMT) and thus promote tumor genesis, development and metastasis.Citation62 From an independent International Cancer Genome Consortium dataset, FAT1 mutation in oral cancer co-occurred with the top mutated genes TP53 and CASP8. Poor overall survival or progression-free survival was observed in patients with FAT1 mutation or altered HER3_pY1289, IRS1, or CAVEOLIN. Pathway analysis revealed dominant ERBB/neuregulin pathways mediated by FAT1 mutations in HNSCC.Citation63 FAT1 plays various roles in different types of cancer and can be used as an oncogene or a tumor suppressor gene. Studies have confirmed that the expression of FAT1 in liver cancer is higher and acts as a tumor protector than in nontumor liver tissue.Citation64,Citation65 Our results show that patients with FAT1 mutation were more prone to liver metastasis, an alternative therapeutic target for SCLC patients. Farago et al reported that combining olaparib and temozolomide in relapsed SCLC could significantly improve the prognosis of SCLC patients.Citation66 After that, they identified a molecular signature predictive of the response to this regimen. Olaparib is PARP inhibitor and has been proved to be effective for BRCA-mutated metastatic breast cancer and ovarian cancer.Citation67,Citation68 We found BRCA2 mutant in our SCLC cohort, which might suggest better sensitivity to olaparib.
Conclusion
Next-generation sequencing demonstrates that the genetic landscape of SCLC is different from that of adenocarcinoma and squamous cell carcinoma. Part mutant genes are linked to clinical factors to some extent. Mutation status of LRP1B, MAP3K13, MSH6 and SPEN has prognostic significance, which might be potential therapeutic targets. We found possible genes and related signaling pathways that affect metastasis. Oncologists might acquire important information to assist in therapeutic options and prognosis evaluation, and SCLC patients might benefit from NGS in clinical practice, and the underlying mechanism deserves further investigation.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Acknowledgment
We are grateful to all peer reviewers and editors for their opinions and suggestions.
Disclosure
The authors declare that they have no conflicts of interest in this work.
Additional information
Funding
References
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7–30. doi:10.3322/caac.21590
- Simon GR, Wagner H. Small cell lung cancer. Chest. 2003;123(1Suppl):259s–271s. doi:10.1378/chest.123.1_suppl.259S
- Asamura H, Kameya T, Matsuno Y, et al. Neuroendocrine neoplasms of the lung: a prognostic spectrum. J Clin Oncol. 2006;24(1):70–76. doi:10.1200/JCO.2005.04.1202
- Offin M, Chan JM, Tenet M, et al. Concurrent RB1 and TP53 alterations define a subset of EGFR-mutant lung cancers at risk for histologic transformation and inferior clinical outcomes. J Thorac Oncol. 2019;14(10):1784–1793. doi:10.1016/j.jtho.2019.06.002
- Rudin CM, Brambilla E, Faivre-Finn C, Sage J. Small-cell lung cancer. Nat Rev Dis Primers. 2021;7(1):3. doi:10.1038/s41572-020-00235-0
- Ortega-Franco A, Ackermann C, Paz-Ares L, Califano R. First-line immune checkpoint inhibitors for extensive stage small-cell lung cancer: clinical developments and future directions. ESMO Open. 2021;6(1):100003. doi:10.1016/j.esmoop.2020.100003
- Tay RY, Heigener D, Reck M, Califano R. Immune checkpoint blockade in small cell lung cancer. Lung Cancer. 2019;137:31–37. doi:10.1016/j.lungcan.2019.08.024
- Qiu MJ, Xia Q, Chen YB, et al. The expression of three negative co-stimulatory B7 family molecules in small cell lung cancer and their effect on prognosis. Front Oncol. 2021;11:600238. doi:10.3389/fonc.2021.600238
- Carvajal-Hausdorf D, Altan M, Velcheti V, et al. Expression and clinical significance of PD-L1, B7-H3, B7-H4 and TILs in human small cell lung Cancer (SCLC). J Immunother Cancer. 2019;7(1):65. doi:10.1186/s40425-019-0540-1
- Liang SK, Hsu CC, Song HL, et al. FOXM1 is required for small cell lung cancer tumorigenesis and associated with poor clinical prognosis. Oncogene. 2021;40:4847–4858.
- Li L, Ng SR, Colón CI, et al. Identification of DHODH as a therapeutic target in small cell lung cancer. Sci Transl Med. 2019;11(517). doi:10.1126/scitranslmed.aaw7852.
- Shue YT, Lim JS, Sage J. Tumor heterogeneity in small cell lung cancer defined and investigated in pre-clinical mouse models. Transl Lung Cancer Res. 2018;7(1):21–31. doi:10.21037/tlcr.2018.01.15
- Saltos A, Shafique M, Chiappori A. Update on the biology, management, and treatment of Small Cell Lung Cancer (SCLC). Front Oncol. 2020;10:1074. doi:10.3389/fonc.2020.01074
- Rudin CM, Poirier JT, Byers LA, et al. Molecular subtypes of small cell lung cancer: a synthesis of human and mouse model data. Nat Rev Cancer. 2019;19(5):289–297. doi:10.1038/s41568-019-0133-9
- Gay CM, Stewart CA, Park EM, et al. Patterns of transcription factor programs and immune pathway activation define four major subtypes of SCLC with distinct therapeutic vulnerabilities. Cancer Cell. 2021;39(3):346–360.e347. doi:10.1016/j.ccell.2020.12.014
- Gazdar AF, Bunn PA, Minna JD. Small-cell lung cancer: what we know, what we need to know and the path forward. Nat Rev Cancer. 2017;17(12):725–737. doi:10.1038/nrc.2017.87
- Brägelmann J, Böhm S, Guthrie MR, Mollaoglu G, Oliver TG, Sos ML. Family matters: how MYC family oncogenes impact small cell lung cancer. Cell Cycle. 2017;16(16):1489–1498. doi:10.1080/15384101.2017.1339849
- Wang S, Zimmermann S, Parikh K, Mansfield AS, Adjei AA. Current diagnosis and management of small-cell lung cancer. Mayo Clin Proc. 2019;94(8):1599–1622. doi:10.1016/j.mayocp.2019.01.034
- Mao X, Zhang Z, Zheng X, et al. Capture-based targeted ultradeep sequencing in Paired tissue and plasma samples demonstrates differential subclonal ctDNA-releasing capability in advanced lung cancer. J Thorac Oncol. 2017;12(4):663–672. doi:10.1016/j.jtho.2016.11.2235
- Li YS, Jiang BY, Yang JJ, et al. Unique genetic profiles from cerebrospinal fluid cell-free DNA in leptomeningeal metastases of EGFR-mutant non-small-cell lung cancer: a new medium of liquid biopsy. Ann Oncol. 2018;29(4):945–952. doi:10.1093/annonc/mdy009
- Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–1760. doi:10.1093/bioinformatics/btp324
- Mckenna A, Hanna M, Banks E, et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20(9):1297–1303. doi:10.1101/gr.107524.110
- Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38(16):e164. doi:10.1093/nar/gkq603
- Newman AM, Bratman SV, Stehr H, et al. FACTERA: a practical method for the discovery of genomic rearrangements at breakpoint resolution. Bioinformatics. 2014;30(23):3390–3393. doi:10.1093/bioinformatics/btu549
- Paz-Ares L, Dvorkin M, Chen Y, et al. Durvalumab plus platinum-etoposide versus platinum-etoposide in first-line treatment of extensive-stage small-cell lung cancer (CASPIAN): a randomised, controlled, open-label, Phase 3 trial. Lancet. 2019;394(10212):1929–1939. doi:10.1016/S0140-6736(19)32222-6
- Roper N, Velez MJ, Chiappori A, et al. Notch signaling and efficacy of PD-1/PD-L1 blockade in relapsed small cell lung cancer. Nat Commun. 2021;12(1):3880. doi:10.1038/s41467-021-24164-y
- Barrows ED, Blackburn MJ, Liu SV. Evolving role of immunotherapy in small cell lung cancer. Semin Cancer Biol. 2022. doi:10.1016/j.semcancer.2022.02.021
- Peifer M, Fernández-Cuesta L, Sos ML, et al. Integrative genome analyses identify key somatic driver mutations of small-cell lung cancer. Nat Genet. 2012;44(10):1104–1110. doi:10.1038/ng.2396
- Meuwissen R, Linn SC, Linnoila RI, Zevenhoven J, Mooi WJ, Berns A. Induction of small cell lung cancer by somatic inactivation of both Trp53 and Rb1 in a conditional mouse model. Cancer Cell. 2003;4(3):181–189. doi:10.1016/S1535-6108(03)00220-4
- Olsen RR, Ireland AS, Kastner DW, et al. ASCL1 represses a SOX9(+) neural crest stem-like state in small cell lung cancer. Genes Dev. 2021;35(11–12):847–869. doi:10.1101/gad.348295.121
- George J, Lim JS, Jang SJ, et al. Comprehensive genomic profiles of small cell lung cancer. Nature. 2015;524(7563):47–53. doi:10.1038/nature14664
- Brown LC, Tucker MD, Sedhom R, et al. LRP1B mutations are associated with favorable outcomes to immune checkpoint inhibitors across multiple cancer types. J Immunother Cancer. 2021;9(3):e001792. doi:10.1136/jitc-2020-001792
- Chen H, Chong W, Wu Q, Yao Y, Mao M, Wang X. Association of LRP1B mutation with tumor mutation burden and outcomes in melanoma and non-small cell lung cancer patients treated with immune check-point blockades. Front Immunol. 2019;10:1113. doi:10.3389/fimmu.2019.01113
- Asano Y, Takeuchi T, Okubo H, et al. Nuclear localization of LDL receptor-related protein 1B in mammary gland carcinogenesis. J Mol Med (Berl). 2019;97(2):257–268. doi:10.1007/s00109-018-01732-2
- Edwards ZC, Trotter EW, Torres-Ayuso P, et al. Survival of head and neck cancer cells relies upon LZK kinase-mediated stabilization of mutant p53. Cancer Res. 2017;77(18):4961–4972. doi:10.1158/0008-5472.CAN-17-0267
- Zhang Q, Li X, Cui K, et al. The MAP3K13-TRIM25-FBXW7α axis affects c-Myc protein stability and tumor development. Cell Death Differ. 2020;27(2):420–433. doi:10.1038/s41418-019-0363-0
- Thirumal Kumar D, Susmita B, Judith E, Priyadharshini Christy J, George Priya Doss C, Zayed H. Elucidating the role of interacting residues of the MSH2-MSH6 complex in DNA repair mechanism: a computational approach. Adv Protein Chem Struct Biol. 2019;115:325–350.
- Ling C, Yang W, Sun H, et al. Rare compound heterozygous mutations in gene MSH6 cause constitutive mismatch repair deficiency syndrome. Clin Case Rep. 2018;6(8):1448–1451. doi:10.1002/ccr3.1564
- Itkonen HM, Kantelinen J, Vaara M, et al. Human DNA polymerase α interacts with mismatch repair proteins MSH2 and MSH6. FEBS Lett. 2016;590(23):4233–4241. doi:10.1002/1873-3468.12475
- Radio FC, Pang K, Ciolfi A, et al. SPEN haploinsufficiency causes a neurodevelopmental disorder overlapping proximal 1p36 deletion syndrome with an episignature of X chromosomes in females. Am J Hum Genet. 2021;108(3):502–516. doi:10.1016/j.ajhg.2021.01.015
- Dossin F, Pinheiro I, Żylicz JJ, et al. SPEN integrates transcriptional and epigenetic control of X-inactivation. Nature. 2020;578(7795):455–460. doi:10.1038/s41586-020-1974-9
- Edelmann J, Holzmann K, Tausch E, et al. Genomic alterations in high-risk chronic lymphocytic leukemia frequently affect cell cycle key regulators and NOTCH1-regulated transcription. Haematologica. 2020;105(5):1379–1390. doi:10.3324/haematol.2019.217307
- Li Y, Lv Y, Cheng C, et al. SPEN induces miR-4652-3p to target HIPK2 in nasopharyngeal carcinoma. Cell Death Dis. 2020;11(7):509. doi:10.1038/s41419-020-2699-2
- Légaré S, Chabot C, Basik M. SPEN, a new player in primary cilia formation and cell migration in breast cancer. Breast Cancer Res. 2017;19(1):104. doi:10.1186/s13058-017-0897-3
- Légaré S, Cavallone L, Mamo A, et al. The estrogen receptor cofactor SPEN functions as a tumor suppressor and candidate biomarker of drug responsiveness in hormone-dependent breast cancers. Cancer Res. 2015;75(20):4351–4363. doi:10.1158/0008-5472.CAN-14-3475
- Feng Y, Bommer GT, Zhai Y, et al. Drosophila split ends homologue SHARP functions as a positive regulator of Wnt/beta-catenin/T-cell factor signaling in neoplastic transformation. Cancer Res. 2007;67(2):482–491. doi:10.1158/0008-5472.CAN-06-2314
- Vogt PK, Bader AG, Kang S. Phosphoinositide 3-kinase: from viral oncoprotein to drug target. Virology. 2006;344(1):131–138. doi:10.1016/j.virol.2005.09.027
- Engelman JA, Luo J, Cantley LC. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat Rev Genet. 2006;7(8):606–619. doi:10.1038/nrg1879
- Samuels Y, Wang Z, Bardelli A, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science. 2004;304(5670):554. doi:10.1126/science.1096502
- Meng F, Zhang L. miR-183-5p functions as a tumor suppressor in lung cancer through PIK3CA inhibition. Exp Cell Res. 2019;374(2):315–322. doi:10.1016/j.yexcr.2018.12.003
- Han N, Cheng QY, Chen B, et al. PIK3CA mutations in resected small cell lung cancer. Adv Clin Exp Med. 2016;25(3):397–402. doi:10.17219/acem/25928
- Samuels Y, Velculescu VE. Oncogenic mutations of PIK3CA in human cancers. Cell Cycle. 2004;3(10):1221–1224. doi:10.4161/cc.3.10.1164
- Kawano O, Sasaki H, Endo K, et al. PIK3CA mutation status in Japanese lung cancer patients. Lung Cancer. 2006;54(2):209–215. doi:10.1016/j.lungcan.2006.07.006
- García-Rostán G, Costa AM, Pereira-Castro I, et al. Mutation of the PIK3CA gene in anaplastic thyroid cancer. Cancer Res. 2005;65(22):10199–10207. doi:10.1158/0008-5472.CAN-04-4259
- Liu Z, Roberts TM. Human tumor mutants in the p110alpha subunit of PI3K. Cell Cycle. 2006;5(7):675–677. doi:10.4161/cc.5.7.2605
- Shibata T, Kokubu A, Tsuta K, Hirohashi S. Oncogenic mutation of PIK3CA in small cell lung carcinoma: a potential therapeutic target pathway for chemotherapy-resistant lung cancer. Cancer Lett. 2009;283(2):203–211. doi:10.1016/j.canlet.2009.03.038
- Morris LG, Kaufman AM, Gong Y, et al. Recurrent somatic mutation of FAT1 in multiple human cancers leads to aberrant Wnt activation. Nat Genet. 2013;45(3):253–261. doi:10.1038/ng.2538
- Dotto GP, Rustgi AK. Squamous cell cancers: a unified perspective on biology and genetics. Cancer Cell. 2016;29(5):622–637. doi:10.1016/j.ccell.2016.04.004
- Santana NO, Lerario AM, Schmerling CK, et al. Molecular profile of Hürthle cell carcinomas: recurrent mutations in the Wnt/β-catenin pathway. Eur J Endocrinol. 2020;183(6):647–656. doi:10.1530/EJE-20-0597
- Hsu TN, Huang CM, Huang CS, et al. Targeting FAT1 inhibits carcinogenesis, induces oxidative stress and enhances cisplatin sensitivity through deregulation of LRP5/WNT2/GSS signaling axis in oral squamous cell carcinoma. Cancers. 2019;11(12):1883. doi:10.3390/cancers11121883
- Lustig B, Behrens J. The Wnt signaling pathway and its role in tumor development. J Cancer Res Clin Oncol. 2003;129(4):199–221. doi:10.1007/s00432-003-0431-0
- Pastushenko I, Mauri F, Song Y, et al. Fat1 deletion promotes hybrid EMT state, tumour stemness and metastasis. Nature. 2021;589(7842):448–455. doi:10.1038/s41586-020-03046-1
- Chen Z, Zhang C, Chen J, et al. The proteomic landscape of growth factor signaling networks associated with FAT1 mutations in head and neck cancers. Cancer Res. 2021. doi:10.1158/0008-5472.can-20-3659
- Xu J, Wang B, Liu ZT, Lai MC, Zhang ML, Zheng SS. miR-223-3p regulating the occurrence and development of liver cancer cells by targeting FAT1 gene. Math Biosci Eng. 2019;17(2):1534–1547. doi:10.3934/mbe.2020079
- Valletta D, Czech B, Spruss T, et al. Regulation and function of the atypical cadherin FAT1 in hepatocellular carcinoma. Carcinogenesis. 2014;35(6):1407–1415. doi:10.1093/carcin/bgu054
- Farago AF, Yeap BY, Stanzione M, et al. Combination olaparib and temozolomide in relapsed small-cell lung cancer. Cancer Discov. 2019;9(10):1372–1387. doi:10.1158/2159-8290.CD-19-0582
- Wang XW, Hu N, Cui L, et al. Durable disease-free survival in a patient with metastatic triple-negative breast cancer treated with olaparib monotherapy. Curr Cancer Drug Targets. 2022;22. doi:10.2174/1568009622666220214092207
- Martin NA, Hanna M, Ehret C, Asiedu G, Jatoi A. Olaparib for ovarian cancer: a single-institution, multi-site qualitative study. Support Care Cancer. 2022. doi:10.1007/s00520-022-06879-w