Abstract
Over the last few decades, the incidence of oral cancer has gradually increased, due to the negative influence of environmental factors and also abnormalities within the genome. The main issues in oral cancer treatment consist in surpassing resistance and recurrence. However, continuous discovery of altered signaling pathways in these tumors provides valuable information for the identification of novel gene candidates targeted in personalized therapy. RNA interference (RNAi) is a natural mechanism that involves small interfering RNA (siRNA); this can be exploited in biomedical research by using natural or synthetic constructs for activation of the mechanism. Synthetic siRNA transcripts were developed as a versatile class of molecular tools that have a diverse range of programmable roles, being involved in the regulation of several biological processes, thereby providing the perspective of an alternative option to classical treatment. In this review, we summarize the latest information related to the application of siRNA in oral malignancy together with molecular aspects of the technology and also the perspective upon the delivery system. Also, the emergence of newer technologies such as clustered regularly interspaced short palindromic repeats/Cas9 or transcription activator-like effector nucleases in comparison with the RNAi approach is discussed in this paper.
Introduction
A group of tumors on the lip, tongue, mouth, and oropharynx are grouped under the name of oral cancer.Citation1 Global epidemiologic data of oral cancer reflect a 20-fold geographic variation, with almost two-thirds of all newly diagnosed cases being reported in developing countries.Citation2,Citation3 Betel chewing, a traditional practice in South-Eastern Asia and in the Pacific Ocean, represents the greatest risk factor, followed by tobacco smoking, human papillomavirus (HPV) infection, alcohol consumption, and nutrient deficiency.Citation4 On a global scale, oral cancer could be considered a deadly disease, considering that out of the 300,000 newly diagnosed cases, 145,000 deaths were reported in 2012.Citation3 The major challenge in the treatment of oral cancer remains the emergence of drug resistance and also recurrence episodes; therefore, improved therapeutic options that surpass these issues are necessary.Citation5,Citation6
In the past, molecular biology studies were centered around protein coding genes, supported by the central dogma of biology, which postulates that messenger RNA (mRNA) only acts as a mediator between a gene and its encoded protein.Citation7–Citation10 The noncoding regions of the genome were considered as nonfunctional entities and classified as “junk DNA” or “dark-matter.”Citation11–Citation15 However, genome sequencing performed over the last 2 decades brought scientists to the conclusion that the phenotypes are generated not only by proteins and their coding genes but also by noncoding regulatory elements that modulate gene expression programs.Citation13,Citation16–Citation18 Noncoding RNAs (ncRNAs) have the common feature of not being translated into proteins, hence their name.Citation19–Citation21 As a very diverse class, these transcripts are classified based on the sequence length, in: long ncRNAs (with >200 nt) and short ncRNAs (with <200 nt), as presented in . The short ncRNAs group contains transcripts, such as microRNAs (miRNAs), small interfering RNAs (siRNAs), and piwi-interacting RNAs.Citation9,Citation19,Citation22,Citation23
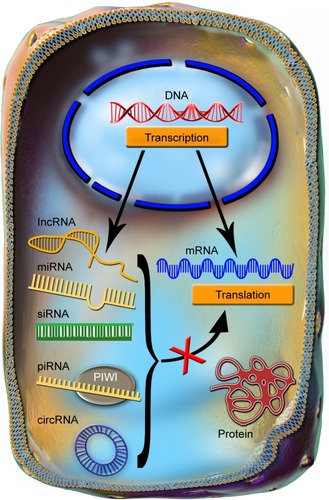
Figure 1 The common features of ncRNAs.
Notes: The ncRNAs are transcribed from the DNA and through multiple processing steps are exported into the cytoplasm, in a manner that bears a higher or lower degree of similarity with the processing of mRNA. In the cytoplasm, the main function of the ncRNAs is to repress the translation of mRNAs into proteins.
Abbreviations: circRNA, circular RNA; lncRNA, long noncoding RNAs; miRNA, microRNAs; mRNA, messenger RNA; ncRNAs, noncoding RNAs; piRNA, piwi-interacting RNA; siRNA, small interfering RNA.
RNA interference (RNAi) was first considered as an exogenous RNA in plants.Citation24,Citation25 It was later discovered that siRNA can have an endogenous origin in many types of cells, human cells included, and that its expression is dysregulated during the development and progression of various pathologies, including cancer.Citation25,Citation26 Both siRNAs and miRNAs are processed by the same proteins: Dicer and RNA-induced silencing complex (RISC). Dicer cleaves the double-stranded precursor into a 23–25 nucleotide double-stranded RNA (dsRNA), with a two-nucleotide overhang at each end, while RISC has two functions: keeping only the guide strand of the dsRNA, while discarding the passenger strand and cleaving the mRNA between its 10th and 11th base, after siRNA full-complementarily binds to the 3′ end ().Citation25,Citation27 The exogenous therapeutic siRNA can be delivered as dsRNA ready to be integrated into the RISC complex or as short-hairpin RNA (shRNA), a precursor of siRNA that is cleaved by Dicer and then loaded into the RISC complex. The dsRNA, referred from now on simply as siRNA, is the most commonly used. One example is represented by siRNAs targeting the epidermal growth factor receptor (EGFR) gene that is overexpressed in oral cancer cells.Citation28
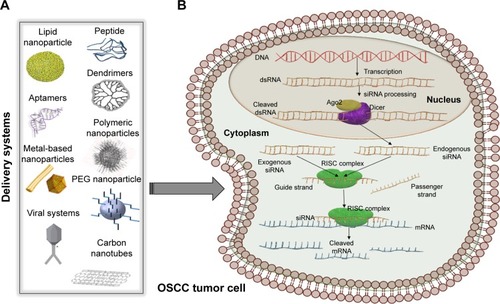
Figure 2 siRNA therapeutic implication.
Notes: (A) Some relevant examples of nanosystems used for siRNA application, comprising a great variety of viral/nonviral delivery systems, some of which are used in oral cancer: lipid nanoparticles, nanorods, peptides, dendrimers, PEG nanoparticles, or adeno-associated viruses with application in oral cancer. (B) Schematic view of siRNA mechanism. siRNAs can have an endogenous origin, being transcribed from the genome and processed by Dicer or it can be delivered by artificial nanosystems. Both the exogenous and the endogenous siRNAs are loaded in the RISC complex, with the help of which only the guide strand is kept for further interaction with its corresponding mRNA.
Abbreviations: dsRNA, double-stranded RNA; mRNA, messenger RNA; OSCC, oral squamous cell carcinoma; PEG, polyethylene glycol; RISC, RNA-induced silencing complex; siRNA, small interfering RNA.
In this review, we analyze the main challenges associated with siRNA therapy, as a modern and effective way of reducing cell proliferation, invasion, metastasis, increasing immune response targeting invasion/migration, preventing metastasis, or even sensitizing cancer cells to chemotherapy. The siRNA therapeutic approach is summarized in two main categories: classical Lipofectamine-transfected siRNAs and nanoparticle-loaded siRNAs as single or multiple drug delivery.
Overcoming the challenges of siRNA therapy
The main challenges associated with siRNA therapy are represented by the sequence design, the high instability of RNAs, the danger of nuclease degradation, endosomal entrapment, and also off-target effects,Citation29 aspects summarized in .
Figure 3 An efficient therapeutic siRNA design has to consider: 1) the siRNA sequence design; 2) modification made to the sequence as to escape the blood nuclease; 3) an efficient nanoparticle size range essential for efficient siRNA delivery; and 4) the off-target effects exogenous siRNA.
Abbreviations: 2′-FANA, 2′-deoxy-2′-fluoro-beta-D-arabinonucleic acid; miRNA, microRNAs; PKR, protein kinase R; RIG-I, retinoic acid-inducible gene I; RISC, RNA-induced silencing complex; siRNA, small interfering RNA; TLR, Toll-like receptor.
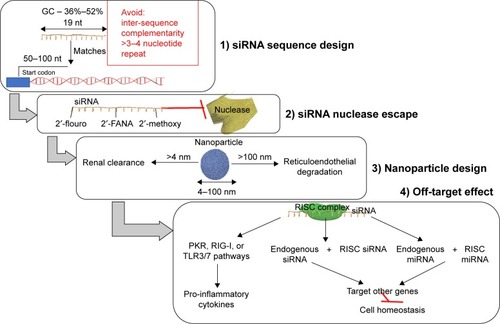
When designing an siRNA sequence, the first step consists in the selection of the targeted gene. The targeted region is composed of 50–100 nucleotides upstream from the start codon, and the single nucleotide polymorphism containing areas are excluded. The recommended length of the siRNA sequence is 19 nucleotides and the guanine–cytosine content should be between 36% and 52%.Citation29,Citation30 The antisense strand has to be enriched in A/U nucleotides at the 5′ end, while the sense strand is advised to contain more G/C nucleotides at the 5′ end.Citation29,Citation31 Inter-sequence complementarity and >3 or 4 repeats of the same nucleotide should be avoided, as it causes changes in the secondary structure of siRNA.Citation30,Citation31
In the blood, there are nucleases that degrade nucleic acids and are characterized by an increased serum half-life.Citation32 The chemical structure of siRNA undergoes some modification in order to be protected particularly from the action of RNA nucleases. Some examples of these modifications are: the insertion of ribonucleic analogs – 2′-flouro, 2′-FANA, 2′-methoxy; 5′-/3′-terminal inverted abasic end caps – or algorithms predicting the nuclease cleaved sites and modifying these with 2′-methoxy.Citation33
Particle size control in delivery represents an important issue. If the nanoparticles are smaller than 4 nm, they are rapidly cleared from the blood via renal filtration and urinary excretion.Citation34 At the same time, if they are larger than 100 nm in diameter, they will be degraded in the reticuloendothelial system and are difficult to be internalized.Citation35 Lipid nanosystems have a long circulation time in the blood and can easily modify their surface, but sometimes they can cause pseudo-allergic reactions. Neutral liposomes have a size of 30–40 nm and are the most commonly used lipid-based nanosystems.Citation36
siRNA therapy is well-tolerated, but in some cases, particularly for the case of transcripts with >23 nucleotides, interferon responses can be activated together with the induction of cell death within in vitro systems. The immune reaction is different, dependent of the type of cells used, and it is hard to predict the in vivo responses.Citation37
Off-target effects are very frequently retrieved in practice. The siRNAs are loaded on the RISC complex and can sometimes target other genes than the intended ones, by an imperfect match. The siRNAs can also be incorporated into dendritic cells and initiate an immune response,Citation38 by stimulating the protein kinase R, retinoic acid-inducible gene I, or Toll-like receptor 3 or 7 pathways for production of pro-inflammatory cytokines.Citation39 Through the off-target effect, the therapeutic siRNA can also impair the endogenous siRNA and the endogenous miRNAs to exert their normal function, ultimately, interfering with the homeostatic state of the cell.Citation38
The trafficking of siRNAs from endosomes into the cytoplasm constitutes a main rate-limiting issue for many delivery approaches.Citation40 siRNA is recognized by surface proteins and internalized into an early endosome that fuses with a sorting endosome and transfers its contents into a late endosome. The late endosome interacts with the lysosome, the pH drops to 4.5, and the siRNA sequence is degraded. The siRNA molecules can form a complex with a special class of lipids and then form a micelle-like structure (called phosphatidylethanolamines [PEs]). The lipoplex fuses rapidly with the anionic membranes, and the content of the endosome is released in the cytoplasm, before lysosomal interaction.Citation40 Lipofectamine, a cationic liposome, used very often in siRNA transfection studies in vitro is a PE-based liposome ().Citation41 Another class of proteins associated with siRNAs are the viroporins, proteins of viral origin that have the ability of forming pores in a lipid bilayer.Citation40 The influenza virus peptide, termed 599, combined with cancerous inhibitor of protein phosphatase 2A (CIP2A) siRNA proved to be able to overcome endosomal entrapment and effectively silenced the CIP2A gene in vitro in the CAL 27 cell line.Citation42
Table 1 Studies using siRNA-targeted delivery as treatment strategies in oral cancer
In order to overcome all these challenges related to siRNAs, several databases were developed. This is the case of siRNAdb, a complex database that furnishes a gene-centric view of siRNA experimental data, providing information related to the siRNA sequence, thermodynamic characteristics, and potential off-target effects.Citation43 siDRM is one of the most promising tools for the design of effective siRNAs.Citation44 RFRCDB-siRNA is another online tool for the design of siRNAs based on the random forest regression model connected with database searching.Citation45 Without any doubt, RNAcentral is the most complex database that has joined 12 new resources and started the importing of new types of data for ncRNAs,Citation46 such as nucleotides modification furnished from MODOMICS and Protein Data Bank, offering also free access.Citation44
Nano-delivery systems for siRNA
The plasma membrane represents a physical barrier for siRNA uptake, due to the high hydrophobicity and negative charge, blocking the crossing of the biological barrier. The delivery methods need to counterwork this limitation and to assist cellular uptake.Citation40
Naked siRNA have a very reduced internalization rate; therefore, in order to increase the therapeutic efficacy, often siRNA is loaded on a carrier alone or in tandem with other therapeutic systems. Also, siRNA may be conjugated with fluorescein isothiocyanate, in order to emit fluorescence, being easily monitored for cellular localization.Citation47
An effective delivery system should be stable at the body temperature and pH variations, have an endocytosis promoting shape, should not be toxic, should exhibit high siRNA loading abilities, and have a size that avoids rapid renal and hepatic clearance.Citation48
All the delivery systems developed for gene therapy can be adapted for siRNA delivery.Citation49 siRNA delivery strategies that use viral particles have a constitutive effect while different nanostructures have only a transitory effect, implying multiple administration doses.Citation36
The viral delivery of siRNA is composed of two main strategies: siRNA is either chemically synthesized and loaded into a viral capsule or it can be expressed from the DNA of a recombinant virus. The major disadvantages of viral siRNA delivery are lower targeted delivery of a specific cell in vivo, the host immune reaction, and the danger of oncogenic transformation of the virus.Citation50 The multidrug resistance in oral cancer can be surpassed through the delivery of siRNA against the multidrug resistance (MDR) gene in adeno-associated viral vector. AAV-siMDR25mer and AAV-siMDR28mer inhibited the MDR1 and MDR2 gene expression and reduced the level of P-glycoprotein, expressed by the MDR1 gene, in KB cells.Citation51 The viral delivery of interference RNA, although highly efficient, presents many challenges when delivered to a living organism, because it causes an inflammatory response following the recognition of the dsRNA longer than 30 nucleotides as nonself.Citation52
Nonviral delivery systems for siRNA are far more diverse and are proposed more often in recent studies.Citation52 These contain several subtypes, among which are liposomes, peptides, immunoliposomes, purified collagen,Citation53 dendrimers,Citation54 gold nanorods,Citation55 carbon nanotubes,Citation56,Citation57 and RNAi microsponges.Citation58 In general, the nanoparticles are PEGylated to mask the unspecific immune response, although it has been proven that the immune system can produce polyethylene glycol (PEG) antibodies.Citation59 The most frequently tested nonviral carriers are liposomes, which have again presented the major challenge of immune reaction.Citation53 The main problems related to siRNA-based drugs are represented by high toxicity.Citation60 The comprehension and elimination of these issues can be considered a major constituent in the progress of safe and effective siRNA therapeutic systems.Citation61
For effective delivery of therapeutic siRNA, the transcripts can be conjugated with cholesterol reaching better tumor retention in vivo. However, the siRNA-cholesterol complex is effective when administered in the tumor proximity; hence, it is unsuitable for systemic delivery.Citation62
Aptamers are oligonucleotides or peptides that target specific molecules and by being bound to siRNA enhance their specificity.Citation48 A nucleotide aptamer selected with the use of Systematic Evolution of Ligands by Exponential Enrichment technology was internalized in the HPV16-infected tonsil epithelial cells more efficiently than in the noninfected cells;Citation63 it was proposed to be conjugated with E6 oncogene targeting siRNA.Citation64 The cationic cell penetrating peptides facilitate endosomal escape or mediate endosomal-free entry into the cell without any need of modifying the siRNA structure.Citation65
The polymers are positively charged; when they are conjugated with the negatively charged nucleic acids, they cause the nucleic acids to condensate and be protected.Citation48 A polymeric nanoparticle galactose-modified trimethyl chitosan-cysteine loaded with vascular endothelial growth factor (VEGF)-targeted siRNA was used for increased specificity and efficiency in inhibiting oral cancer cell proliferation in vitro and tumor growth in vivo.Citation49 Dendrimers are also artificial polymers that have a positive charge and are highly branched at the end-chains.
Polyamidoamine (PAMAM) is a dendrimer that, when conjugated with siRNA, can have both nuclear and perinuclear localization.Citation48 The shRNA-mediated silencing of human telomerase reverse transcriptase loaded on PAMAM is a more potent knockdown method with an increased therapeutic efficacy.Citation66
The targeted delivery of a complex composed of CIP2A siRNA, EGFR binding-peptide, and an endosome-disruptive peptide was proven to silence more efficiently the CIP2A oncogene, in vitro in the oral squamous cell carcinoma (OSCC) cell lines CAL 27 and squamous cell carcinoma (SCC)-15 and in vivo in xenografted mouse tumors of transfected CAL 27 cells.Citation28 Recently, it was proven that the co-delivery of doxorubicin and MDR1 siRNA by using polymer polyethylenimine was related to an increased response to treatment, demonstrated both in vitro and in vivo. More details over the targeted delivery of siRNA as a therapeutic option in oral cancer are provided in .
Therapeutic assessment of siRNA in oral cancer
The essential characteristic of siRNA is its highly specific targeting of a single gene transcript. The various studies that use siRNA in cancer focus on the identification of a particular gene function or on the development of new therapies by silencing the overexpressed genes with oncogenic role ().
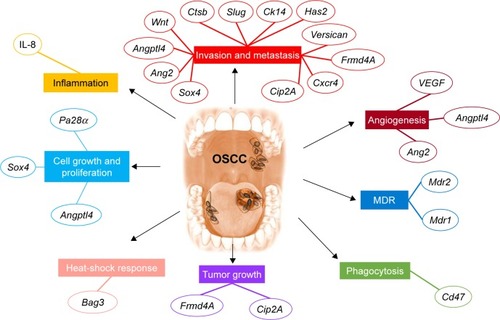
Figure 4 The main pathway targeted using siRNA delivery systems in oral cancer.
Notes: Examples of siRNA therapeutics alter multiple hallmarks applied for oral cancer. Cell growth and proliferation are impaired by siRNA for Pa28α, Sox4, or Angptl4. Local inflammation is reduced by IL-8. Invasion and metastasis are decreased by siRNA for Sox4, Ang2, Angptl4, Wnt, Ctsb, Slug, Ck14, Has2, Versican, Frmd4A, Cxcr4, and Cip2A. Angiogenesis is inhibited by siRNA targeting VEGF, Angptl4, and Ang2. MDR is inhibited by siRNA for Mdr1 and Mdr2. Phagocytosis is stimulated by siRNA for Cd47. Tumor growth is reduced in the presence of siRNA for Cip2A and Frmd4A. The heat-shock response is decreased by siRNA for Bag3.
Abbreviations: IL-8, interleukin-8; MDR, multidrug resistance; siRNA, small interfering RNA; VEGF, vascular endothelial growth factor.
Recently, the identification of subcellular targets in oral cancer cells guided the well-founded progression in the field of “siRNA-targeted therapy.” These newly designed siRNA systems are directed to target cellular components involved in signal transduction pathways that control epigenetics effectors, cell proliferation, cell cycle, apoptosis, immune response, or angiogenesis.Citation20,Citation70–Citation72
The balance of histone acetylation and deacetylation is an important regulator of epigenetic pathways, affecting the mechanisms of gene expression regulation.Citation73 Alteration or mutation within the histone deacetylase 8 (HDAC8) is associated with tumor progression caused by induction of aberrant transcription factors that regulate important cellular function.Citation73 HDAC8 was found to be overexpressed in OSCC cancer cells. By targeting HDAC8 with siRNA, the expression of this enzyme was reduced, leading to reduced proliferation and increased apoptosis within the cells.Citation74 After silencing the Frmd4A gene, the CAL 27 cells remained in the G0 phase with impaired proliferative, invasive, and migration capacities.Citation75
Molecular analysis showed that a number of genes were affected by the treatment and that these genes were involved in the cell cycle, in the following signaling pathways: JAK/STAT, p53, and extracellular matrix proteins.Citation67
SOX4 is an important regulator of the oncogenic PI3K/AKT and MAPK signaling pathways, which sustains cell proliferation and differention.Citation76,Citation77 SOX4 functions as an oncogene in the OSCC event described after siRNA treatment, followed by a decline in the proliferation, invasion, and migration of UM1 cancer cells.Citation78 A reduction in the mutant p53 expression caused by its corresponding siRNA increased the apoptotic and necrotic SCC-4 cells and also lowered their metastasis capability.Citation79
Previous studies reveal the complex effects of immune response mechanisms in OSCC.Citation80–Citation83 IL-8 is an important pro-inflammatory interleukin, involved in head and neck SCC progression, since its knockdown by an analog antisense ncRNA affected the cellular viability and the colony formation capacity mediated by NOD signaling pathway.Citation81 IL-8 was also found to be overexpressed in the saliva of OSCC patients, when compared to healthy subjects and patients with precancerous lesions.Citation82
siRNA knockdown of C-X-C chemokine receptor type 4 (Cxcr4) in OSCC decreased cell proliferation, while increasing the apoptosis rate and causing cell cycle arrest. Also, Cxcr4 supports epithelial to mesenchymal transition (EMT) progress and lymph node metastasis in tongue squamous cell carcinoma (TSCC); thus, the silencing of this gene by its corresponding siRNA led to the regression of these events in vitro as well as in xenografted tumors in mice.Citation83
Cytokeratin 14 (Ck14) is a cytoskeletal component of the epithelial tissue and an important promoter of local invasion and metastasis. The Ck14 siRNA resulted in a 80% drop in the collective invasion of cells belonging to a three-dimensional salivary adenoid cystic carcinoma (SACC) culture.Citation84 Hyaluronic synthases 2 (Has2) is overexpressed only in cancer-associated fibroblasts (CAFs) and not in normal fibroblasts. When this gene is knocked down by one of its two specific siRNAs in CAFs, and then the CAFs co-cultured with oral OSCC cells, the cancer invasion and migration abilities were decreased. This process is mediated by increased expression level of matrix metalloproteinase (MMP)-1 and MMP-3 and decreased expression of tissue inhibitor of metalloproteinase 1 and MMP-10.Citation85
In some cases, multiple siRNAs can be used as a way of following the interaction of multiple elements from a signaling pathway. The regulation of EMT, a key process due to the fact that oral cancer cells are characterized by high epithelial plasticity, can be exploited to develop novel targeted therapies. EMT is related to inhibition/loss of E-cadherin expression, reduction of cellular adhesion, a fact that promotes invasion/migration, and finally leads to metastatic disease.Citation86 For instance, by knocking down the MMP-10 and the Slug gene, it was proven that transforming growth factor β induces EMT and invasion of human oral squamous cell carcinoma (HSC)-4 cells, through Slug-dependent activation of MMP-10.Citation87
Sometimes, siRNA can be used to study the role of a gene in OSCC pathology. The knocking down of Cdh11 gene enhanced the proliferation and invasion of UM-SCC-29 and UM-SCC-47 cells, thus providing proof of its function as a tumor suppressor.Citation88 The methods by which the above-mentioned siRNA effects were determined are listed in .
Table 2 Relevant studies of siRNA as a silencing mechanism in OSCC using commercial delivery systems used to furnish new mechanistic insights
siRNA was used to target angiopoietin-like protein 4 mRNA in TSCC cells, as a way of inhibiting its role in cellular growth and colony formation, with the perspective of improving the survival rate of patients.Citation89 Following siRNA silencing of Ang2, the expression of Vimentin, Snail, and Twist genes was downregulated at the same time as E-cadherin was upregulated. Thereby, the TCA8113 cells would present a reduction in their capacity to invade and migrate through the EMT process.Citation90 The same pattern was observed in the case of siRNA inhibition of superoxide dismutase 2 in SACC-LM that led to E-cadherin increased levels as well as the decreased expression of migration-associated genes Vimentin, Slug, Snail, p-ERK, and MMP-2.Citation91
Aberrant Notch4 expression has an essential function in OSCC tumorigenesis and promotes metastasis by perineural spread,Citation92 becoming an important prognosis marker and therapeutic target.Citation92 Notch4 siRNA caused the 1.4-fold drop in proliferation and 1.9-fold drop in cell motility, after 48-hour post-transfection in an oral cancer cell line, HSC-3.Citation92
Combination of siRNA with other therapies
Nano-delivery of WNT siRNA in KB cells interrupted the WNT, β-catenin, and Vimentin pathway, hindering mesenchymal transition and by including photodynamic therapy, it could advance into a novel combinatorial therapeutic strategy.Citation97 The combination of plasmonic photothermal therapy (PPTT) and heat-shock protein (HSP) siRNA is another instance of this kind of approach. By removing the protective role of Hsp70, which opposes heat-induced aggregation of proteins, thereby preventing cell death, these ncRNAs enhance the beneficial effects of PPTT.Citation98 An important remark is the fact that a few years ago it was discovered that the KB cells have probably been contaminated with the HeLa cell line of cervical cancer, since it shows high genetic similarity with it; the controversy still remains around this cell line, because phenotypically it maintained the same morphology as the originally isolated cells of OSCC.Citation99 Hence, studies using the KB cell line as oral cancer model should be regarded with a rational trace of doubt regarding its application in oral cancer.
Nanoparticles composed of PEG, chlorine e6, and polyethylenimine loaded with WNT siRNA decrease the in vitro viability and invasion capacity of KB cells and the effects were even more pronounced when gene silencing was combined with photodynamic therapy, thus proving the fact that combinatorial therapy is still the best option.Citation100 Similar results were obtained in xenografted TSCC cell lines, SCC-4 and SAS treated with calcium phosphate lipid nanoparticles containing VEGF siRNA, in which the tumor growth was reduced due to efficient silencing of the pro-angiogenic factor VEGF.Citation101 The same type of nanoparticles combined with photodynamic therapy and loaded with HIF1α siRNA again proved to be more effective than single therapy in SCC-4 and SAS cells in vitro and in xenografted tumors in vivo.Citation102 siRNA-mediated silencing of Bag3 is more efficient in vivo in xenografted tumors of CAL 27 cells, by combining ncRNA with gold nanorods and applying photodynamic therapy.Citation55 A summary of the data and details are provided in .
Table 3 Studies using siRNA in combination with other therapies in oral cancer
Clinical trial in oral cancer using siRNA therapy
The main challenge for siRNA delivery is the proper choice of a delivery route. In the clinical trials listed for therapeutic delivery systems in oral cancer, the majority of them (three out of four) use the adenovirus injected locally as a carrier of the therapeutic agent, while a single clinical trial loaded lipid nanoparticles with the siRNA for p53, which are again injected locally.
A way of preventing transformation of premalignant oral lesions may be the local delivery of the adenoviral containing wild-type p53 gene, the Ad5CMV-p53 gene. In a clinical trial first proposed in 2006, the injection of oral lesion of Ad5CMV-p53 was followed by a rinse of the mouth. It was demonstrated that Ad5CMV-p53 is safe and can be repetitively administered for improvement of the therapeutic efficacy, lacking signs of severe toxicity. Its presence in the circulation of the adenovirus at 24 hours posttreatment, and the presence of p53 transgene in the tumor tissue were demonstrated. This certifies the utility of this therapeutic system for the mutated p53 gene,Citation103 being well-tolerated by patients with oral cancers.Citation104
The study was terminated and another was performed.Citation105,Citation106 siRNA delivery was carried out alone, before surgical tumor resection or in combination with standard chemotherapy, in order to observe how it would affect the survival rate of patients with advanced oral or maxillofacial malignancies.Citation107 The antisense DNA of EGFR gene was delivered in lipid nanoparticles, in a clinical trial for advanced SCC of the head and neck,Citation108,Citation109 being well-tolerated, showing only reduced inflammation at the injection site.Citation109
siRNA therapies versus novel genomic editing tools
RNAi is a natural mechanism that has the capacity to regulate gene expression via an endogenous systematic system with effects on the expression level of a particular phenotype. It still remains an important technology for the study of gene function despite several disadvantages related to the hypomorphic phenotypes, some inconsistencies in complete loss-of-function and also due to genetic mutation.Citation110
A basically distinct method to regulate gene function was developed based on the comprehension of the mechanism of action of the programmable DNA-binding proteins, retrieved in the literature as transcription activator-like effectors nucleases (TALEN).Citation110 Another important tool for genome editing is represented by the clustered regularly interspaced short palindromic repeats (CRISPR)/Cas system.Citation110–Citation112 CRISPR/Cas9 is currently the most popular system used and is based on a programmable DNA nuclease similar to TALEN.Citation112
In contrast to RNAi, CRISPR or TALEN seem to be able to have a consistent and robust knockdown effect upon the targeted DNA,Citation110 but the main problem related to these two approaches consists in the possible mutagenic effect achieved during procedure, an effect that is maintained persistently in the DNA sequence.
The subject regarding the advantages and disadvantages between the newly implemented technologies, and RNAi is currently under major debate; however, the RNAi strategy has several advantages that make this approach a strong contender in different experimental settings. One of these advantages consists in the simplicity of the method, in that RNAi does not require additional elements for the generation of knockdowns. The fact that the majority of the targeted cells already contain the RNAi silencing machinery, that is actually the full equipment necessary, makes the implementation of such strategies easier and also more cost efficient. Moreover, RNAi targets the transcriptomic landscape and not the genomic DNA sequences, such as CRISPR or TALEN; therefore, the accessibility to the targeted sequence is not hampered by the chromatin state.Citation110 Also, RNAi does not target the transcription start site (TSS), an act that facilitates the use of the technology in organisms without annotated TSSs.Citation113
Conclusion and perspectives
siRNA therapeutic development in oral cancer has to overcome some major challenges related especially to the delivery methods and potential off-target effects. Also, the danger of being degraded by bloodstream nucleases, loaded onto the reticuloendothelial system, endosomal escape, avoidance of the immune response, and RISC interaction are important aspects to be considered in the future.
The siRNA delivery issue can be overcome through chemical modification of the exogenous sequence, loading of the particle in a viral capsule, or loading siRNA on different types of nanoparticles.
Often, siRNA is used as a means of evaluating the function of a particular gene, for the study of particular molecular mechanisms. During the first evaluations of the therapeutic potential of siRNA against certain oral cancer tumor promoting genes: CIP2A, CXCR4, VEGF, MDR1, and MDR2, specific cancer hallmarks were targeted. One step further in the development of siRNA therapeutics is the concept of combining siRNA therapy with other therapies such as photodynamic therapy or PPTT. Lipid nanoparticles targeting EGFR were proved to have a significant therapeutic potential, along with a clinical trial for the silencing of the p53 gene exploiting RNAi mechanisms. These two clinical trials have demonstrated only minimal drawbacks of using a more targeted approach on oral cancer malignancies and opened the path for a brighter future of oral cancer therapies.
Disclosure
The authors report no conflicts of interest in this work.
References
- D’SouzaGMcNeelTSFakhryCUnderstanding personal risk of oropharyngeal cancer: risk-groups for oncogenic oral HPV infection and oropharyngeal cancerAnn Oncol201728123065306929059337
- WarnakulasuriyaSGlobal epidemiology of oral and oropharyngeal cancerOral Oncol2009454–530931618804401
- FerlayJSoerjomataramIDikshitRCancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012Int J Cancer20151365E359E38625220842
- PettiSLifestyle risk factors for oral cancerOral Oncol2009454–534035018674956
- IrimieAIBraicuCCojocneanu-PetricRBerindan-NeagoeICampianRSNovel technologies for oral squamous carcinoma biomarkers in diagnostics and prognosticsActa Odontol Scand201573316116825598447
- PileczkiVBraicuCGhermanCDBerindan-NeagoeITNF-alpha gene knockout in triple negative breast cancer cell line induces apoptosisInt J Mol Sci201214141142023263670
- WangKCChangHYMolecular mechanisms of long noncoding RNAsMol Cell201143690491421925379
- SetoyamaTLingHNatsugoeSCalinGANon-coding RNAs for medical practice in oncologyKeio J Med201160410611322200634
- Berindan-NeagoeICalinGAMolecular pathways: microRNAs, cancer cells, and microenvironmentClin Cancer Res201420246247625325512634
- Berindan-NeagoeIMonroig PdelCPasculliBCalinGAMicroRNAome genome: a treasure for cancer diagnosis and therapyCA Cancer J Clin201464531133625104502
- AlexanderRPFangGRozowskyJSnyderMGersteinMBAnnotating non-coding regions of the genomeNat Rev Genet201011855957120628352
- FrancastelCHubéFManiSSantulliGTaubeJSzweykowska-KulinskaZThe non-coding RNA journal club: highlights on recent papers – 2Noncoding RNA201512167
- CalinGVazquezFWelcome to the new journal Non-Coding RNA!Noncoding RNA20151113
- LingHVincentKPichlerMJunk DNA and the long non-coding RNA twist in cancer geneticsOncogene201534395003501125619839
- ZaharieFMuresanMSPetrushevBExosome-carried microRNA-375 inhibits cell progression and dissemination via Bcl-2 blocking in colon cancerJ Gastrointestin Liver Dis201524443544326697569
- FerracinMGautheretDHubéFThe non-coding RNA journal club: highlights on recent papersNoncoding RNA20151187
- AmaralPPMattickJSNoncoding RNA in developmentMamm Genome2008197–845449218839252
- MunkerRCalinGAMicroRNA profiling in cancerClin Sci (Lond)2011121414115821526983
- CatanaCSPichlerMGiannelliGMaderRMBerindan-NeagoeINon-coding RNAs, the Trojan horse in two-way communication between tumor and stroma in colorectal and hepatocellular carcinomaOncotarget2017817295192953428392501
- BraicuCCatanaCCalinGABerindan-NeagoeINCRNA combined therapy as future treatment option for cancerCurr Pharm Des201420426565657425341933
- Pop-BicaCGuleiDCojocneanu-PetricRBraicuCPetrutBBerindan-NeagoeIUnderstanding the role of non-coding RNAs in bladder cancer: from dark matter to valuable therapeutic targetsInt J Mol Sci2017187E151428703782
- BraicuCCojocneanu-PetricRChiraSClinical and pathological implications of miRNA in bladder cancerInt J Nanomedicine20151079180025653521
- IrimieAIBraicuCSoneaLA looking-glass of non-coding RNAs in oral cancerInt J Mol Sci20171812E262029206174
- HamiltonAJBaulcombeDCA species of small antisense RNA in posttranscriptional gene silencing in plantsScience1999286544195095210542148
- CarthewRWSontheimerEJOrigins and mechanisms of miRNAs and siRNAsCell2009136464265519239886
- DanaHChalbataniGMMahmoodzadehHMolecular mechanisms and biological functions of siRNAInt J Biomed Sci2017132485728824341
- LamJKChowMYZhangYLeungSWsiRNA versus miRNA as therapeutics for gene silencingMol Ther Nucleic Acids20154e25226372022
- Alexander-BryantAAZhangHAttawayCCDual peptide-mediated targeted delivery of bioactive siRNAs to oral cancer cells in vivoOral Oncol20177212313128797448
- FakhrEZareFTeimoori-ToolabiLPrecise and efficient siRNA design: a key point in competent gene silencingCancer Gene Ther2016234738226987292
- TilesiFFradianiPSocciVWillemsDAscenzioniFDesign and validation of siRNAs and shRNAsCurr Opin Mol Ther200911215616419330721
- TaferHBioinformatics of siRNA designMethods Mol Biol2014109747749024639173
- HickersonRPVlassovAVWangQStability study of unmodified siRNA and relevance to clinical useOligonucleotides200818434535418844576
- ChernolovskayaELZenkovaMAChemical modification of siRNACurr Opin Mol Ther201012215816720373259
- ChoiHSLiuWMisraPRenal clearance of nanoparticlesNat Biotechnol200725101165117017891134
- MieleESpinelliGPMieleENanoparticle-based delivery of small interfering RNA: challenges for cancer therapyInt J Nanomedicine201273637365722915840
- IrimieAISoneaLJurjAFuture trends and emerging issues for nanodelivery systems in oral and oropharyngeal cancerInt J Nanomedicine2017124593460628721037
- GavrilovKSaltzmanWMTherapeutic siRNA: principles, challenges, and strategiesYale J Biol Med201285218720022737048
- JacksonALLinsleyPSRecognizing and avoiding siRNA off-target effects for target identification and therapeutic applicationNat Rev Drug Discov201091576720043028
- MengZLuMRNA interference-induced innate immunity, off-target effect, or immune adjuvant?Front Immunol2017833128386261
- DominskaMDykxhoornDMBreaking down the barriers: siRNA delivery and endosome escapeJ Cell Sci2010123Pt 81183118920356929
- DalbyBCatesSHarrisAAdvanced transfection with Lipofectamine 2000 reagent: primary neurons, siRNA, and high-throughput applicationsMethods20043329510315121163
- CantiniLAttawayCCButlerBAndinoLMSokoloskyMLJakymiwAFusogenic-oligoarginine peptide-mediated delivery of siRNAs targeting the CIP2A oncogene into oral cancer cellsPLoS One201389e7334824019920
- ChalkAMWarfingeREGeorgii-HemmingPSonnhammerELLsiRNAdb: a database of siRNA sequencesNucleic Acids Res200533Database issueD131D13415608162
- GongWRenYZhouHWangYKangSLiTsiDRM: an effective and generally applicable online siRNA design toolBioinformatics200824202405240618718944
- JiangPWuHDaYRFRCDB-siRNA: improved design of siRNAs by random forest regression model coupled with database searchingComput Methods Programs Biomed200787323023817644215
- RNAcentral ConsortiumRNAcentral: an international database of ncRNA sequencesNucleic Acids Res201543Database issueD123D12925352543
- OckerMNeureiterDLuedersMVariants of bcl-2 specific siRNA for silencing antiapoptotic bcl-2 in pancreatic cancerGut20055491298130816099798
- TatipartiKSauSKashawSKIyerAKsiRNA delivery strategies: a comprehensive review of recent developmentsNanomaterials (Basel)201774E7728379201
- HanLTangCYinCOral delivery of shRNA and siRNA via multifunctional polymeric nanoparticles for synergistic cancer therapyBiomaterials201435154589460024613049
- OliveiraSStormGSchiffelersRMTargeted delivery of siRNAJ Biomed Biotechnol2006200646367517057365
- XuDMcCartyDFernandesAFisherMSamulskiRJJulianoRLDelivery of MDR1 small interfering RNA by self-complementary recombinant adeno-associated virus vectorMol Ther200511452353015771955
- AkhtarSBenterIFNonviral delivery of synthetic siRNAs in vivoJ Clin Invest2007117123623363218060020
- LiSDHuangLGene therapy progress and prospects: non-viral gene therapy by systemic deliveryGene Ther200613181313131916953249
- BiswasSTorchilinVPDendrimers for siRNA deliveryPharmaceuticals (Basel)20136216118324275946
- WangBKYuXFWangJHGold-nanorods-siRNA nanoplex for improved photothermal therapy by gene silencingBiomaterials201678273926646625
- NeagoeIBBraicuCMateaCEfficient siRNA delivery system using carboxilated single-wall carbon nanotubes in cancer treatmentJ Biomed Nanotechnol20128456757422852466
- GhermanCTudorMCConstantinBPharmacokinetics evaluation of carbon nanotubes using FTIR analysis and histological analysisJ Nanosci Nanotechnol20151542865286926353506
- LeeJBHongJBonnerDKPoonZHammondPTSelf-assembled RNA interference microsponges for efficient siRNA deliveryNat Mater201211431632222367004
- YangQLaiSKAnti-PEG immunity: emergence, characteristics, and unaddressed questionsWiley Interdiscip Rev Nanomed Nanobiotechnol20157565567725707913
- JudgeAMacLachlanIOvercoming the innate immune response to small interfering RNAHuman Gene Ther200819211112418230025
- RobbinsMJudgeAMacLachlanIsiRNA and innate immunityOligonucleotides20091928910219441890
- ChernikovIVGladkikhDVMeschaninovaMICholesterol-containing nuclease-resistant siRNA accumulates in tumors in a carrier-free mode and silences MDR1 geneMol Ther Nucleic Acids2017620922028325287
- GourroncFARockeyWMThielWHGiangrandePHKlingelhutzAJIdentification of RNA aptamers that internalize into HPV-16 E6/E7 transformed tonsillar epithelial cellsVirology20134461–232533324074596
- ButzKRistrianiTHengstermannADenkCScheffnerMHoppe-SeylerFsiRNA targeting of the viral E6 oncogene efficiently kills human papillomavirus-positive cancer cellsOncogene200322385938594512955072
- WangJLuZWientjesMGAuJLDelivery of siRNA therapeutics: barriers and carriersAAPS J201012449250320544328
- LiuXHuangHWangJDendrimers-delivered short hairpin RNA targeting hTERT inhibits oral cancer cell growth in vitro and in vivoBiochem Pharmacol2011821172321453684
- YuTWuYHuangYRNAi targeting CXCR4 inhibits tumor growth through inducing cell cycle arrest and apoptosisMol Ther201220239840722108861
- XuLYeudallWAYangHFolic acid-decorated polyamidoamine dendrimer exhibits high tumor uptake and sustained highly localized retention in solid tumors: its utility for local siRNA deliveryActa Biomater20175725126128438704
- WangDXuXZhangKCodelivery of doxorubicin and MDR1-siRNA by mesoporous silica nanoparticles-polymerpolyethylenimine to improve oral squamous carcinoma treatmentInt J Nanomedicine20181318719829343957
- BraicuCPileczkiVIrimieABerindan-NeagoeIp53siRNA therapy reduces cell proliferation, migration and induces apoptosis in triple negative breast cancer cellsMol Cell Biochem20133811–2616823881244
- AldeaMCraciunLTomuleasaCRepositioning metformin in cancer: genetics, drug targets, and new ways of deliveryTumour Biol20143565101511024504677
- AldeaMDPetrushevBSoritauOMetformin plus sorafenib highly impacts temozolomide resistant glioblastoma stem-like cellsJ BUON201419250251124965413
- RoperoSEstellerMThe role of histone deacetylases (HDACs) in human cancerMol Oncol200711192519383284
- AhnMYYoonJHHistone deacetylase 8 as a novel therapeutic target in oral squamous cell carcinomaOncol Rep201737154054628004115
- ZhengXJiaBLinXFRMD4A: a potential therapeutic target for the treatment of tongue squamous cell carcinomaInt J Mol Med20163851443144927666346
- Ramezani-RadPGengHHurtzCSOX4 enables oncogenic survival signals in acute lymphoblastic leukemiaBlood2013121114815523152540
- LiuYCuiLHuangJSOX4 promotes progression in OLP-associated squamous cell carcinomaJ Cancer20167111534154027471569
- GeorgakopoulouEATroupisTGTroupisGGorgoulisVGUpdate of the cancer-associated molecular mechanisms in oral lichen planus, a disease with possible premalignant natureJ Buon201116461361622331711
- IrimieAIBraicuCPileczkiVKnocking down of p53 triggers apoptosis and autophagy, concomitantly with inhibition of migration on SSC-4 oral squamous carcinoma cellsMol Cell Biochem20164191–2758227370646
- WinckFVPrado RibeiroACRamos DominguesRInsights into immune responses in oral cancer through proteomic analysis of saliva and salivary extracellular vesiclesSci Rep201551630526538482
- ChanLPWangLFChiangFYLeeKWKuoPLLiangCHIL-8 promotes HNSCC progression on CXCR1/2-mediated NOD1/RIP2 signaling pathwayOncotarget2016738618206183127557518
- PunyaniSRSathawaneRSSalivary level of interleukin-8 in oral precancer and oral squamous cell carcinomaClin Oral Investig2013172517524
- DuanYZhangSWangLTargeted silencing of CXCR4 inhibits epithelial–mesenchymal transition in oral squamous cell carcinomaOncol Lett20161232055206127602138
- GaoXLWuJSCaoMXCytokeratin-14 contributes to collective invasion of salivary adenoid cystic carcinomaPLoS One2017122e017134128152077
- ZhangZTaoDZhangPHyaluronan synthase 2 expressed by cancer-associated fibroblasts promotes oral cancer invasionJ Exp Clin Cancer Res201635118127884164
- PatilSRaoRSGanaviBSMesenchymal–epithelial transition in oral cancerJ Int Oral Health201579iii
- HinoMKamoMSaitoDTransforming growth factor-beta1 induces invasion ability of HSC-4 human oral squamous cell carcinoma cells through the Slug/Wnt-5b/MMP-10 signalling axisJ Biochem2016159663164026861993
- PiaoSInglehartRCScanlonCSRussoNBanerjeeRD’SilvaNJCDH11 inhibits proliferation and invasion in head and neck cancerJ Oral Pathol Med2017462899727397103
- HuangZXieJLinSThe downregulation of ANGPTL4 inhibits the migration and proliferation of tongue squamous cell carcinomaArch Oral Biol20167114414927505034
- LiCLiQCaiYOverexpression of angiopoietin 2 promotes the formation of oral squamous cell carcinoma by increasing epithelial–mesenchymal transition-induced angiogenesisCancer Gene Ther201623929530227492854
- ChangBYangHJiaoYSOD2 deregulation enhances migration, invasion and has poor prognosis in salivary adenoid cystic carcinomaSci Rep201662591827181103
- MkHPrinceSMohanAMKrishnanKVDeviAAssociation of Notch4 with metastasis in human oral squamous cell carcinomaLife Sci2016156384627197026
- FengXJiangYXieLOverexpression of proteasomal activator PA28α serves as a prognostic factor in oral squamous cell carcinomaJ Exp Clin Cancer Res2016353526892607
- SakakuraKTakahashiHKairaKRelationship between tumor-associated macrophage subsets and CD47 expression in squamous cell carcinoma of the head and neck in the tumor microenvironmentLab Invest2016969994100327322955
- WeiTCongXWangXTInterleukin-17A promotes tongue squamous cell carcinoma metastasis through activating miR-23b/versican pathwayOncotarget2017846663668028035060
- YangWEHoCCYangSFCathepsin B expression and the correlation with clinical aspects of oral squamous cell carcinomaPLoS One2016113e015216527031837
- MaCShiLHuangYNanoparticle delivery of Wnt-1 siRNA enhances photodynamic therapy by inhibiting epithelial–mesenchymal transition for oral cancerBiomater Sci20175349450128070573
- AliMRAliHRRankinCREl-SayedMATargeting heat shock protein 70 using gold nanorods enhances cancer cell apoptosis in low dose plasmonic photothermal therapyBiomaterials20161021827318931
- JiangLZengXWangZChenQCell line cross-contamination: KB is not an oral squamous cell carcinoma cell lineEur J Oral Sci20091171909119196324
- GhahhariNMBabashahSInterplay between microRNAs and WNT/β-catenin signalling pathway regulates epithelial-mesenchymal transition in cancerEur J Cancer201551121638164926025765
- LecarosRLHuangLLeeTCHsuYCNanoparticle delivered VEGF-A siRNA enhances photodynamic therapy for head and neck cancer treatmentMol Ther201624110611626373346
- ChenWHLecarosRLTsengYCHuangLHsuYCNanoparticle delivery of HIF1α siRNA combined with photodynamic therapy as a potential treatment strategy for head-and-neck cancerCancer Lett20153591657425596376
- TolcherAWHaoDde BonoJPhase I, pharmacokinetic, and pharmacodynamic study of intravenously administered Ad5CMV-p53, an adenoviral vector containing the wild-type p53 gene, in patients with advanced cancerJ Clin Oncol200624132052205816648505
- INGN 201: Ad-p53, Ad5CMV-p53, adenoviral p53, p53 gene therapy-introgen, RPR/INGN 201Drugs R D20078317618717472413
- M.D. Anderson Cancer CenterAdenovirus for Oral Premalignancies Available from: https://clinicaltrials.gov/ct2/show/NCT00410865?term=p53&cond=oral+cancer&draw=1&rank=2. NLM identifier: NCT00410865Accessed January 10, 2018
- National Cancer Institute (NCI)Gene Therapy in Preventing Cancer in Patients With Premalignant Carcinoma of the Oral Cavity or Pharynx Available from: https://clinicaltrials.gov/ct2/show/study/NCT00064103?term=p53&cond=oral+cancer&draw=1&rank=3. NLM identifier: NCT00064103Accessed January 10, 2018
- Shenzhen SiBiono GeneTech Co., LtdrAd-p53 Gene Therapy for Advanced Oral and Maxillofacial Malignant Tumors (rAd-p53) Available from: https://clinicaltrials.gov/ct2/show/study/NCT00902083?term=p53&cond=oral+cancer&draw=1&rank=6. NLM identifier: NCT00902083Accessed January 10, 2018
- University of PittsburghGene Therapy in Treating Patients With Advanced Head and Neck Cancer Available from: https://clinicaltri-als.gov/ct2/show/study/NCT00009841?term=gene&cond=Oral+Cancer&draw=3. NLM identifier: NCT00009841Accessed January 10, 2018
- ZengQKanterPMDhirRGoodingWEHuangLGrandisJRLack of toxicity of EGFR antisense gene therapyJ Exp Ther Oncol20022317418612415634
- BoettcherMMcManusMTChoosing the right tool for the job: RNAi, TALEN, or CRISPRMol Cell201558457558526000843
- ChiraSGuleiDHajitouABerindan-NeagoeIRestoring the p53 ‘guardian’ phenotype in p53-deficient tumor cells with CRISPR/Cas9Trends Biotechnol Epub2018222
- GuleiDBerindan-NeagoeICRISPR/Cas9: a potential life-saving tool. What’s next?Mol Ther Nucleic Acids2017933333629246311
- CarninciPSandelinALenhardBGenome-wide analysis of mammalian promoter architecture and evolutionNat Genet200638662663516645617
