Abstract
Purpose
Long noncoding RNAs (lncRNAs) are known to function as regulators in the development and occurrence of various tumors. MALAT1 is a highly conserved lncRNA and has vital functions in diverse tumors, including pancreatic cancer (PC). However, the underlying molecular regulatory mechanism involved in the occurrence and development of PC remains largely unknown. Thus, it is important to explore MALAT1 in PC and elucidate its function, which might offer a new perspective for clinical diagnosis and therapy.
Methods
First, we used the Gene Expression Omnibus, Oncomine, and The Cancer Genome Atlas databases to determine the clinical diagnostic and prognostic values of MALAT1. We next used our own GeneChip and The Cancer Genome Atlas database to collect the possible target genes of MALAT1 and further utilized a bioinformatics analysis to explore the underlying significant pathways that might be crucial in PC. Finally, we identified several key target genes of MALAT1 and hope to offer references for future research.
Results
We found that the expression of MALAT1 was significantly elevated in patients with PC. A receiver operating characteristics curve analysis showed a moderate diagnostic value (area under the curve =0.75, sensitivity =0.66, specificity =0.72). A total of 224 important overlapping genes were collected, and six hub genes (CCND1, MAPK8, VEGFA, FOS, CDH1, and HSP90AA1) were identified, of which CCND1, MAPK8, and VEGFA, are important genes in PC. Several pathways, including the mTOR signaling pathway, pathways in cancer, and the MAPK signaling pathway, were suggested to be the vital MALAT1 pathways in PC.
Conclusion
MALAT1 is suggested to be a promising diagnostic biomarker in PC. Six hub genes (CCND1, MAPK8, VEGFA, FOS, CDH1, and HSP90AA1), and specifically CCND1, MAPK8, and VEGFA, might be key MALAT1 target genes in PC. Due to their possible clinical significance in PC, several pathways, such as the mTOR signaling pathway, pathways in cancer, and the MAPK signaling pathway, are worthy of further study.
Introduction
Pancreatic cancer (PC) is the seventh most common cause of cancer mortality and causes 330,000 deaths per year worldwide, accounting for approximately 4.0% of all cases of cancer.Citation1 Currently, while the incidence of PC tends to linearly increase, the mortality of PC is high, and the prognosis is dismal. It is estimated that patients who present with early disease and have a negative resection margin only have a 5-year survival rate of 24%.Citation2 There is an even higher mortality for unresectable patients. Currently, PC lacks a sensitive diagnosis marker to monitor the disease at an early stage and to predict the prognosis. Therefore, PC is in desperate need of a new index.
Long noncoding RNAs (lncRNAs) do not encode proteins and were once regarded as transcriptional noise or cloning artifacts.Citation3,Citation4 According to recent research, lncRNAs are thought to be regulators of cancer development, which make them a promising target for cancer treatment.Citation5,Citation6 MALAT1, with a length of over 8,000 nucleotides, is a highly conserved lncRNA that is derived from chromosome 11q13. It controls cancer cell proliferation, differentiation, apoptosis, and invasion by regulating the synthesis of proteins and the expression of genes. MALAT1 is well known as the first lncRNA, indicating the poor prognosis of non-small-cell lung cancer.Citation7 Currently, emerging research has shown that MALAT1 plays a crucial role in ovarian cancer,Citation8 esophageal cancer,Citation9 prostate cancerCitation10 breast cancer,Citation11 thyroid cancer,Citation12 nasopharyngeal carcinoma,Citation13 and other types of malignant diseases. Previously, studies have found that elevated MALAT1 levels are involved in PC proliferation and metastasis through the stimulation of autophagy.Citation14 Nevertheless, the potential molecular mechanism is still unclear and needs additional research. Over the last few years, there have been numerous novel techniques related to cancer mechanism and therapy exploration emerging. For instance, molecular docking and molecular dynamics simulation, which helps identify the cancer-associated single nucleotide polymorphisms and their possible molecular mechanism.Citation15–Citation17 And other analysis including protein–ligand interaction analysis, principal component analysis, shape analysis of binding pocket, and kinase inhibitor screening are also used to explore the therapeutic molecules for cancers.Citation18,Citation19 Researchers are also trying to figure out a methodology to optimize a synergistic and clinically achievable combination of multiple agents for cancer clinical therapy.Citation20 Despite their particular perspectives in uncovering the cancer mechanism and therapy, the complicated and demanding operation of the experimental techniques themselves might limit their further development. As a result, we hope to use computational methods to explore the cancer mechanism by virtue of online databases. Thus, in this paper, we performed a comprehensive study to further explore the clinical significance and underlying molecular mechanism of MALAT1 in PC on a computational level.
To explore the relationship between MALAT1 and PC, we collected gene expression data and clinical data from patients using The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO). The relationship between MALAT1 and PC was further analyzed based on the TCGA and GEO data. For the purpose of exploring the molecular mechanism, we analyzed the overlapping genes in the differentially expressed genes after MALAT1 knockout and the coexpressed genes from TCGA. We performed a bioinformatics functional analysis for the overlapping genes and used Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), and Protein–Protein Interaction (PPI) databases.
Taken together, we hope to understand the clinical value of MALAT1 in PC and the molecular mechanism of MALAT1. We hope that the data shown here may offer new perspectives in future research and clinical application.
Materials and methods
Selection of microarrays in GEO datasets and profiles
For the purpose of understanding the diagnostic value of MALAT1 in PC, we systematically retrieved the GEO microarrays performed before December 2016 for a meta-analysis. The retrieval strategies were as follows: Pancrea* AND (adenocarcinoma OR carcinoma OR cancer OR neoplasm OR tumor OR tumour OR neoplas* OR malignan*). The inclusion criteria were as follows: 1) the enrolled subjects included patients with PC and normal control samples, 2) MALAT1 was expressed, detected, and available for extraction in the enrolled subjects, and 3) there were more than three subjects.
Validation of MALAT1 expression in PC based on the Oncomine database
The Oncomine database (http://www.oncomine.org) incorporates 264 independent datasets that include 35 cancer types and supports various methods of analysis, including molecular concepts analysis, interactome analysis, and meta-analysis.Citation21 Thus, we adopted the Oncomine database to further validate the expression of MALAT1 in PC. The differential expression analysis was directly performed using Oncomine online analysis tools.
Identification of the clinical prognostic value of MALAT1 in PC
TCGA (http://cancergenome.nih.gov) data portal is one of the largest available public resources offering genomic, transcriptomic, methylomic, and copy number variation datasets for more than 20 cancer types.Citation22,Citation23 We evaluated the TCGA survival of patients with PC in two online analysis websites, cBioPortal for Cancer GenomicsCitation24,Citation25 (http://www.cbioportal.org/) and OncoLncCitation26 (http://www.oncolnc.org/), which link to the TCGA database.
Identification of key differentially expressed genes associated with MALAT1 in PC
For the purpose of investigating the genes influenced by MALAT1, we performed GeneChip analysis after MALAT1 was knocked down. The human PC Miapaca-2 cell line was purchased from the Institute of Biochemistry and Cell Biology of the Chinese Academy of Sciences (Wuhan, People’s Republic of China), and was maintained routinely in Dulbecco’s Modified Eagle’s Medium (Thermo Fisher Scientific, Waltham, MA, USA). The four siRNAs for MALAT1 were designed on the internet (http://design.RNAi.jp/) and synthesized by Shanghai Jima company. Lentivirus MALAT1-shRNA was synthesized using lentivirus, packaged, and screened. The knockdown rate of MALAT1-stable cells was 76% in Miapaca-2 cell line with the best siRNA sequence (F:GGCAGCTTTAACAGATAACA; R:CCGTCCGACAAGGGTCATTCA). The three groups of control vector cells and MALAT1-shRNA stable cells were tested with GeneChip prime view human kit (Affymetrix., 901838, Santa Clara, CA, USA). Using MALAT1 knockdowns, we first acquired the differentially expressed genes. After performing GO analysis, we obtained the enriched GO data and the corresponding genes. Furthermore, we collected MALAT1 and mRNA gene coexpression data from the TCGA database. We adopted 0.15 as the cut-off value for screening the significant coexpression genes. Aiming at determining the most likely key differentially expressed MALAT1-associated genes in PC, we further correlated the enriched GO items with correlative genes and TCGA coex-pression genes. The overlapping genes we obtained were used for further analysis.
Bioinformatics analysis of the overlapping genes
For the overlapping genes, The Database for Annotation, Visualization, and Integrated Discovery version 6.7 (https://david-d.ncifcrf.gov/), an online bioinformatics functional enrichment resource for a list of genes analysis,Citation27,Citation28 was used to perform GO enrichment and KEGG pathway analysis. The analysis was achieved via uploading the overlapping genes though the Functional Annotation porch. We chose those items whose P-values were below 0.05 for further study. Further, we used BiNGO, a plugin of Cytoscape 3.40,Citation29 to visualize the GO enrichment pathway, which includes biological process (BP), cellular component (CC), and molecular function (MF). Meanwhile, we also aimed to identify the most crucial target genes of MALAT1 from the overlapping genes. The STRING database version 10 (http://string-db.org/) is aimed at providing a critical assessment and integration of PPI through direct (physical) and indirect (functional) associations network construction.Citation30 Thus, STRING database was further used to perform a protein–protein interaction network analysis. The PPI network construction was achieved through StringApp, a plugin of Cytoscape 3.40, which links to STRING database. Interaction score (0.7) was adopted as a high confidence cut-off value to determine the PPI. Hub genes, which are the key genes in PC, were obtained from the network.
Statistical analysis
Stata 12.0 (Stata Corp., College Station, TX, USA) was used to perform a GEO microarray meta-analysis in this study. Receiver operating characteristic (ROC) curve analyses were utilized to evaluate the diagnostic value of MALAT1 in PC and normal tissue. Kaplan–Meier curves and the log-rank test were applied to analyze the effect of MALAT1 on the survival outcome of patients with PC. Only P<0.05 was considered statistically significant.
Results
Characteristics of the included datasets
A total of 27 eligible datasets published from 2008 to 2016 in the GEO database were used in this study. The characteristics of the selected datasets are shown in . In total, 764 PC samples and 469 healthy controls were used.
Table 1 Characteristics of the GEO datasets included in the meta-analysis
Clinical diagnostic value of MALAT1 as a biomarker in PC based on the GEO datasets
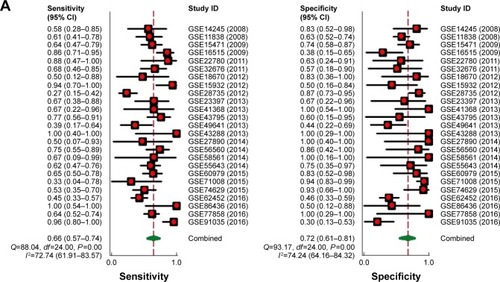
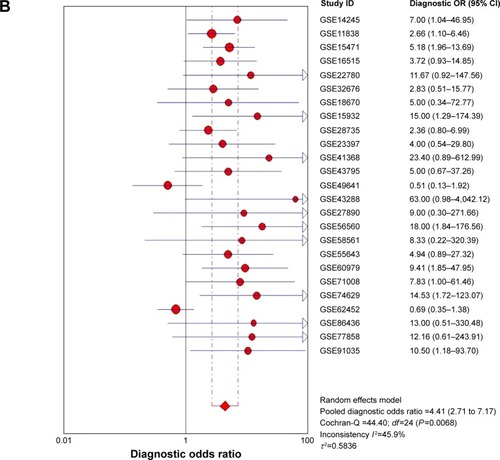
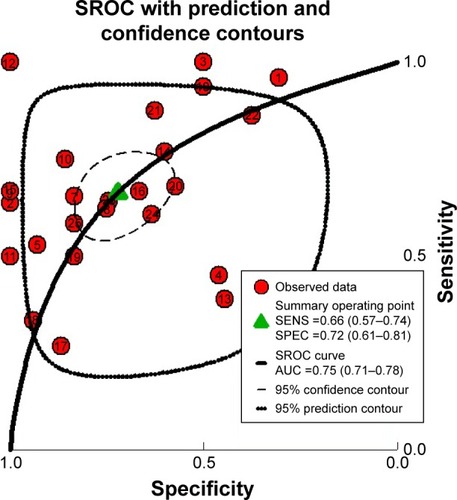
After pooling the data from the 27 eligible datasets, we found that there was significant heterogeneity among the datasets (I2=61.1%, P=0.000, ). As a result, we performed a sensitivity analysis to screen the possible data-sets that might cause the heterogeneity. As can be seen in , two datasets, namely, GSE15471 and GSE62165, may contribute to significant heterogeneity. Thereby, we removed the two datasets and recalculated the pooled standard mean difference. As shown in , the heterogeneity appears to have disappeared (I2=44.9%, P=0.009). The pooled standard mean difference was 0.23 (95% confidence interval [CI]: 0.10–0.37, P=0.001), which indicates a significant difference in the effect of MALAT1 between patients with PC and healthy control patients. The expression of MALAT1 was significantly upregulated. Publication bias among the 25 datasets was assessed through Begg’s funnel plot. As shown in , the funnel plot was basically considered symmetrical. Taking P>0.05 as cut-off value, we found that P=0.414 was obtained by Begg’s test, which indicated that no significant publication bias existed. Furthermore, a summary ROC curve revealed that the area under the curve was 0.75 (95% CI: 0.71–0.78), and the overall sensitivity and specificity was 0.66 (95% CI: 0.57–0.74) and 0.72 (95% CI: 0.61–0.81) ( and ). The pooled diagnostic odds ratio was also calculated as 4.41 (95% CI: 2.71–7.17), which indicates that upregulated MALAT1 results in a higher risk of PC ().
Figure 1 Forest plot of 27 GEO datasets evaluating the SMD of MALAT1 in the PC group and the healthy control group (a fixed-effect model).
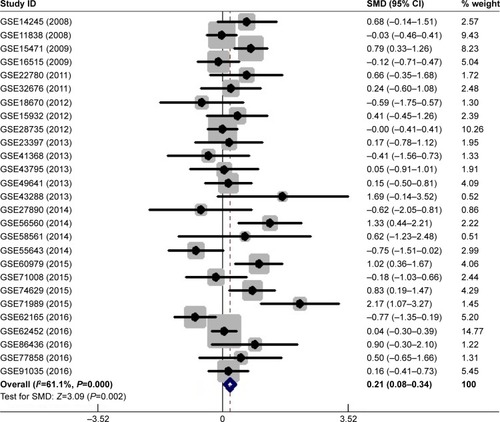
Figure 2 Sensitivity analysis to evaluate the heterogeneity among 27 GEO datasets.
Abbreviations: CI, confidence interval; GEO, Gene Expression Omnibus.
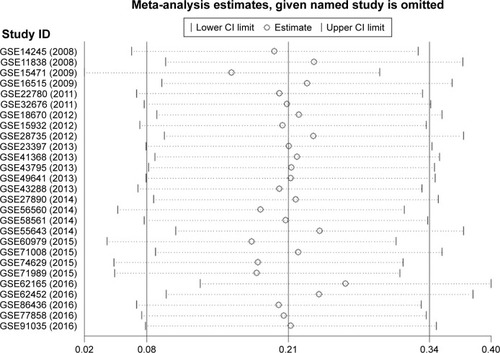
Figure 3 Forest plot of 25 GEO datasets evaluating the SMD of MALAT1 in the PC group and the healthy control group after omitting two datasets (GSE15471 and GSE62165) (a fixed-effect model).
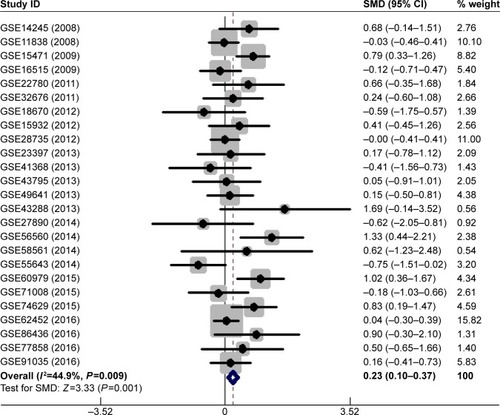
Figure 4 Begg’s funnel plot evaluating the publication bias among the 25 datasets.
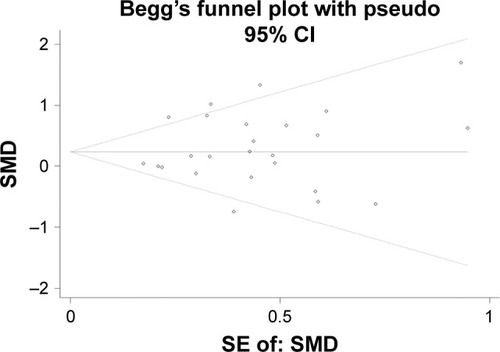
Figure 5 Summary of the ROC curve of 25 datasets evaluating the diagnostic value of MALAT1 in PC.
MALAT1 expression difference validation based on the Oncomine database
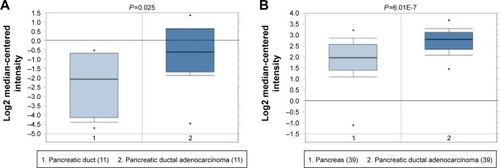
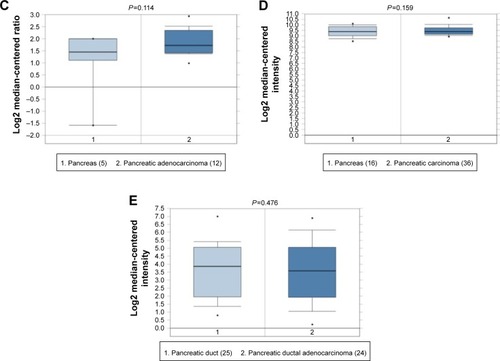
Five PC datasets in the Oncomine database were adopted for the validation of MALAT1 expression (Grutzmann’s dataset [http://www.ebi.ac.uk/arrayexpress/experiments/E-MEXP-950]; Badea’s datasets [http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE15471]; lacobuzio-Donahue’s dataset [http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE3654; http://genome-www.stanford.edu/pancreatic1/index.shtml]; Pie’s dataset [http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE16515]; Ishikawa’s dataset [http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE1542]). In Grutzmann’s and Badea’s datasets, the expression of MALAT1 was significantly elevated in PC, which was in accordance with the results of the meta-analysis (). The P-values for these datasets were 0.025 and <0.01, respectively. However, the remaining three datasets did not show a statistically significant difference in MALAT1 expression ().
Figure 7 Box plot validating the expression of MALAT1 in the PC group and the healthy control group based on the Oncomine database.
Abbreviation: PC, pancreatic cancer.
Prognostic value of MALAT1 based on the TCGA database
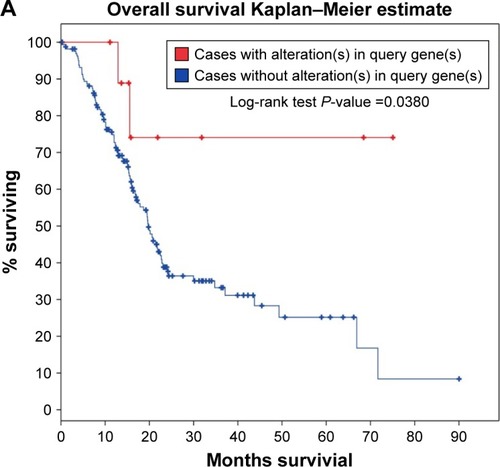
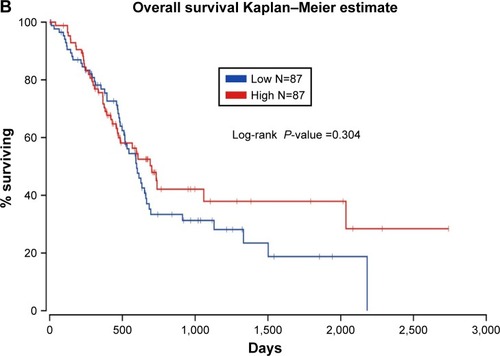
For the analysis using the cBioPortal for Cancer Genomics, the survival data for a total of 178 PC patients was analyzed. We found 10 of 178 patients had alterations in the expression of MALAT1 (Supplementary material 1). Thus, we evaluated the survival of the cases with MALAT1 alteration compared to those without MALAT1 alteration. The log-rank test showed that the P-value was 0.0380 (). As reported on the Oncolnc website, we analyzed the survival of patients with high MALAT1 expression and compared it to that of patients with low MALAT1 expression in a total of 174 patients with PC. It showed a trend that high MALAT1 expression indicates better survival, although no statistical significance was observed (P=0.304, ).
Collection of the overlapping genes
After knocking down MALAT1, we performed a GO enrichment analysis and collected 656 enriched GO gene symbols. Furthermore, using 0.15 as P-value cut-off, we obtained 5,887 coexpressed genes from the TCGA database. We then intersected the two part genes and gathered 224 overlapping genes (Supplementary material 2), which might become the crucial targets genes of MALAT1 in PC.
Bioinformatics analysis of the overlapping genes
As displayed in , the overlapping genes contain three enriched GO pathway categories: BP, CC, and MF. In the BP category, the enriched items were mainly intracellular transport, regulation of phosphorus metabolic process, regulation of phosphate process, etc. In the CC category, membrane-enclosed lumen, nuclear lumen, and nuclear chromosome, among others, were the main components of the enrichment. In regards to the MF category, protein dimerization activity, nucleoside binding, and purine nucleoside binding ranked as the top three enriched items. All of the enriched GO pathways, including the BP, CC, and MF categories, were further visualized through Cytoscape 3.40 and the results are displayed in Supplementary materials 3 and 4 and , respectively. In the KEGG pathway analysis, we identified the mTOR signaling pathway, pathways in cancer, and the MAPK signaling pathway as the top three enriched pathways (). Interestingly, we also found that the PC pathway was one of the significantly enriched pathways. Five target genes (VEGFC, CCND1, PIK3CB, VEGFA, and MAPK8) were involved in the PC pathway. A PPI was constructed (Supplementary material 5) and six hub genes (CCND1, MAPK8, VEGFA, FOS, CDH1, and HSP90AA1) were identified ().
Figure 9 Significant MF network constructed by Cytoscape 3.40.
Abbreviations: GO, Gene Ontology; MF, molecular function.
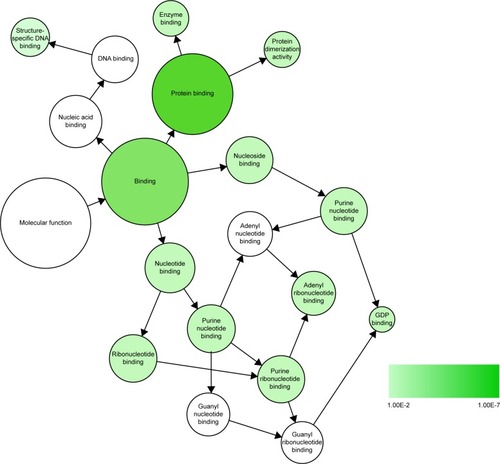
Figure 10 Hub genes obtained from the PPI network.
Abbreviation: PPI, protein–protein interaction.
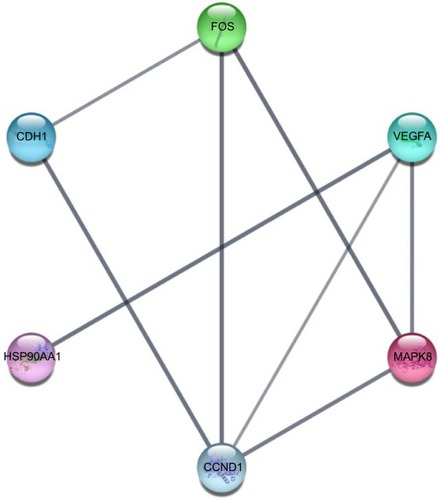
Table 2 Significant GO and KEGG items of the overlapping genes
Discussion
In this paper, we first discovered that the expression of MALAT1 was significantly upregulated in PC patients by using a meta-analysis of 27 published GEO microarrays, which included a total of 764 patients with PC and 469 healthy controls. Further validation via the Oncomine database was consistent with the GEO microarray meta-analysis. Using the TCGA database, we acquired the survival data of the patients and investigated the influence of the alteration of MALAT1 expression on the patients’ prognosis. Furthermore, using our own GeneChip and TCGA database, we collected and intersected the enriched GO genes after knocking down MALAT1 and the coexpressed genes in TCGA. Further bioinformatics analysis of the overlapping genes, which aims to explore the underlying molecular regulatory mechanism, was performed.
During the past few years, alteration of MALAT1 has been found to be involved in malignancies and other diseases. Shuai et alCitation31 performed a meta-analysis that revealed that MALAT1 served as a promising biomarker for the prognosis of numerous malignancies, including digestive system cancers, urinary system cancers, and respiratory system cancers. In our study, we observed upregulated MALAT1, which is consistent with other previous reports, such as those by Li et al,Citation14 Pang et al,Citation32 and Liu et al.Citation33 Elevated MALAT1 has also been reported in gastric cancer,Citation34 hepatocellular carcinoma,Citation35 gallbladder cancer,Citation36 esophageal cancer,Citation37 and clear cell renal cell carcinoma.Citation38 Considering all of the above information, MALAT1 appears to play a distinct role in different tumors, which provides a brand new perspective for future cancer therapy.
In the present study, we discovered that MALAT1 showed a moderate performance in diagnosing PC after meta-analysis of a large number of PC, and corresponding control, samples. The area under the curve for the summary ROC curve was 0.75, with a sensitivity of 0.66 and a specificity of 0.72. Thus far in the published literature, there is no report evaluating the diagnostic value of MALAT1 in PC through comprehensive meta-analysis. As a consequence, the information in this study will provide a valuable reference for clinical PC diagnosis. To date, several studies have reported that higher MALAT1 expression suggests poorer survival.Citation31,Citation32 Regarding our results, we found that patients with MALAT1 alteration had better survival than patients without MALAT1 alteration (P=0.0380). We also found that the high expression of MALAT1 suggested better survival in the Kaplan–Meier curves. However, the P-values did not show statistical significance. Therefore, the exact prognostic value of MALAT1 in PC still remains unknown.
Since lncRNAs exert their regulatory function by specifically binding with target genes, we also aimed to determine possible targets of MALAT1 and to further explore the possible molecular pathways. Through a GO enrichment analysis, some top-ranked GO items, such as intracellular transport, membrane-enclosed lumen, and protein dimerization activity, were found to be possible crucial events in PC development. Regarding the KEGG pathway analysis, several significant pathways, such as the mTOR signaling pathway, pathways in cancer, and the MAPK signaling pathway, were discovered. Importantly, the PC pathway was also one of the enriched KEGG pathways. Five genes, VEGFC, CCND1, PIK3CB, VEGFA, and MAPK8, are target genes of MALAT1, which might also be key target genes in PC. Interestingly, according to previous studies, we found that the mTOR signaling pathway has been reported to be involved in PC.Citation39,Citation40 Additionally, the MAPK signaling pathway also has been reported to play a role in PC.Citation41 Therefore, to some extent, the pathways identified here might provide more evidence for elucidating the potential molecular mechanism in PC. Finally, the PPI network suggests that six hub genes (CCND1, MAPK8, VEGFA, FOS, CDH1, and HSP90AA1) might serve as pivotal target genes of MALAT1 in PC. We discovered that CCND1, MAPK8, and VEGFA also participate in the PC pathway, as mentioned. As a consequence, we concentrate on the top three genes (CCND1, MAPK8, and VEGFA), which may be important genes, for detailed discussion.
CCND1, also known as BCL1, encodes a highly conserved cyclin protein, which exerts regulatory functions on CDK kinases. Mutation, amplification, and overexpression of CCND1 alter cell cycle progression, which might contribute to tumorigenesis. As a result, CCND1 alteration was frequently discovered in various tumors. The CCND1 genotype has been reported to be related to the risk for various cancers, such as colorectal cancer,Citation42 acute lymphoblastic leukemia,Citation43 glioma,Citation44 esophageal squamous cell carcinoma,Citation45 gastric cancer,Citation46 and nasopharyngeal carcinoma.Citation47,Citation48 Moreover, CCND1 was also reported to play different roles in cancer survival, such as in lung adenocarcinoma,Citation49,Citation50 prostate cancer,Citation51 meningiomaCitation52 gastric adenocarcinoma,Citation53 and colorectal cancer.Citation54 In regards to PC, Deharvengt et alCitation55 reported that CCND1 is a target of a B lentivirus-delivered shRNA, which exerts suppressive effects on the growth, invasiveness, tumorigenicity, and proangiogenic potential of PC. Another study found that CCND1 was a strong prognostic biomarker for PC survival.Citation56 Taken together, it is obvious that CCND1 is involved in various cancers, including PC. However, whether it is related to MALAT1 has not been reported to date, and further studies are required.
MAPK8, also named JNK or JNK1, belongs to the MAPK kinase family that acts as an integration point in numerous cellular processes, such as proliferation, differentiation, transcription regulation, and development. Increasing amounts of evidence have proven that MAPK8 has a crucial role in a wide variety of cancers. Chang et alCitation57 discovered that JNK1 activation was capable of predicting poor prognosis in hepatocellular carcinoma. Moreover, Okada et alCitation58 reported that JNK can inhibit temozolomide, thus acting as a rational therapeutic biomarker in glioblastoma. JNK activity has also been reported in endometrial cancer,Citation59 melanoma,Citation60 colon cancer,Citation61 etc. In regard to PC, Sahu et alCitation62 reported that JNK was involved in benzyl isothiocyanate (BITC)-mediated G(2)/M arrest to mediate apoptosis in human PC cells. Another study also found that JNK1 is activated by Irofulven treatment, which induces apoptosis in human PC cells.Citation63 Therefore, based on the abovementioned information, MAPK8 could possibly serve as a target gene of MALAT1 in PC, but additional studies are needed.
VEGFA, also known as VPF, VEGF, and MVCD1, is a member of the PDGF/VEGF growth factor family. Encoding a heparin-binding protein, it induces the proliferation and migration of vascular endothelial cells, which is fundamental for physiological and pathological angiogenesis. Upregulated VEGFA is common in many tumors and is related to tumor stage and progression. Liu et alCitation64 reported that VEGFA polymorphisms might play a potential role in the development and clinical outcome of hepatocellular carcinoma. VEGFA has also been found to be inhibited by miR-1, which exerts inhibitory functions in osteosarcoma cells, as well as promotes proliferation, migration, and invasion.Citation65 Furthermore, Zeng et alCitation66 found that downregulation of VEGFA can inhibit proliferation, migration, and invasion and can promote apoptosis in renal clear cell carcinoma. VEGFA in tumors has also been studied in breast cancer,Citation67,Citation68 laryngeal carcinoma,Citation69 colorectal cancer,Citation70,Citation71 bladder cancer,Citation72 and lung cancer.Citation73 In PC, Liu et alCitation74 found that VEGFA was involved in the Twist/miR-497/VEGFA axis, which is significantly correlated to metastasis and angiogenesis in PC. VEGFA secretion has also been reported to be regulated by myoferlin and affects tumor-associated angiogenesis in PC.Citation75 In summary, as VEGFA has been shown to be important in cancers, it is suggested that VEGFA is a target of MALAT1 in PC. However, more studies are needed.
There are limitations in this study. Here, we have attempted to explore the relationship between MALAT1 and PC clinical parameters based on the TCGA database. However, no statistical significance was observed. Therefore, it still remains to be seen if there is a correlation between MALAT1 and PC. In addition, in this study, the exploration occurred at a bioinformatics level. Further experiments are needed to validate these assumptions.
In sum, we discovered that upregulated MALAT1 acts as a biomarker for the diagnosis of PC. The alteration or upregulation of MALAT1 expression tends to predict better prognostic outcome in PC. Several pathways, such as the mTOR signaling pathway, pathways in cancer, and the MAPK signaling pathway, might be targeted by MALAT1 and participate in the PC process. Six hub genes (CCND1, MAPK8, VEGFA, FOS, CDH1, and HSP90AA1), and specifically CCND1, MAPK8 and VEGFA, were identified as key target genes of MALAT1 in PC. Here, we provided clinical reference values and underlying molecular mechanisms for MALAT1 in PC. These findings still need to be validated in future studies.
Conclusion
MALAT1 seems to act as a promising diagnostic biomarker in PC. Six hub genes (CCND1, MAPK8, VEGFA, FOS, CDH1, and HSP90AA1), specifically CCND1, MAPK8, and VEGFA, might be key correlative genes of MALAT1 in PC. Several pathways related to MALAT1, such as the mTOR signaling pathway, pathways in cancer, and the MAPK signaling pathway may play a crucial role in PC. The information we have obtained might shed light on clinical application and future research. Nevertheless, more experimental studies are needed to further validate these findings.
Acknowledgments
The study was supported by the Fund of National Natural Science Foundation of China (NSFC 81560448), the Natural Science Foundation of Guangxi, China (2014GXNSF BA118167), and Guangxi Medical University Students Innovative Project (201610598100). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Disclosure
The authors report no conflicts of interest in this work.
References
- LeiQZhengHBiJWhole grain intake reduces pancreatic cancer risk: a meta-analysis of observational studiesMedicine (Baltimore)2016959e274726945361
- AmbeCMMahipalAFulpJChenLMalafaMPEffect of metformin use on survival in resectable pancreatic cancer: a single-institution experience and review of the literaturePLoS One2016113e015163226967162
- BeenJVMackenbachJPMillettCBasuSSheikhATobacco control policies and perinatal and child health: a systematic review and meta-analysis protocolBMJ Open201559e008398
- ShiXSunMWuYPost-transcriptional regulation of long noncoding RNAs in cancerTumour Biol201536250351325618601
- YuanJYueHZhangMTranscriptional profiling analysis and functional prediction of long noncoding RNAs in cancerOncotarget2016778131814226812883
- LizJEstellerMlncRNAs and microRNAs with a role in cancer developmentBiochim Biophys Acta20161859116917626149773
- WeiYNiuBRole of MALAT1 as a prognostic factor for survival in various cancers: a systematic review of the literature with meta-analysisDis Markers2015201516463526420912
- LeiRXueMZhangLLinZLong noncoding RNA MALAT1-regulated microRNA 506 modulates ovarian cancer growth by targeting iASPPOnco Targets Ther201710354628031721
- LiZZhouYTuBBuYLiuAXieCLong noncoding RNA MALAT1 affects the efficacy of radiotherapy for esophageal squamous cell carcinoma by regulating Cks1 expressionJ Oral Pathol Med Epub20161209
- AielloABacciLReAMALAT1 and HOTAIR long non-coding rnas play opposite role in estrogen-mediated transcriptional regulation in prostate cancer cellsSci Rep201663841427922078
- JadalihaMZongXMalakarPFunctional and prognostic significance of long non-coding RNA MALAT1 as a metastasis driver in ER negative lymph node negative breast cancerOncotarget2016726404184043627250026
- HuangJKMaLSongWHMALAT1 promotes the proliferation and invasion of thyroid cancer cells via regulating the expression of IQGAP1Biomed Pharmacother2016831727470543
- HuaWFZhongQXiaTLRBM24 suppresses cancer progression by upregulating miR-25 to target MALAT1 in nasopharyngeal carcinomaCell Death Dis201679e235227584791
- LiLChenHGaoYLong noncoding RNA MALAT1 promotes aggressive pancreatic cancer proliferation and metastasis via the stimulation of autophagyMol Cancer Ther20161592232224327371730
- KamarajBRajendranVSethumadhavanRPurohitRIn-silico screening of cancer associated mutation on PLK1 protein and its structural consequencesJ Mol Model201319125587559924271645
- KumarAPurohitRUse of long term molecular dynamics simulation in predicting cancer associated SNPsPLoS Comput Biol2014104e100331824722014
- RajendranVStructural analysis of oncogenic mutation of isocitrate dehydrogenase 1Mol biosyst20161272276228727194485
- RajendranVGopalakrishnanCPurohitRImpact of point mutation P29S in RAC1 on tumorigenesisTumour Biol20163711152931530427699663
- BaranskiZBooijTHKuijjerMLMEK inhibition induces apoptosis in osteosarcoma cells with constitutive ERK1/2 phosphorylationGenes Cancer2015611–1250351226807203
- YuDKahenECubittCLIdentification of synergistic, clinically achievable, combination therapies for osteosarcomaSci Rep201551699126601688
- RhodesDRKalyana-SundaramSMahavisnoVOncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profilesNeoplasia20079216618017356713
- DengMBragelmannJSchultzeJLPernerSWeb-TCGA: an online platform for integrated analysis of molecular cancer data setsBMC Bioinformatics2016177226852330
- KrasnovGSDmitrievAAMelnikovaNVCrossHub: a tool for multi-way analysis of The Cancer Genome Atlas (TCGA) in the context of gene expression regulation mechanismsNucleic Acids Res2016447e6226773058
- GaoJAksoyBADogrusozUIntegrative analysis of complex cancer genomics and clinical profiles using the cBioPortalSci Signal20136269pl123550210
- CeramiEGaoJDogrusozUThe cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics dataCancer Discov20122540140422588877
- AnayaJOncoLnc: linking TCGA survival data to mRNAs, miRNAs, and lncRNAsPeer J Computer Science20162e67
- Huang daWShermanBTLempickiRASystematic and integrative analysis of large gene lists using DAVID bioinformatics resourcesNat Protoc200941445719131956
- Huang daWShermanBTLempickiRABioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene listsNucleic Acids Res200937111319033363
- ShannonPMarkielAOzierOCytoscape: a software environment for integrated models of biomolecular interaction networksGenome Res200313112498250414597658
- SzklarczykDFranceschiniAWyderSSTRING v10: protein-protein interaction networks, integrated over the tree of lifeNucleic Acids Res201543Database issueD447D45225352553
- ShuaiPZhouYGongBLong noncoding RNA MALAT1 can serve as a valuable biomarker for prognosis and lymph node metastasis in various cancers: a meta-analysisSpringerPlus201651172127777857
- PangEJYangRFuXBLiuYFOverexpression of long non-coding RNA MALAT1 is correlated with clinical progression and unfavorable prognosis in pancreatic cancerTumour Biol20153642403240725481511
- LiuJHChenGDangYWLiCJLuoDZExpression and prognostic significance of lncRNA MALAT1 in pancreatic cancer tissuesAsian Pac J Cancer Prev20141572971297724815433
- LeeNKLeeJHIvanCMALAT1 promoted invasiveness of gastric adenocarcinomaBMC Cancer20171714628077118
- MalakarPShiloAMogilavskyALong noncoding RNA MALAT1 promotes hepatocellular carcinoma development by SRSF1 up-regulation and mTOR activationCancer Res20177751155116727993818
- WangSHZhangWJWuXCLong non-coding RNA Malat1 promotes gallbladder cancer development by acting as a molecular sponge to regulate miR-206Oncotarget2016725378573786727191262
- HuangCYuZYangHLinYIncreased MALAT1 expression predicts poor prognosis in esophageal cancer patientsBiomed Pharmacother20168381327470544
- ZhangHMYangFQChenSJCheJZhengJHUpregulation of long non-coding RNA MALAT1 correlates with tumor progression and poor prognosis in clear cell renal cell carcinomaTumour Biol20153642947295525480417
- HeLWuYLinLHispidulin, a small flavonoid molecule, suppresses the angiogenesis and growth of human pancreatic cancer by targeting vascular endothelial growth factor receptor 2-mediated PI3K/Akt/mTOR signaling pathwayCancer Sci2011102121922521087351
- KagawaSTakanoSYoshitomiHAkt/mTOR signaling pathway is crucial for gemcitabine resistance induced by Annexin II in pancreatic cancer cellsJ Surg Res2012178275876722726648
- WuYYMaTLGeZJJWA gene regulates PANC-1 pancreatic cancer cell behaviors through MEK-ERK1/2 of the MAPK signaling pathwayOncol Lett2014841859186325202426
- HuangCYTsaiCWHsuCMChangWSShuiHABauDTThe significant association of CCND1 genotypes with colorectal cancer in TaiwanTumour Biol20153686533654025809706
- XueYRongLTongNWangMZhangZFangYCCND1 G870A polymorphism is associated with toxicity of methotrexate in childhood acute lymphoblastic leukemiaInt J Clin Exp Pathol201589115941160026617896
- ZongHCaoLMaCAssociation between the G870A polymorphism of Cyclin D1 gene and glioma riskTumour Biol20143588095810124840634
- HussainSMYThakurNAssociation of cyclin D1 gene polymorphisms with risk of esophageal squamous cell carcinoma in Kashmir Valley: a high risk areaMol Carcinog201150748749821268129
- KuoHWHuangCYFuCKThe significant association of CCND1 genotypes with gastric cancer in TaiwanAnticancer Res20143494963496825202078
- ShihLCTsaiCWTsaiMHAssociation of cyclin D1 genotypes with nasopharyngeal carcinoma riskAnticancer Res20123231093109822399638
- LiaoDWuYPuXCyclin D1 G870A polymorphism and risk of nasopharyngeal carcinoma: a case-control study and meta-analysisPLoS One2014911e11329925409185
- XuPZhaoMLiuZElevated nuclear CCND1 expression confers an unfavorable prognosis for early stage lung adenocarcinoma patientsInt J Clin Exp Pathol2015812158871589426884860
- LeeEJinDLeeBBNegative effect of cyclin D1 overexpression on recurrence-free survival in stage II-IIIA lung adenocarcinoma and its expression modulation by vorinostat in vitroBMC Cancer20151598226681199
- TsaiHMoraisCLAlshalalfaMCyclin D1 loss distinguishes prostatic small-cell carcinoma from most prostatic adenocarcinomasClin Cancer Res201521245619562926246306
- ChengGZhangLLvWDongCWangYZhangJOverexpression of cyclin D1 in meningioma is associated with malignancy grade and causes abnormalities in apoptosis, invasion and cell cycle progressionMedical Oncol2015321439
- MaLWangXLanFPrognostic value of differential CCND1 expression in patients with resected gastric adenocarcinomaMedical Oncol2015321338
- LiYWeiJXuCZhaoZYouTPrognostic significance of cyclin D1 expression in colorectal cancer: a meta-analysis of observational studiesPLoS One201494e9450824728073
- DeharvengtSJGunnJRPickettSBKorcMIntratumoral delivery of shRNA targeting cyclin D1 attenuates pancreatic cancer growthCancer Gene Ther201017532533319851352
- BachmannKNeumannAHinschACyclin D1 is a strong prognostic factor for survival in pancreatic cancer: analysis of CD G870A polymorphism, FISH and immunohistochemistryJ Surg Oncol2015111331632325470788
- ChangQChenJBeezholdKJCastranovaVShiXChenFJNK1 activation predicts the prognostic outcome of the human hepatocellular carcinomaMol Cancer200986419686584
- OkadaMSatoAShibuyaKJNK contributes to temozolomide resistance of stem-like glioblastoma cells via regulation of MGMT expressionInt J Oncol201444259159924316756
- RenoEMHaughianJMJacksonTAThorneAMBradfordAPc-Jun N-terminal kinase regulates apoptosis in endometrial cancer cellsApoptosis200914680982019424800
- AlexakiVIJavelaudDMauvielAJNK supports survival in melanoma cells by controlling cell cycle arrest and apoptosisPigment Cell Melanoma Res200821442943818541008
- MahalingamDKeaneMPirianovGMehmetHSamaliASzegezdiEDifferential activation of JNK1 isoforms by TRAIL receptors modulate apoptosis of colon cancer cell linesBr J Cancer200910091415142419352384
- SahuRPZhangRBatraSShiYSrivastavaSKBenzyl isothiocyanate-mediated generation of reactive oxygen species causes cell cycle arrest and induces apoptosis via activation of MAPK in human pancreatic cancer cellsCarcinogenesis200930101744175319549704
- WangWWatersSJMacDonaldJRIrofulven (6-hydroxym ethylacylfulvene, MGI 114)-induced apoptosis in human pancreatic cancer cells is mediated by ERK and JNK kinasesAnticancer Res2002222A55956412014623
- LiuFLuoLWeiYAssociation of VEGFA polymorphisms with susceptibility and clinical outcome of hepatocellular carcinoma in a Chinese han populationOncotarget2017810164881649728147320
- NiuJSunYGuoQNiuDLiuBmiR-1 Inhibits cell growth, migration, and invasion by targeting VEGFA in osteosarcoma cellsDis Markers20162016706898627777493
- ZengFCZengMQHuangLDownregulation of VEGFA inhibits proliferation, promotes apoptosis, and suppresses migration and invasion of renal clear cell carcinomaOnco Targets Ther201692131214127110129
- Vieira-Monteiro HdeAFreitas-AlvesDRSobral-LeiteMPrognostic evaluation of VEGFA genotypes and haplotypes in a cohort of Brazilian women with non metastatic breast cancerCancer Biol Ther201617667468327195611
- WuJYuanPMaoQmiR-613 inhibits proliferation and invasion of breast cancer cell via VEGFABiochem Biophys Res Commun2016478127427827449609
- XuLShenBChenTDongPmiR-203 is involved in the laryngeal carcinoma pathogenesis via targeting VEGFA and Cox-2Onco Targets Ther201694629463727555783
- ZhangWZouCPanLMicroRNA-140-5p inhibits the progression of colorectal cancer by targeting VEGFACell Physiol Biochem20153731123113326402430
- TakahashiNIwasaSTaniguchiHPrognostic role of ERBB2, MET and VEGFA expression in metastatic colorectal cancer patients treated with anti-EGFR antibodiesBr J Cancer201611491003101127002940
- GaoYWuKChenYBeyond proliferation: KLF5 promotes angiogenesis of bladder cancer through directly regulating VEGFA transcriptionOncotarget2015641437914380526544730
- ZhangSDMcCruddenCMKwokHFPrognostic significance of combining VEGFA, FLT1 and KDR mRNA expression in lung cancerOncol Lett20151031893190126622771
- LiuAHuangCCaiXXuJYangDTwist promotes angiogenesis in pancreatic cancer by targeting miR-497/VEGFA axisOncotarget2016718258012581427015364
- FahmyKGonzalezAArafaMMyoferlin plays a key role in VEGFA secretion and impacts tumor-associated angiogenesis in human pancreas cancerInt J Cancer2016138365266326311411