Abstract
Epidermal growth factor, latrophilin, and seven transmembrane domain-containing protein 1 on chromosome 1 (ELTD1), an orphan adhesion G-protein coupled receptor, was reported as a regulator of angiogenesis, also involved in cancer progression and development. More recently, ELTD1 was identified as a potential new tumor marker for high-grade glioma. ELTD1, belongs to the G-protein coupled receptor superfamily that comprises the biggest receptor family in the human genome. Following the discovery of ELTD1 almost a decade ago, only a few research groups have attempted to find its role in normal and tumor cells, important information about this receptor remaining still unknown. The ELTD1 ligand has not currently been identified and intracellular signaling studies have not yet been performed in normal or tumor cells. Although the current published data on ELTD1 function and structure are rather limited, this receptor seems to be very important, not only as biomarker, but also as molecular target in glioblastoma. This review summarizes and discusses the current knowledge on ELTD1 structure, function, and its role in both physiological and tumoral angiogenesis.
Keywords:
Introduction
Since the early ages of oncology, medical researchers have strived to discover new biomarkers, which can be correlated with early stages of cancer development, tumor progression, and resistance to therapy. With the discovery of targeted therapy, some of these markers became interesting therapeutic targets due to their implication in intricate pathophysiological processes involved in cancer growth, invasion, or resistance to curative approaches. One of the most studied pathophysiological events involved in cancer development and progression is angiogenesis. Depriving nascent cancer cells of their nutrients provided by aberrant, newly formed blood vessels became an attractive approach that, in theory, could deter or even reverse neoplastic growth. However, both early and more recent trials have yielded limited results due to the complex nature of microvasculature in each cancer type, the multiple pathways implicated in cell signaling, and the vast number of genes responsible for both normal and cancer angiogenesis.Citation1–Citation5 Together, these variables can confer both natural and acquired resistance to anti-angiogenic therapy, leading to mixed clinical results.Citation6–Citation8 Many angiogenic inhibitors, designated for cancer treatment, are currently in preclinical and clinical use. The vascular endothelial growth factor receptor family is a well-studied angiogenic class of receptorsCitation9–Citation11 that have been constantly used to develop new cancer therapies for quite a while now.Citation12–Citation14 The therapeutic results, however, for many malignant diseases, in which vascular endothelial growth factor receptor dysfunction is believed to drive tumor angiogenesis, have been ineffective. Future approaches should include inhibition of multiple pathways involved in cancer microvasculature formation or the discovery of new angiogenesis markers, more specific for each cancer type.Citation15–Citation18
One such possible candidate is epidermal growth factor, latrophilin, and seven transmembrane domain-containing protein 1 on chromosome 1 (ELTD1), a biomarker recently linked to both normal and pathological angiogenesis.Citation19,Citation20 An editorial article linked ELTD1 levels to how the human organism responds to hematopoietic stem cell transplant by developing graft-versus-host disease.Citation21 Using complementary DNA microarray, another study also found that ELTD1 was significantly upregulated in patients with ulcerative colitis with dysplastic lesions when compared with normal, nondysplastic, and colic mucosa.Citation22 Upregulation of the ELTD1 gene was observed in studies describing genomic adaption to high altitudes in the Andean population,Citation23 cannabis use disorders,Citation24 genetic predisposition for obesity in both humans and pigs,Citation25 and even resistance to therapy in cattle afflicted by ticks.Citation26
Initially mentioned in 2001, as a receptor normally regulated in the human heart,Citation27 ELTD1 has been recently linked to the progression and development of glioblastoma multiforme (GBM),Citation28,Citation29 a particularly aggressive brain tumor with a very bleak prognosis.
Although it was discovered 14 years ago,Citation27 in the scientific literature, only a few published works have dealt directly with this topic. The current review is an analysis of the information currently available on ELTD1.
Origin and structure of the ELTD1 receptor
Ontogenetically, ELTD1 belongs to the G-protein coupled receptor (GPCR) superfamily, which comprises the biggest receptor family in the human genome. Also called the 7α-helicases transmembrane receptors, the GPCR superfamily transducts signals from the extracellular to the intracellular domain using guanine-binding proteins. The GPCR superfamily is involved in a multitude of human physiological processes, pertaining to different areas such as chemotaxis, smell, hormone secretion, taste, vision, or inflammation.Citation30–Citation32
Containing over 900 members, the GPCR superfamily comprises five major families: glutamate (22 members), rhodopsin (701 members), adhesion (33 members), frizzled (11 members), and secretin (15 members).Citation33
The second biggest of the five families forming the GPCR super group, the adhesion family was initially considered as part of the secretin family, but was later revealed to be a distinct group. As a particular feature, this family presents an unusually elongated N-terminal ectodomain, which is often perceived as combinations of different adhesion-linked motifs: cadherins,Citation34 thrombospondins,Citation35,Citation36 leucine-rich repeats,Citation37 and epithelial growth factor (EGF)-like domain.Citation38,Citation39 Members of the adhesion family are implicated in multiple cellular functions such as cytoskeletal regulation,Citation40–Citation43 planar cell polarity,Citation42,Citation44,Citation45 cell adhesion and migration,Citation46–Citation48 cellular proliferation and death,Citation35,Citation49,Citation50 hematopoiesis, immunity,Citation51–Citation53 and angiogenesis.Citation19,Citation54–Citation57 The adhesion family includes the latrophilin-like subfamily, with its four members: the orphan ELTD1 receptor, as well as the latrophilin 1, 2, and 3 receptors.Citation58
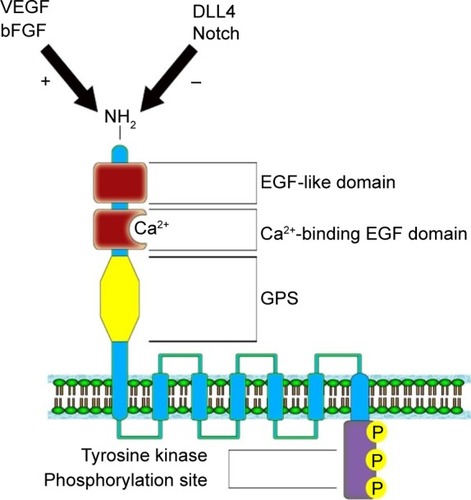
Nechiporuk et alCitation27 mapped the ELTD1 gene on the 1p33–p32 band, belonging to chromosome 1. The gene contains 15 exons having two splice variants: a full-length version and a shorter variant represented by a fraction of a transmembrane domain.Citation59 The gene encodes a 3,527 nucleotide transcript, which further translates into a 690 amino acid protein. The ELTD1 receptor is formed of a large extracellular domain, a seven transmembrane domain coupled with a short cytoplasmic tail. The extracellular domain contains an N-terminal region, an EGF-like domain, a calcium-binding EGF domain, and a GPCR proteolysis site. The cytoplasmic domain contains a tyrosine kinase phosphorylation site, suggested to be involved in receptor intracellular signaling ().Citation27 Unfortunately, no information is yet available in current literature on the downstream signaling pathways implicated in signal transduction or the ligands that bind to the ELTD1 receptor.
Figure 1 Structure of the ELTD1 receptor. The VEGF/bFGF signaling pathway has a positive effect on ELTD1 expression, while DLL4/Notch pathway determines a reduction in protein expression.Citation60–Citation62
ELTD1 implication in angiogenesis
Due to the low number of studies concerning ELTD1, its roles in both physiological and pathological angiogenesis are quite unknown. In the first study published in this area, Nechiporuk et alCitation27 asserted that the ELTD1 receptor was highly expressed in cardiomyocytes and smooth muscle cells of both blood vessels and bronchi, in murine models. ELTD1 regulation was also linked to the switch between hyperplastic and hypertrophic evolution in the development of cardiomyocytes during the physiological differentiation process, in both human and rat models. The receptor’s presence in the coronary arteries also raises further question marks regarding its implication in coronary angiogenesis.Citation27 More recently, another study mentioned that ELTD1 was actively implicated in cardiac hypertrophy, in rat models.Citation63 This study also pointed out that ELTD1 impairment was linked to the faulty cardiomyocytic remodeling process, produced by cardiac pressure overload. ELTD1 gene knockout resulted in an abrupt rise in myocardial fibrosis and hypertrophy of cardiac myocytes.
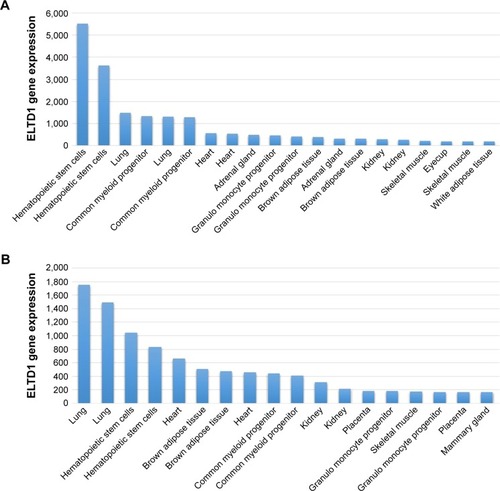
ELTD1 was also reported to be involved in brain angiogenesis. A study by Wallgard et alCitation1 of 58 genes involved in the development of microvasculature in different tissues indicated that ELTD1 expression within the endothelial cells of small blood vessels of the cerebrum increased by 254-fold in comparison to other areas of the brain. This ranked ELTD1 first as a potential marker for microvascular development, making it a particularly interesting target for future therapeutic approaches to target angiogenesis. ELTD1 was also found to be expressed in nonpathological samples from tissues, organs, or cell lines belonging to murine models and the results were reported in The Human Protein Atlas and the Biogps Online Database. Although results shown in each dataset vary, the highest levels of ELTD1 gene expression were obtained from the same organs, tissues, and cell lines: lungs, hematopoietic stem cells, common myeloid progenitors, heart, adrenal glands, kidneys, or granulomonocyte progenitors (). The relative abundance of ELTD1 in some tissues may prove valuable in the future, unlocking further steps into understanding how angiogenesis differs in different organs. It also hints to a hope that ELTD1 may play a larger role in angiogenesis, not only in the brain, but also in other major organs.
Figure 2 ELTD1 gene expression in tissues, organs, or cell lines in the 1,418,058 (A) and 1,418,059 (B) databases. The samples were obtained from mice. Data were obtained from the Biogps online database (http://biogps.org/#goto=genereport&id=170,757, accessed October 17, 2015).Citation64 Results are represented as the 20 most highly expressed gene levels for each probe in order to highlight the most relevant data.
ELTD1 roles in cancer
GBM has long been considered an immense obstacle for modern medicine, being both the most common and the most aggressive brain tumor to date.Citation65 Considered a grade IV glioma by the World Health Organization, GBM has an extremely poor prognosis with a vast majority of patients dying within 2 years of diagnosis, with less than 5% of them surviving more than 3 years.Citation66
Therapeutic approaches involving chemotherapy, radiotherapy, targeted therapy, and/or combined regimes have so far yielded very limited results with a poor contribution to GBM patient survival.Citation67 So far, no viable biomarker has been selected as a strong candidate among the numerous gene alterations, protein overexpressions/suppressions, and abnormal signaling pathways in glioblastoma.Citation28 One of the most well-known markers for GBM is the epithelial growth factor receptor (EGFR). EGFR amplification, along with mutations such as EGFRvIII/ΔEGFR, has long been considered as poor prognostic markers for GBM patients, but also viable therapeutic targets.Citation68–Citation70 Other biomarkers include mutations of the isocitrate dehydrogenase enzyme (IDH) with its two isoforms IDH1 and IDH2, considered the genetic signature for secondary GBMs,Citation71 O6-methylguanine DNA methyltransferase protein expression and methylation status, which are considered markers for slower tumor progression,Citation72 PTEN gene mutation.Citation73,Citation74 Circulating biomarkers such as extracellular vesicles were recently mentioned as valuable biomarkers for GBM.Citation75 Extracellular vesicles are membrane-surrounded structures, secreted by cells in the biological fluids and consist of a mixture of several proteins, lipids, nucleic acids, glycans, and other biomolecules. These metabolites are suggested to be tumor-specific, thus considered to be good circulating biomarkers for GBM progression and response to the therapy.Citation75 In a study by Towner et alCitation29 conducted on human and rodent glioma samples, higher levels of ELTD1 were found in high-grade gliomas compared with low-grade gliomas. Higher levels of ELTD1 were also associated with the mesenchymal subtype compared with the proneural subtype. However, the authors didn’t find any correlations between ELTD1 levels and survival rates in both the human and murine models.Citation29
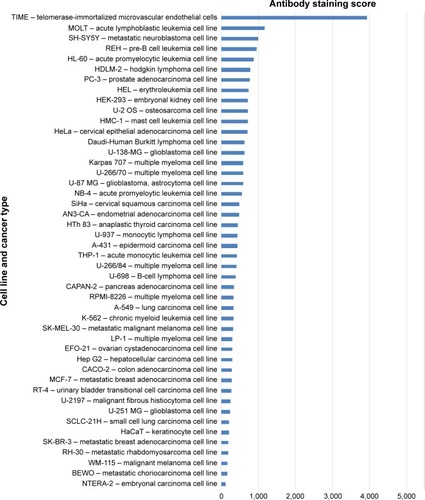
To analyze things from a broader perspective, an online database has also compared various tumor cell lines to see how ELTD1 levels correlate with different types of cancer (). Surprisingly, in contrast with the previous studies we mentioned, the results placed ELTD1 expression in GBM lines midway through the list. More surprisingly, immunohistochemical staining did not find any ELTD1 levels in glioma tumor samples (http://www.proteinatlas.org/ENSG00000162618-ELTD1/cancer, accessed October 17, 2015). However, it should be noted that the database refers only to staining in glioma, without mentioning if the samples were taken from high- or low-grade gliomas. As Towner et alCitation29 pointed out, higher levels of ELTD1 were associated with the more aggressive, high-grade gliomas in contrast to low-grade ones.
Figure 3 ELTD1 protein levels in human cancer cell lines. The results show the intensity of antibody staining (protein levels) for each cell line. Results were obtained from the Human Protein Atlas Database (http://www.proteinatlas.org/ENSG00000162618-ELTD1/cell/HPA025229, accessed October 17, 2015).Citation77
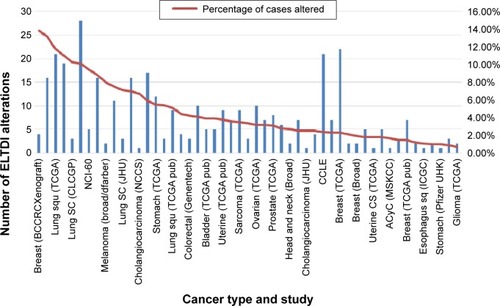
In order to better understand how the receptor works, alterations to the ELTD1 genetic sequence in different cancers were also taken into consideration (). By comparison with other cancer types, ELTD1 alterations were almost nonexistent in malignant gliomas. These findings suggest that while a high level of ELTD1 expression may be indicative of the presence or aggressiveness of a malignant glioma, mutation levels are not linked to any aspect of tumor evolution.
Figure 4 ELTD1 genetic alterations (mutations, amplifications, and deletions) in different human tumor samples. Results are shown as the number of alterations and percentage of alterations per samples analyzed in each individual study and only shows cancer types where genetic alterations are present. The results were obtained from the BioPortal for Cancer Genomics database (http://bit.ly/1PyuYcR, accessed October 17, 2015).Citation78
ELTD1 has also been implicated in the angiogenesis of other types of cancer. A study of various tumor samples (renal, colorectal, ovarian, and head and neck) showed a match between the upregulation of the gene encoding ELTD1 and tumor development.Citation19 Tissues of tumoral origin presented higher levels of protein expression when compared with normal tissue samples, while vascular smooth muscle cells showed higher ELTD1 staining when compared with endothelial cells. Increased ELTD1 staining was also correlated with higher microvascular density levels and reduced hypoxia signature in some forms of cancer/tumor.Citation19 The article also points out that ELTD1 impairment in endothelial cells halved the tumor cells’ capacity to sprout new microvessels. Furthermore, Masiero et al demonstrated, using both in vivo and in vitro models, that the two main antagonistic pathways involved in angiogenesis (VEGF/bFGF and DLL4-Notch)Citation60–Citation62 had an opposing effect on ELTD1 levels (the former increased ELTD1 expression while the latter determined the exact opposite)Citation19 (). A study by Dieterich et alCitation76 also demonstrated that in human glioblastomas, increased VEGF-A signaling produced higher levels of ELTD1 gene expression in endothelial cells. Perhaps with a better understanding of the processes behind how ELTD1 works, which are currently unknown, such as ligand–receptor interaction and signaling pathways or the effect mutations have on the receptor function, we may draw a parallel between ELTD1 and the well-studied VEGF and in terms of involvement in both normal and neoplastic angiogenesis. However, because of the scarcity of information regarding ELTD1 due to the relatively recent discovery of the receptor, further studies are required to confirm its involvement in these mechanisms.
Conclusion
Recent studies have linked ELTD1 levels to both normal and pathological angiogenesis, such as cardiac hypertrophy in mice and even the development of nouvelle microvessels in various cancer types. One association that stands out is the link between ELTD1 and the evolution of GBM, recently pointed out in various studies. If proven feasible, this association may provide not only a viable biomarker for this highly aggressive cancer, but also a target for future therapeutic agents. These therapeutic approaches should focus on either silencing the ELTD1 gene using methods such as small interfering RNA or blocking the receptor with antibodies. However, with little knowledge about the ligands binding to the receptor or the pathways involved in downstream signal transduction, targeting ELTD1 might still prove problematic.
With several other roles besides vascular proliferation in cancer such as graft-versus-host disease, cannabis addiction, obesity or adaption to altitude in certain populations and notably the presence in a large number of normal tissues, ELTD1 may play a larger role than expected in the human body. Hopefully, increased attention to this receptor may shed some light on its multiple functions leading to a better understanding on how some pathological mechanisms are interrelated and how they can be tackled.
Author contributions
All authors contributed toward the literature search, data analysis, drafting and critically revising the paper and agree to be accountable for all aspects of the work.
Acknowledgments
The paper was financed by “Program of Excellence in multidisciplinary doctoral and postdoctoral research in chronic diseases”, Contract No POSDRU/159/1.5/S/133377, cofinanced project from the European Social Fund by Operational Sectoral Programme Human Resources Development 2007–2013 and Executive Agency for Higher Education, Research, Development and Innovation Funding Romania, Grant PN-II-ID-PCE-2011-3-1041, Romania.
Disclosure
The authors report no conflicts of interest in this work.
References
- WallgardELarssonEHeLIdentification of a core set of 58 gene transcripts with broad and specific expression in the microvasculatureArterioscler Thromb Vasc Biol20082881469147618483404
- van BeijnumJRNowak-SliwinskaPHuijbersEJThijssenVLGriffioenAWThe great escape; the hallmarks of resistance to antiangiogenic therapyPharmacol Rev201567244146125769965
- McIntyreAHarrisALMetabolic and hypoxic adaptation to anti-angiogenic therapy: a target for induced essentialityEMBO Mol Med20157436837925700172
- RapisardaAMelilloGOvercoming disappointing results with anti-angiogenic therapy by targeting hypoxiaNat Rev Clin Oncol20129737839022525710
- Glade BenderJCooneyEMKandelJJYamashiroDJVascular remodeling and clinical resistance to antiangiogenic cancer therapyDrug Resist Updat200474–528930015533766
- BergersGHanahanDModes of resistance to anti-angiogenic therapyNat Rev Cancer20088859260318650835
- FieldKMJordanJTWenPYRosenthalMAReardonDABevacizumab and glioblastoma: scientific review, newly reported updates, and ongoing controversiesCancer20151217997100725263092
- HaalandBChopraAAcharyyaSFayAPLopes GdeLComparative effectiveness of approved first-line anti-angiogenic and molecularly targeted therapeutic agents in the treatment of good and intermediate risk metastatic clear cell renal cell carcinomaBMC Cancer20141459225127891
- PradeepCRSunilaESKuttanGExpression of vascular endothelial growth factor (VEGF) and VEGF receptors in tumor angiogenesis and malignanciesIntegr Cancer Ther20054431532116282508
- ByrneAMBouchier-HayesDJHarmeyJHAngiogenic and cell survival functions of vascular endothelial growth factor (VEGF)J Cell Med200594777794
- TandleALibuttiSKAntiangiogenic therapy: targeting vascular endothelial growth factor and its receptorsClin Adv Hematol Oncol200311414816227959
- FontanellaCOngaroEBolzonelloSGuardascioneMFasolaGAprileGClinical advances in the development of novel VEGFR2 inhibitorsAnn Transl Med201421212325568876
- AprileGRijavecEFontanellaCRihawiKGrossiFRamucirumab: preclinical research and clinical developmentOnco Targets Therapy2014719972006
- SharmaTDhingraRSinghSAflibercept: a novel VEGF targeted agent to explore the future perspectives of anti-angiogenic therapy for the treatment of multiple tumorsMini Rev Med Chem201313453054023317499
- ZhaoYAdjeiAATargeting angiogenesis in cancer therapy: moving beyond vascular endothelial growth factorOncologist20152066067326001391
- AlessiPLealiDCamozziMCantelmoAAlbiniAPrestaMAnti-FGF2 approaches as a strategy to compensate resistance to anti-VEGF therapy: long-pentraxin 3 as a novel antiangiogenic FGF2-antagonistEur Cytokine Netw200920422523420167562
- MedingerMPasswegJAngiogenesis in myeloproliferative neoplasms, new markers and future directionsMemo2014720621025544863
- SecordAANixonABHurwitzHIThe search for biomarkers to direct antiangiogenic treatment in epithelial ovarian cancerGynecol Oncol2014135234935825178997
- MasieroMSimoesFCHanHDA core human primary tumor angiogenesis signature identifies the endothelial orphan receptor ELTD1 as a key regulator of angiogenesisCancer cell201324222924123871637
- FavaraDMBanhamAHHarrisALA review of ELTD1, a pro-angiogenic adhesion GPCRBiochem Soc Trans20144261658166425399586
- HarkenseeCOkaAOnizukaMMicrosatellite scanning of the immunogenome associates MAPK14 and ELTD1 with graft-versus-host disease in hematopoietic stem cell transplantationImmunogenetics201365641742723474535
- PekowJDoughertyUHuangYGene signature distinguishes patients with chronic ulcerative colitis harboring remote neoplastic lesionsInflamm Bowel Dis201319346147023388545
- EichstaedtCAAntaoTPaganiLCardonaAKivisildTMorminaMThe Andean adaptive toolkit to counteract high altitude maladaptation: genome-wide and phenotypic analysis of the CollasPLoS One201493e9331424686296
- AgrawalAPergadiaMLSacconeSFAn autosomal linkage scan for cannabis use disorders in the nicotine addiction genetics projectArch Gen Psychiatry200865671372118519829
- LeeKTByunMJKangKSNeuronal genes for subcutaneous fat thickness in human and pig are identified by local genomic sequencing and combined SNP association studyPLoS One201162e1635621311593
- Porto NetoLRBunchRJHarrisonBEBarendseWDNA variation in the gene ELTD1 is associated with tick burden in cattleAnim Genet2011421505520880337
- NechiporukTUrnessLDKeatingMTETL, a novel seven-transmembrane receptor that is developmentally regulated in the heart. ETL is a member of the secretin family and belongs to the epidermal growth factor-seven-transmembrane subfamilyJ Biol Chem200127664150415711050079
- NicolaidisSBiomarkers of glioblastoma multiformeMetabolism2015643 Suppl 1S22S2725468141
- TownerRAJensenRLColmanHELTD1, a potential new biomarker for gliomasNeurosurgery2013721779023096411
- EoHSChoiJPNohSJHurCGKimWA combined approach for the classification of G protein-coupled receptors and its application to detect GPCR splice variantsComput Biol Chem200731424625617631418
- BaldwinJMStructure and function of receptors coupled to G proteinsCurr Opin Cell Biol1994621801908024808
- LefkowitzRJThe superfamily of heptahelical receptorsNat Cell Biol200027E133E13610878827
- SchiothHBFredrikssonRThe GRAFS classification system of G-protein coupled receptors in comparative perspectiveGen Comp Endocrinol20051421–29410115862553
- FormstoneCJ7TM-Cadherins: developmental roles and future challengesAdv Exp Med Biol2010706143621618823
- KaurBBratDJDeviNSVan MeirEGVasculostatin, a proteolytic fragment of brain angiogenesis inhibitor 1, is an antiangiogenic and antitumorigenic factorOncogene200524223632364215782143
- ShiratsuchiTNishimoriHIchiseHNakamuraYTokinoTCloning and characterization of BAI2 and BAI3, novel genes homologous to brain-specific angiogenesis inhibitor 1 (BAI1)Cytogenet Cell Genet1997791–21031089533023
- PickeringCHagglundMSzmydynger-ChodobskaJThe Adhesion GPCR GPR125 is specifically expressed in the choroid plexus and is upregulated following brain injuryBMC Neurosci200899718834514
- KwakkenbosMJKopENStaceyMThe EGF-TM7 family: a postgenomic viewImmunogenetics2004551065566614647991
- HamannJKoningNPouwelsWEMR1, the human homolog of F4/80, is an eosinophil-specific receptorEur J Immunol200737102797280217823986
- OdaKShiratsuchiTNishimoriHIdentification of BAIAP2 (BAI-associated protein 2), a novel human homologue of hamster IRSp53, whose SH3 domain interacts with the cytoplasmic domain of BAI1Cytogenet Cell Genet1999841–2758210343108
- GaoFBBrenmanJEJanLYJanYNGenes regulating dendritic outgrowth, branching, and routing in DrosophilaGenes Dev199913192549256110521399
- UsuiTShimaYShimadaYFlamingo, a seven-pass trans-membrane cadherin, regulates planar cell polarity under the control of FrizzledCell199998558559510490098
- LiWGaoFBActin filament-stabilizing protein tropomyosin regulates the size of dendritic fieldsJ Neurosci200323156171617512867499
- SeifertJRMlodzikMFrizzled/PCP signalling: a conserved mechanism regulating cell polarity and directed motilityNat Rev Genet20078212613817230199
- DevenportDOristianDHellerEFuchsEMitotic internalization of planar cell polarity proteins preserves tissue polarityNat Cell Biol201113889390221743464
- FredrikssonRLagerstromMCLundinLGSchiothHBThe G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprintsMol Pharmacol20036361256127212761335
- KoiralaSJinZPiaoXCorfasGGPR56-regulated granule cell adhesion is essential for rostral cerebellar developmentJ Neurosci200929237439744919515912
- YonaSLinHHDriPLigation of the adhesion-GPCR EMR2 regulates human neutrophil functionFASEB J200822374175117928360
- KohJTKookHKeeHJExtracellular fragment of brain-specific angiogenesis inhibitor 1 suppresses endothelial cell proliferation by blocking alphavbeta5 integrinExp Cell Res2004294117218414980512
- CorkSMKaurBDeviNSA proprotein convertase/MMP-14 proteolytic cascade releases a novel 40 kDa vasculostatin from tumor suppressor BAI1Oncogene201231505144515222330140
- CapassoMDurrantLGStaceyMGordonSRamageJSpendloveICostimulation via CD55 on human CD4+ T cells mediated by CD97J Immunol200617721070107716818763
- HuangYSChiangNYHuCHActivation of myeloid cell-specific adhesion class G protein-coupled receptor EMR2 via ligation-induced translocation and interaction of receptor subunits in lipid raft micro-domainsMol Cell Biol20123281408142022310662
- LeemansJCte VeldeAAFlorquinSThe epidermal growth factor-seven transmembrane (EGF-TM7) receptor CD97 is required for neutrophil migration and host defenseJ Immunol200417221125113114707087
- CullenMElzarradMKSeamanSGPR124, an orphan G protein-coupled receptor, is required for CNS-specific vascularization and establishment of the blood-brain barrierProc Nati Acad Sci U S A20111081457595764
- AndersonKDPanLYangXMAngiogenic sprouting into neural tissue requires Gpr124, an orphan G protein-coupled receptorProc Nati Acad Sci U S A2011108728072812
- WangTWardYTianLCD97, an adhesion receptor on inflammatory cells, stimulates angiogenesis through binding integrin counter-receptors on endothelial cellsBlood200510572836284415576472
- KuhnertFMancusoMRShamlooAEssential regulation of CNS angiogenesis by the orphan G protein-coupled receptor GPR124Science2010330600698598921071672
- BjarnadottirTKFredrikssonRHoglundPJGloriamDELagerstromMCSchiothHBThe human and mouse repertoire of the adhesion family of G-protein-coupled receptorsGenomics2004841233315203201
- FlicekPAmodeMRBarrellDEnsembl 2014Nucleic Acids Research201442Database issueD749D75524316576
- LiJLHarrisALCrosstalk of VEGF and Notch pathways in tumour angiogenesis: therapeutic implicationsFront Biosci20091430943110
- ThurstonGKitajewskiJVEGF and Delta-Notch: interacting signalling pathways in tumour angiogenesisBr J Cancer20089981204120918827808
- HellstromMPhngLKGerhardtHVEGF and Notch signaling: the yin and yang of angiogenic sproutingCell Adh Migr20071313313619262131
- XiaoJJiangHZhangRAugmented cardiac hypertrophy in response to pressure overload in mice lacking ELTD1PLoS One201275e3577922606234
- WuCOrozcoCBoyerJBioGPS: an extensible and customizable portal for querying and organizing gene annotation resourcesGenome Biol20091011R13019919682
- GrossmanSABataraJFCurrent management of glioblastoma multiformeSemin Oncol200431563564415497116
- KrexDKlinkBHartmannCLong-term survival with glioblastoma multiformeBrain2007130Pt 102596260617785346
- StuppRHegiMEMasonWPEffects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trialLancet Oncol200910545946619269895
- SangarVFunkCCKusebauchUCampbellDSMoritzRLPriceNDQuantitative proteomic analysis reveals effects of epidermal growth factor receptor (EGFR) on invasion-promoting proteins secreted by glioblastoma cellsMol Cell Proteomics201413102618263124997998
- GanHKCvrljevicANJohnsTGThe epidermal growth factor receptor variant III (EGFRvIII): where wild things are alteredFEBS J2013280215350537023777544
- CrespoIVitalALGonzalez-TablasMMolecular and Genomic Alterations in Glioblastoma MultiformeAm J Pathol201518571820183325976245
- StieberDAbdul RahimSANiclouSPNovel ways to target brain tumour metabolismExpert Opin Ther Targets201115101227123921635150
- SonodaYYokosawaMSaitoRO(6)-Methylguanine DNA methyltransferase determined by promoter hypermethylation and immunohistochemical expression is correlated with progression-free survival in patients with glioblastomaInt J Clin Oncol201015435235820232102
- PhillipsHSKharbandaSChenRMolecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesisCancer Cell20069315717316530701
- CohenALColmanHGlioma biology and molecular markersCancer Treat Res2015163153025468223
- MahmoudiKEzrinAHadjipanayisCSmall extracellular vesicles as tumor biomarkers for glioblastomaMol Aspects Med2015459710226118341
- DieterichLCMellbergSLangenkampETranscriptional profiling of human glioblastoma vessels indicates a key role of VEGF-A and TGFbeta2 in vascular abnormalizationJ Pathol2012228337839022786655
- UhlenMOksvoldPFagerbergLTowards a knowledge-based Human Protein AtlasNat Biotechnol201028121248125021139605
- CeramiEGaoJDogrusozUThe cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics dataCancer Discov20122540140422588877
