Abstract
Background
Over the decades, new antibacterial agents have been developed in an attempt to combat drug resistance, but they remain unsuccessful. Recently, a novel class of bacterial gene expression regulators, bacterial small RNAs (sRNAs), has received increasing attention toward their involvement in antibiotic resistance. This systematic review aimed to discuss the potential of these small molecules as antibacterial drug targets.
Methods
Two investigators performed a comprehensive search of MEDLINE, EmBase, and ISI Web of Knowledge from inception to October 2016, without restriction on language. We included all in vitro and in vivo studies investigating the role of bacterial sRNA in antibiotic resistance. Risk of bias of the included studies was assessed by a modified guideline of Systematic Review Center for Laboratory Animal Experimentation (SYRCLE).
Results
Initial search yielded 432 articles. After exclusion of non-original articles, 20 were included in this review. Of these, all studies examined bacterial-type strains only. There were neither relevant in vivo nor clinical studies. The SYRCLE scores ranged from to 5 to 7, with an average of 5.9. This implies a moderate risk of bias. sRNAs influenced the antibiotics susceptibility through modulation of gene expression relevant to efflux pumps, cell wall synthesis, and membrane proteins.
Conclusion
Preclinical studies on bacterial-type strains suggest that modulation of sRNAs could enhance bacterial susceptibility to antibiotics. Further studies on clinical isolates and in vivo models are needed to elucidate the therapeutic value of sRNA modulation on treatment of multidrug-resistant bacterial infection.
Introduction
The presence of drug resistance in causative microbes is responsible for excessive mortality and length of hospital stay.Citation1 In a matched cohort comparing bacteremic patients with third-generation cephalosporin-resistant Escherichia coli to those with -susceptible isolates, the former exhibited nearly fivefold increased risk for death in 30 days and had an excessive length of stay of 5 days attributable to infection with a resistant bacteria.Citation2 In critically ill patients, the 28-day mortality rate was twofold higher for patients infected with resistant bacteria comapred with those infected with a susceptible strain.Citation3,Citation4
The presence of resistant microbes limits the choice of antimicrobial chemotherapy. Traditional antimicrobials inhibit the essential synthesis of bacterial components, such as cell wall, membrane, and proteins.Citation5,Citation6 These involve a sophisticated network of enzymes encoded by multiple gene loci. Genetic sequence alteration of these loci is one of the mechanisms through which drug resistance is developed.Citation7 Citation8 These may involve synthesis of a novel protein altering the affinity to an existing drug or a novel enzyme, which is able to replace the functional enzyme otherwise inhibited by an antibiotic.Citation9 Alternatively, drug resistance can be spread horizontally through conjugation, which transfers the drug resistance gene from the donor cell to the recipient cell by directly connecting cell surfaces with pili or adhesins.Citation10,Citation11
This traditional gene-protein paradigm in development of antibiotic resistance has recently been challenged by the recognition of an intermediate nucleotide, the small bacterial regulatory RNAs. The small RNAs (sRNAs), also known as regulatory sRNAs, exist in bacteria with a length of 50–500 nucleotides.Citation12 Based on base-pairing algorithm, they are classified into cis- and trans-encoded sRNAs. Cis-encoded sRNAs are transcribed from the same loci as the mRNAs on the opposite strand of DNA and bind to their cognate mRNA targets with perfect complementarity, resulting in either transcriptional termination or translational initiation.Citation12 Their actions are independent of chaperone. On the other hand, trans-encoded sRNAs do not display high specificity to a given mRNA. Instead, a given trans-encoded sRNA interacts with multiple mRNA targets.Citation13,Citation14 These trans-sRNAs are functionally dependent on chaperones.Citation15 Hfq and CsrA are essential chaperones for the activity of numerous trans-sRNAs. When Hfq and CsrA activity were inhibited and subsequently inhibited their downstream sRNA networks, bacterial infectiveness and susceptibility to antibiotics were reduced.Citation16,Citation17 The distribution of cis- and trans-sRNAs differs between Gram-positive and Gram-negative organisms.Citation18 The biological mechanism of sRNA has been comprehensively reviewed.Citation14 These sRNAs are important in controlling bacterial gene expression in response to extracellular stress and maintaining microbial cell homeostasis.Citation19,Citation20 By complementary binding to C-terminus of RNase or a promoter region of a transcription factor, the sRNA molecule either actively degrades the encoded proteins or halts the transcription of the encoded proteins.Citation21,Citation22
Accumulating evidence suggests that sRNA is differentially expressed in bacterial transition from colonization to active infectionCitation23,Citation24 and may represent one of the mechanisms for the organism to adapt to the changing environment as disease develops.Citation25 Environmental stress such as exposure of antibiotics contributes to the physiological change of bacterial cells. Exposure of selective bacterial strains to antibiotics has revealed a panel of differentially expressed sRNAs.Citation26–Citation29 More importantly, overexpression of some of these sRNAs increases antibiotic susceptibility of the otherwise highly resistant bacterial strains.Citation30,Citation31 In this review, we performed a systematic search of the literature to discuss the therapeutic potential of these sRNAs in combating against the big problem of multidrug-resistant bacterial infection.
Methods
Literature search
We performed a search of published literature indexed in three electronic bibliographic databases: PubMed Medline, Web of Knowledge, and Embase. These search engines have previously been reported for use in systematic search.Citation32 The search terms included a combination of “antibiotic resistance”, “riboswitch”, and “bacterial small RNA”. Manual search of relevant articles was performed to identify other relevant articles. The details of manual search are as follows: titles of the studies were screened for match with predesigned inclusion criteria. The duplicate studies were eliminated among three databases. After that, abstracts of the eligible studies were screened by two authors and full-text articles were evaluated to determine whether they fulfilled the eligibility criteria. Publications from inception to October 2016 were included. Two authors (HC and JH) performed the literature search and eligibility assessment independently. Disagreements were resolved by consensus or discussion with senior authors. We followed suggestions from the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement to conduct the literature search.Citation32
Selection criteria and data abstraction
We included all in vitro and in vivo studies deciphering the role of bacterial sRNAs in antibiotic resistance. Review articles, editorials, commentaries, conference abstracts, and book chapters were excluded. Studies reporting bacterial ribosomal RNAs or other RNAs outside the 50–500 nucleotides range were excluded.
HC performed the data abstraction, which was then verified by another researcher (JH). The following data were abstracted: author, year of publication, sample size, methods for detecting sRNAs, antibiotics used, and functional determination of sRNAs ().
Table 1 Summary of the included studies
Quality assessment of the included studies
The quality of the included studies was evaluated using a modified guideline from SYstematic review Center for Laboratory animal Experimentation (SYRCLE), an instrument based on the Cochrane Collaboration Risk of Bias tool.Citation33,Citation34 An item concerning random housing of animals was removed in the modified version. The remaining items assess selection bias, performance bias, detection bias, attribution bias, and reporting bias. These factors are common among in vitro and in vivo studies. Higher SYRCLE scores indicate better quality article. The maximum score is 9.
Results
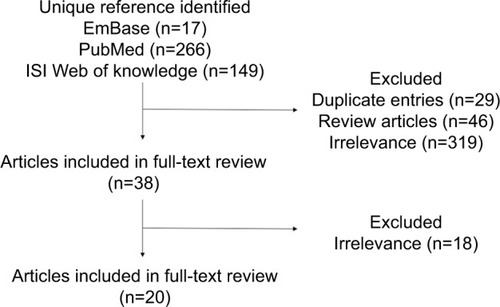
The initial search for the literature using three search engines yielded 432 eligible articles examining the bacterial small regulatory RNAs in the modulation of antibiotic resistance. Based on screening of the titles, abstracts, and keywords, we excluded 29 duplicated entries, 46 reviews, and 337 studies that are irrelevant to bacterial sRNA and antibiotics resistance, in which 32% of the studies are related to small molecule–or protein-mediating activity in antibiotics resistance without mentioning sRNA, 26% to ribosomal RNAs in antibiotics resistance, 20% to the point mutation of bacterial strains without involvement of sRNA, 8% to a platform or method to screen or investigate the RNAs or protein in antibiotics resistance, and 4% to new antibiotic–resistant clinical isolates or species. Eventually, 20 articles were included in our systematic review ().
Quality assessment of the included studies
Overall, the SYRCLE scores ranged from 5 to 7, with an average of 5.9. This implies a moderate risk of bias (). None of the studies fulfilled the criteria for allocation concealment and experimental blinding. Four studies applied inadequate generation of allocation sequences.Citation35–Citation38 Several studies have demonstrated randomization for outcome assessment.Citation26,Citation27,Citation39,Citation40
Table 2 Summary of study quality assessment
The species of bacteria included in the studies
First, we reported the species of bacteria that are related to the sRNAs. The role of sRNAs has been examined largely in laboratory reference strains but not in clinical isolates. There were no relevant in vivo or clinical studies. A wide range of bacterial species were investigated in these studies, including Escherichia coli, Acinetobacter baumannii, Pseudomonas aeruginosa, Vibrio cholera, Salmonella typhimurium, Enterococcus faecalis, Neisseria menin-gitidis, Staphylococcus aureus, and Clostridium acetobu-tylicum. Seven studies were performed on E. coli,Citation30,Citation35,Citation39,Citation41,Citation44, three on Pseudomonads,Citation26,Citation31,Citation45 and two on Salmonella spp.Citation27,Citation46 The remaining genera were examined by a single study.Citation29,Citation35,Citation36,Citation47,Citation48
Experimental techniques for included studies to investigate bacterial sRNAs
Second, experimental methods are essential to explore sRNAs or its targets. We classified the experimental techniques used in the included studies. Five studies performed functional investigation on bacterial sRNAs by profiling their expression patterns after exposure of bacterial strains to specific antibiotics.Citation26–Citation29,Citation47 The remaining articles employed genetic silencing and modulation approach to evaluate the role of sRNA. Antibiotic resistance-encoding genes were either deleted or introduced into the experimental strains.Citation30,Citation31,Citation35–Citation49 This was followed by overexpression of sRNAs of interest in order to detect changes of antibiotic susceptibility.Citation35–Citation49
Mechanism of sRNAs in antibiotics resistance
Third, we elucidated the mechaisms of sRNAs in antibiotics resistance. The modulation of antibiotic sensitivity by sRNAs is pertinent to the synthesis of efflux pump, cell wall, transporters, and outer membrane proteins. The role of sRNAs in antibiotic resistance is summarized in and . The biological mechanism of sRNA is depicted in .
Figure 2 Small bacterial RNAs interact with canonical (in blue) and unknown mechanisms (in red) in acquisition of antimicrobial resistance phenotype. The pointed arrowheads denote putative stimulatory effect whereas the blunted arrowheads represent inhibitory actions.
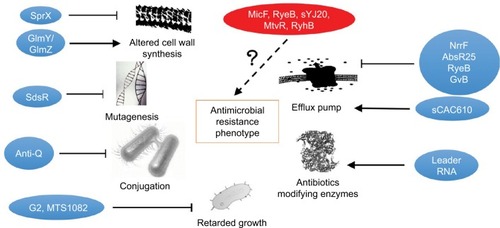
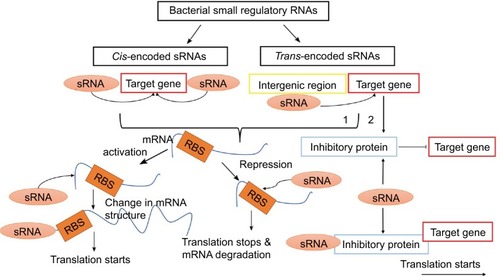
Figure 3 The biological action of small bacterial RNAs. These molecules are classified into cis- and trans-encoded sRNAs. By binding to non-coding region of mRNA or a gene per se, sRNAs either activate or repress the expression of a given protein.
Abbreviation: RBS, RNA binding site.
The included studies have shown that regulated expression of bacterial sRNAs is important in sustaining antibiotic- resistant phenotypes and conferring bacterial survival advantage.Citation26–Citation31,Citation47 Interestingly, experimental alteration of antibiotic-responsive sRNAs considerably increased bacterial susceptibility to various antibiotics.Citation30,Citation31,Citation49
Discussion
Although new antibacterial agents have been developed in an attempt to combat drug resistance, this remains largely unsuccessful. As soon as a new agent enters the market, the targeted bacteria develop novel strategies to minimize their susceptibility, shortening the lifespan of newly introduced antibiotics. This warrants continuous search for novel bacterial targets with less likelihood to induce drug resistance.
Bacterial sRNAs are regulatory molecules in response to the constant changes of the surrounding environment.Citation30,Citation42,Citation43 As such, sRNAs are important for bacterial adaptability in the presence of harmful substances. By translational bypassing, these short-sequence nucleotides respond promptly to antimicrobial challenge.Citation14 With non-specific and imperfect binding to mRNAs, these sRNA molecules can modulate the activity of multiple mRNAs simultaneously.Citation50 Given the essential pathways and multiple targets regulated by a given sRNA, modulation of this moiety may be a promising therapeutic approach with less likelihood of antimicrobial resistance development. This is because the sRNAs modulate the translation process of antmicriobial resistant genes instead of whole bacterial genome.Citation51 In addition, sRNA targeting mRNA sequences distinct from those in humans will exhibit excellent selective toxicity. This will provide an alternative therapy to highly toxic antibiotics such as aminoglycosides.
sRNAs involved in mutation-induced antibiotic resistance
After exposure to antibiotics in sublethal concentration, bacterial growth is hindered and mutagenesis is favored.Citation47,Citation52 For instance, the expression of sRNAs G2 and MTS1082 in Mycobacterium tuberculosis was repressed at stationary phase of growth, which is associated with their antibiotic-tolerant phenotype.Citation47 In E. coli, the presence of growth-limiting stressor induced a transcription factor known as RpoS regulon.Citation52 This complex leads to subsequent expression of sdsR, an RpoS-dependent sRNA.Citation52,Citation53 This negatively regulated the ampicillin-induced mutagenesis by binding to mutS, an mRNA molecule encoding protein for mismatch detection.Citation53
Pro-survival benefits via modulating sRNA expression
The sRNA molecule is important for bacterial survival in the presence of antimicrobial challenge. In Staphylococcus aureus, exposed to a combination of vancomycin, linezolid, ceftobiprole, and tigecycline, expression of specific sRNAs was prominent. The change in the expression of these sRNAs maintains protein synthesis and ribosomal function in the presence of antimicrobial challenge, thereby ensuring bacterial survival.Citation26,Citation27,Citation31 Upon the exposure of antibiotics, genes coding proteins including metabolism, transport, transcriptional regulation, and ribosome proteins are unregulated.Citation26 In the presence of tigecycline and tetracycline, sRNA sYJ20 was upregulated in Salmonella typhimurium. Deletion of sYJ20 reduced the bacterial survival rate upon exposure to tigecycline.Citation28 sRNA sYJ20 regulated the antibiotic tolerance through modulation of tigecycline and tetracycline relevant encoding genes.
Increasing antimicrobial sensitivity by targeting sRNAs
Modulation of sRNAs has been shown to increase bacterial sensitivity to antibiotics. Experimental upregulation of sRNA, MtvR, enhanced the sensitivity of E. coli to tetracycline, chloramphenicol, ciprofloxacin, tobramycin, gentamycin, and ampicillin by two- to eightfold.Citation30 In Pseudomonas aerugi-nosa, overexpression of MtvR significantly suppressed their growth upon exposure to chloramphenicol, ciprofloxacin, tetracycline, tobramycin, gentamycin, and ampicillin.Citation31 Conversely, sRNA-induced modulation of antibiotic susceptibility was reversed by depletion of the respective sRNAs.Citation30,Citation31
In response to ampicillin and norfloxacin, the expression pattern of RyhB sRNA changed considerably in E. coli. Ampicillin and norfloxacin share common pathways in the production of highly deleterious hydroxyl radicals in the tricarboxylic acid cycle (TCA).Citation54 The RyhB sRNA targets AcnA, a gene encoding the TCA cycle-related enzyme in E. coli.Citation49,Citation55 Perturbation of the TCA cycle-related enzymes triggers TCA cycle dysfunctions and subsequently makes bacteria less susceptible to β-lactam antibiotics by altering its cell surface properties.Citation56
Although accumulating evidence has suggested that modulating sRNA expression alters bacterial susceptibility to antibiotics, these appear to be strain specific. In addition, laboratory reference strains may not reflect clinical situation as type strains likely behave differently from clinical strains.Citation57 To extrapolate these findings into therapeutic drug development, the candidate sRNA should be universally responsive to a given antibiotic in different strains of the same bacterial genus, or at least be consistently responsive in the same species. Clinical isolates should be thoroughly tested with complete profile of pharmacokinetics and pharmacodynamics of the exogenous sRNA modulators established.
Reversal of efflux pump-mediated resistance
The presence of multidrug efflux pump represents one of the important mechanisms conferring antibiotic resistance. It prevents the accumulation of drugs in bacterial cells, enabling the bacteria to survive. In Acinetobacter baumannii, an efflux pump regulating sRNA-designated AbsR25 was initially recognized by its high hybridization energy.Citation36 In E. coli, AcrAB-TolC represents a major efflux pump conferring multidrug-resistant phenotype.Citation58 Deletion of tolC rendered the mutants more sensitive to antibiotics as compared to the wild-type counterparts.Citation59 The sRNA sdsR, or alternatively known as RyeB, post-transcriptionally represses the expression of tolC by base-pairing with its 5′-untranslated region (5′-UTR).Citation45 MicF represents another important sRNA which is presumably involved in the regulation of efflux pumps. The MicF is a two-component response regulator contributing to carbenicillin resistance. In E. coli, the sRNA MicF response regulator inhibited the transcription of the ompF gene encoding porins, which reduced the pore-forming antibiotics such as β-lactams to enter the cell interior.Citation44 In Neisseria species, trans-acting sRNA NrrF played a crucial role in transcriptional control of efflux pump synthesis. Experimental overex-pression of NrrF decreased the antibiotic-induced expression of efflux pumps.Citation48
Other transporters in bacteria were correlated with the efflux pumps and further implicated the antibiotic resistance in bacteria.Citation60 In Clostridium acetobutylicum, the sRNA sCAC610 regulates ATP binding cassette transporter genes.Citation29 In the presence of clindamycin, the level of sCAC610 was significantly upregulated. Bioinformatics analysis revealed that sCAC610 modulated efflux pump function through cis-regulation.Citation29 D-cycloserine (DCS) is a second-line antibiotic for the treatment of multidrug-resistant Mycobacterium tuberculosis.Citation61 In this organism, an sRNA molecule GcvB suppressed the expression of cycA, a gene encoding an enzyme for degradation of DCS.Citation39,Citation62
Collectively, targeted delivery of sRNA agonists or antagonists may represent alternative approach to combat against multidrug-resistant Enterobacteriaceae and tubercle bacilli.
Reversal of resistance to cell wall inhibitors
Resistance to cell wall synthesis inhibitors predominantly involves overproduction of degradative enzymes and formation of alternative membrane protein with low affinity to antibiotics. Penicillins and cephalosporins are major antibiotic classes under this category. Recently, it has been appreciated that some sRNAs are activated upon encountering cell wall synthesis inhibitors in an attempt to restore cell wall integrity. Bacilysin, a peptide antibiotic, inhibits glucosamine-6-phosphate synthase (GlmS) activity and thereby interferes with cell wall synthesis. However, suppression of GlmS by bacilysin in E. coli activates the expression of two sRNAs, namely GlmY and GlmZ, which in return restore the expression of GlmS, this resulting in a resistance phenotype.Citation42 In this connection, co-administration of bacilysin with GlmY/GlmZ antagonist may reduce the therapeutic dose of the antibiotics concerned.
The increasing prevalence of methicillin-resistant Staphylococcus aureus with reduced susceptibility to vancomycin represents another important public health threat. To date, the mechanisms contributing to this non-susceptibility remain unclear, although point mutation and thickening of cell walls have been suggested.Citation7 Comparative proteomic studies revealed that a small RNA molecule SprX inhibits the expression of stage V sporulation protein G (SpoVG), a moiety contributing to non-susceptibility to glycopeptides.Citation46,Citation62 It remains unclear under which condition this RNA molecule is activated. However, this suggests that co-administration of SprX activators with glycopeptides may reduce the likelihood of resistance development during therapy. Whether this combination results in synergy or antagonism will need to be elucidated.
Reversal of minimal membrane permeability-mediated resistance
One of the important mechanisms by which Gram-negative bacteria gain survival benefits in face of antibiotic challenge is reducing their outer membrane permeability to aqueous drugs by loss of porins.Citation43,Citation63 The control of the porins’ expression has been studied thoroughly in E. coli. It has been shown that mutation of soxRS reduced the expression of a major drug target porin, namely OmpF, through a sRNA molecule called micF.Citation64 As such, inhibition of micF sensitizes bacteria to aqueous antibiotics.Citation43 Serial passage of laboratory strains of E. coli has revealed the acquisition of an alternative porin called Chip in compensation for the loss of drug target porins OmpC and OmpF.Citation41 Further molecular characterization led to identification of a sRNA known as ChiX, which downregulates the expression of Chip porins and restores the bacterial susceptibility to carbapenams.Citation41
Reversal of other drug resistance mechanisms
Bacterial riboswitch is a cis-encoded sRNA situated at the non-coding region of mRNA.Citation35,Citation38,Citation40 The aptamer domain of this RNA molecule binds to specific metabolites, thereby exerting post-transcriptional control of their expression.Citation65,Citation66 In Enterobactericeae, the presence of a riboswitch with theophylline-specific aptamer sequence mediates the production of a range of β-lactamases which further confer β-lactam antibiotic resistance.Citation35 Therefore, β-lactam antibiotic resistance can be eliminated by reduction of this riboswitch activity. In aminoglycoside resistance, the major mechanism is attributable to the presence of nucleotidyltransferases, which modify the drug and render it ineffective. These enzymes are mainly encoded by aac and aad genes. The 5′ leader RNA, a riboswitch, of the aac/aad genes interacts with aminoglyco-sides and in turn activates the expression of the modifying enzymes.Citation38,Citation40,Citation67 In this regard, development of an antagonist, which prevents the binding of aminoglycosides to the leader sequence, may be useful to avoid the development of inducible resistance. Alternatively, modification of side chains of the next-generation aminoglycoside can facilitate their interaction with 30S ribosomal subunit rather than the 5′ leader sequence leading to transcription of degradative enzymes.
Transfer of resistance determinants across bacterial species by conjugation has been increasingly recognized as a novel strategy by which bacteria acquire drug resistance.Citation68 This may be an important reason, if not the only one, which accounts for epidemiology peculiarity of antibiotic resistance. In Enterococcus faecalis, a small RNA molecule called anti-Q has been shown to suppress the conjugation of the plasmid pCF10 by termination of prgX repression and prgQ conjugation operons.Citation37
In this connection, conjugation suppressor sRNAs may be employed to prevent inter-bacterial transfer of resistance determinants. Development of air spray containing these suppressor molecules may be applicable for use in environments such as intensive care units to prevent interspecies transfer of resistance determinants.
Concluding remarks and future perspectives
Experimental studies using type strains have suggested that targeting sRNA increased their susceptibility to antibiotics in the otherwise resistance strains. Unlike conventional antibiotics, sRNA is a powerful master regulator in gene regulation which inhibited the translation of bacterial protein synthesis. sRNA has a great potential to develop as a novel drug or become a novel acceessory ingredient in medical use. Given multiple targets of a single sRNA molecule, modulating sRNAs in bacterial strains in vivo using bacteriophages or nanoparticles may be a promising therapeutic approach in clinical use. Co-administration of sRNA agonist or antagonist with conventional antibiotics may represent an alternative option to minimize the emergence of resistance during therapy. Further studies with clinical isolates and in vivo models are necessary to elucidate the real clinical situation and therapeutic potential of sRNA modulators.
Acknowledgments
This study was funded by Health and Medical Research Fund (15140132); Commissioned Research on Control of Infectious Disease (Phase III) (CU-15-B2), Food and Health Bureau, Hong Kong; and internal research fund of the Department of Anesthesia and Intensive Care, Faculty of Medicine, the Chinese University of Hong Kong. The authors would like to thank Wai T Wong, Thomas Kwong, Gordon Choi, Czarina CH Leung, Sharon Tsang, Tony Gin, Jun Yu, and Gary Tse for their contribution to this manuscript. As they did not fulfill the criteria for authorship according to Good Publication Practice 3 guideline, we apologise for not listing them as co-authors.
Disclosure
The authors report no conflicts of interest in this work.
References
- de KrakerMEWolkewitzMDaveyPGClinical impact of antimicrobial resistance in European hospitals: excess mortality and length of hospital stay related to methicillin-resistant Staphylococcus aureus bloodstream infectionsAntimicrob Agents Chemother20115541598160521220533
- de KrakerMEWolkewitzMDaveyPGBurden of antimicrobial resistance in European hospitals: excess mortality and length of hospital stay associated with bloodstream infections due to Escherichia coli resistant to third-generation cephalosporinsJ Antimicrob Chemother20116639840721106563
- CombesALuytCEFagonJYImpact of pieracillin resistance on the outcome of Pserudomonas ventilator-associated pneumoniaIntensive Care Med2006321970197816957901
- TabahAKoulentiDLauplandKCharacteristics and determinants of outcome of hospital-acquired bloodstream infections in intensive care units: the EUROBACT International Cohort StudyIntensive Care Med2012381930194523011531
- NikolaidisIFavini-StabileSDessenAResistance to antibiotics targeted to the bacterial cell wallProtein Sci20142324325924375653
- BakheetTMDoigAJProperties and identification of antibiotic drug targetsBMC Bioinformatics20101119520406434
- LevySBThe 2000 Garrod lecture. Factors impacting on the problem of antibiotic resistanceJ Antimicrob Chemother200249253011751763
- DoddangoudarVCBoostMVTsangDNTracking changes in the vraSR and graSR two component regulatory systems during the development and loss of vancomycin non-susceptibility in a clinical isolateClin Microbiol Infect2011171268127221375655
- KohanskiMADwyerDJCollinsJJHow antibiotics kill bacteria: from targets to networksNat Rev Microbiol20108642343520440275
- SmillieCGarcillan-BarciaMPFranciaMVMobility of plasmidsMicrobiol Mol Biol Rev20107443445220805406
- WozniakRAWaldorMKIntegrative and conjugative elements: mosaic mobile genetic elements enabling dynamic lateral gene flowNat Rev Microbiol2010855256320601965
- KwendaSGorshkovVRameshAMDiscovery and profiling of small RNAs responsive to stress conditions in the plant pathogen Pectobacterium atrosepticumBMC Genomics2016174726753530
- VogelJWagnerEGTarget identification of small noncoding RNAs in bacteriaCurr Opin Microbiol20071026227017574901
- WatersLSStorzGRegulatory RNAs in bacteriaCell200913661562819239884
- EllisMJTrusslerRSHanifordDBA cis-encoded sRNA, Hfq and mRNA secondary structure act independently to suppress IS200 transpositionNucleic Acids Res2015436511652726044710
- OlivaGSahrTBuchrieserCSmall RNAs, 5′ UTR elements and RNA-binding proteins in intracellular bacteria: impact on metabolism and virulenceFEMS Microbiol Rev201539333134926009640
- MuhlenSDerschPAnti-virulence strategies to target bacterial infectionsCurr Top Microbiol Immunol201639814718326942418
- JousselinAMetzingerLFeldenBOn the facultative requirement of the bacterial RNA chaperone, HfqTrends Microbiol20091739940519733080
- VogelJLuisiBFHfq and its constellation of RNANat Rev Microbiol2011957858921760622
- ZhouYXieJThe roles of pathogen small RNAsJ Cell Physiol201122696897320945366
- MoritaTMakiKAibaHRNase E-based ribonucleoprotein complexes: mechanical basis of mRNA destabilization mediated by bacterial non-coding RNAsGenes Dev2005192176218616166379
- BouvierMSharmaCMMikaFSmall RNA binding to 5′ mRNA coding region inhibits translational initiationMol Cell20083282783719111662
- NithyaRAhmedSAHoeCHNon-protein coding RNA genes as the novel diagnostic markers for the discrimination of Salmonella species using PCRPLoS One201510e011866825774907
- SongJLaysCVandeneschFThe expression of small regulatory RNAs in clinical samples reflects the different life styles of Staphylococcus aureus in colonization vs. infectionPLoS One20127e3729422629378
- XiaLXiaWLiSIdentification and expression of small non-coding RNA, L10-Leader, in different growth phases of Streptococcus mutansNucleic Acid Ther20122217718622468692
- Molina-SantiagoCDaddaouaAGomez-LozanoMDifferential transcriptional response to antibiotics by Pseudomonas putida DOT-T1EEnviron Microbiol2015173251326225581266
- HowdenBPBeaumeMHarrisonPFAnalysis of the small RNA transcriptional response in multidrug-resistant Staphylococcus aureus after antimicrobial exposureAntimicrob Agents Chemother2013573864387423733475
- YuJSchneidersTTigecycline challenge triggers sRNA production in Salmonella enterica serovar TyphimuriumBMC Microbiol20121219522958399
- ChenYLIndurthiDCJonesSWSmall RNAs in the genus ClostridiummBio20112e003401021264064
- KimTBakGLeeJSystematic analysis of the role of bacterial Hfq-interacting sRNAs in the response to antibioticsJ Antimicrob Chemther20157016591668
- RamosCGGriloAMSousaSARegulation of Hfq mRNA and protein levels in Escherichia coli and Pseudomonas aeruginosa by the Burkholderia cenocepacia MtvR sRNAPLoS One20149e9881324901988
- LiberatiAAltmanDGTetzlaffJThe PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate health care interventions: explanation and elaborationJ Clin Epidemiol200962e13419631507
- HooijmansCRRoversMMde VriesRBSYRCLE’s risk of bias tool for animal studiesBMC Med Res Methodol2014144324667063
- HoJChanHWongHThe involvement of regulatory non-coding RNAs in sepsis: a systematic reviewCrit Care20162038327890015
- LiuLWangSEngineered riboswitch as a gene-regulatory platform for reducing antibiotic resistanceMethods Mol Biol2014111125125824549625
- SharmaRAryaSPatilSDIdentification of novel regulatory small RNAs in Acinetobacter baumanniiPLoS One20149e9383324705412
- JohnsonCMChenYQLeeHIdentification of a conserved branched RNA structure that functions as a factor-independent terminatorProc Natl Acad Sci U S A20141113573357824550474
- JiaXZhangJSunWXRiboswitch control of aminoglycoside antibiotic resistanceCell2013152688123332747
- BaisaGStaboNJWelchRACharacterization of Escherichia coli D-cycloserine transport and resistant mutantsJ Bacteriol20131951389139923316042
- ChenDMurchieAIAn aminoglycoside sensing riboswitch controls the expression of aminoglycoside resistance acetyltransferase and adenyltransferasesBiochim Biophys Acta2014183995195824631585
- KnoppMAnderssonDIAmelioration of the fitness costs of antibiotic resistance due to reduced outer membrane permeability by upregulation of alternative porinsMol Biol Evol2015323252326326358402
- KhanMAGopelYMilewskiSTwo small RNAs conserved in Enterobacteriaceae provide intrinsic resistance to antibiotics targeting the cell wall biosynthesis enzyme glucosamine-6-phosphate synthaseFront Microbiol2016790827379045
- ChouJHGreenbergJTDempleBPosttranscriptional repression of Escherichia coli OmpF protein in response to redox stress: positive control of the micF antisense RNA by the soxRS locusJ Bacteriol1993175102610317679383
- AllenHKAnRHandelsmanJA response regulator from a soil metagenome enhances resistance to the beta-lactam antibiotic carbenicillin in Escherichia coliPLoS One201510e012009425782011
- ParkerAGottesmanSSmall RNA regulation of TolC, the outer membrane component of bacterial multidrug transportersJ Bacteriol20161981101111326811318
- EyraudATattevinPChabelskayaSA small RNA controls a protein regulator involved in antibiotic resistance in Staphylococcus aureusNucleic Acids Res2014424892490524557948
- JeevesREMarriottAAPullanSTMycobacterium tuberculosis is resistant to isoniazid at a slow growth rate by single nucleotide polymorphisms in katG codon Ser315PLoS One201510e013825326382066
- JacksonLAPanJCDayMWControl of RNA stability by NrrF, an iron-regulated small RNA in Neisseria gonorrhoeaeJ Bacteriol20131955166517324039262
- KimJNKwonYMGenetic and phenotypic characterization of the RyhB regulon in Salmonella typhimuriumMicrobiol Res2013168414922824499
- GottesmanSMicros for microbes: non-coding regulatory RNAs in bacteriaTrends Genet20052139940415913835
- StorzGVogelJWassarmanKMRegulation by small RNAs in bacteria: expanding frontiersMol Cell201143688089121925377
- LombardoMJAponyiIRosenbergSMGeneral stress response regulator RpoS in adaptive mutation and amplification in Escherichia coli.Genetics200416666968015020458
- GutierrezALauretiLCrussardSBeta-lactam antibiotics promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelityNat Commun20134161023511474
- KohanskiMADwyerDJHayeteBA common mechanism of cellular death induced by bactericidal antibioticsCell200713079781017803904
- MasseEGottesmanSA small RNA regulates the expression of genes involved in iron metabolism in Escherichia coliProc Natl Acad Sci U S A2002994620462511917098
- ThomasVCKinkeadLCJanssenAA dysfunctional tricarboxylic acid cycle enhances fitness of Staphylococcus epidermidis during beta-lactam stressmBio201344 pii: e00437-13
- FuxCAShirtliffMStoodleyPCan laboratory reference strains mirror “real-world” pathogenesis?Trends Microbiol200513586315680764
- NikaidoHStructure and mechanism of RND-type multidrug efflux pumpsAdv Enzymol Relat Areas Mol Biol20117716021692366
- AugustusAMCelayaTHusainFAntibiotic-sensitive TolC mutants and their suppressorsJ Bacteriol20041861851186014996816
- MousaJJBrunerSDStructural and mechanistic diversity of multidrug transportersNat Prod Rep2016331255126727472662
- RiccardiGPascaMRBuroniSMycobacterium tuberculosis: drug resistance and future perspectivesFuture Microbiol2009459761419492969
- MeierSGoerkeCWolzCSigmaB and the sigmaB-dependent arlRS and yabJ-spoVG loci affect capsule formation in Staphylococcus aureusInfect Immun2007754562457117635871
- FernandezLHancockREAdaptive and mutational resistance: role of porins and efflux pumps in drug resistanceClin Microbiol Rev20122566168123034325
- MizunoTChouMYInouyeMA unique mechanism regulating gene expression: translational inhibition by a complementary RNA transcript (micRNA)Proc Natl Acad Sci U S A198481196619706201848
- DebRoySGebbieMRameshAA riboswitch-containing sRNA controls gene expression by sequestration of a response regulatorScience201434593794025146291
- TernanNGSmall regulatory RNA molecules in bacteriaOA Microbiol201311
- HeWZhangXZhangJRiboswitch control of induction of aminoglycoside resistance acetyl and adenyl-transferasesRNA Biol2013101266127323880830
- WendlandtSLiJHoJEnterococccal multi-resistance gene cluster in methicillin-resistant Staphyloccus aureus from various origins and geographical locaitonsJ Antimicrob Chemother201433