Abstract
Breast cancer is the most common cancer among women, accounting for about 30% of all cancers. In contrast, breast cancer is a rare disease in men, accounting for less than 1% of all cancers. Up to 10% of all breast cancers are hereditary forms, caused by inherited germ-line mutations in “high-penetrance,” “moderate-penetrance,” and “low-penetrance” breast cancer susceptibility genes. The remaining 90% of breast cancers are due to acquired somatic genetic and epigenetic alterations. A heterogeneous set of somatic alterations, including mutations and gene amplification, are reported to be involved in the etiology of breast cancer. Promoter hypermethylation of genes involved in DNA repair and hormone-mediated cell signaling, as well as altered expression of micro RNAs predicted to regulate key breast cancer genes, play an equally important role as genetic factors in development of breast cancer. Elucidation of the inherited and acquired genetic and epigenetic alterations involved in breast cancer may not only clarify molecular pathways involved in the development and progression of breast cancer itself, but may also have an important clinical and therapeutic impact on improving the management of patients with the disease.
Introduction
Breast cancer is currently the most common cancer among women, accounting for about 30% of all cancers.Citation1 In contrast, breast cancer is a rare disease in men, accounting for less than 1% of all cancers.Citation2 The age-specific incidence rates for breast cancer in women increase rapidly until the age of 50 years, and then increase at a slower rate for older women, while incidence rates for breast cancer in men increase linearly and steadily with age. Overall, current epidemiologic and pathologic data, such as age-frequency distribution, age-specific incidence rate patterns, and prognostic factor profiles, suggest that male breast cancer is similar to postmenopausal female breast cancer.Citation2 It is generally accepted that breast cancer may represent the same disease entity in both genders, and common hormonal, genetic, and environmental risk factors are involved in the pathogenesis of breast cancer in women and men. Hormonal changes, such as increased estrogen exposure due to diabetes, obesity or liver disease, and environmental and lifestyle factors, such as carcinogen exposure or alcohol intake, are associated with risk of developing breast cancer.Citation3 However, the major predisposition factor for breast cancer is a positive family history of the disease. Patients of both genders with a positive first-degree family history have a twofold increased risk, which increases to more than fivefold with the number of affected relatives and early onset relatives, thus suggesting a relevant genetic component in breast cancer risk.Citation4 It is estimated that up to 10% of all breast cancers are hereditary forms, caused by inherited germ-line mutations in breast cancer susceptibility genes. Commonly, inherited mutations are loss-of-function mutations that occur in tumor suppressor genes involved in DNA repair and cell cycle checkpoint activation.Citation1 The remaining 90% of breast cancers are due to acquired somatic, genetic, and epigenetic alterations.Citation5 Genetic alterations include gain-of-function mutations, amplification, deletions, and rearrangements occurring in genes which stimulate cell growth, division, and survival.Citation6 Epigenetic deregulation, mainly due to promoter methylation, may also contribute to the abnormal expression of these genes.Citation7 In addition, the involvement of micro RNAs (miRNAs) in modulating gene expression in the development of breast cancer has been recently reported.Citation8 The focus of this review will be on the most relevant inherited and acquired alterations in the development of breast cancer in both genders. We did a systematic literature search using PubMed to provide a synopsis of the current understanding and future directions of research in this field. We selected original articles and reviews published up to April 2011. The following search key terms were used to query the PubMed website: “inherited breast cancer,” “breast cancer AND susceptibility,” “breast cancer AND somatic alterations,” “breast cancer AND epigenetic,” “breast cancer AND miRNA.” The abstracts resulting from these queries were individually analyzed for relevance.
Inherited susceptibility to breast cancer
To date, 5%–10% of all breast cancers are caused by inherited germ-line mutations in well identified breast cancer susceptibility genes.Citation1 According to their mutation frequency and the magnitude of their impact in breast cancer susceptibility, these genes can be divided into “high-penetrance,” “moderate-penetrance,” and “low-penetrance” genes ().Citation9 Variants in the two major high-risk breast cancer genes, ie, BRCA1 and BRCA2, occur rarely in the population, but confer a high risk of breast cancer to the individual.Citation1P53 and PTEN, two genes involved in rare syndromes (Li-Fraumeni and Cowden syndromes, respectively), also confer a high risk of breast cancer.Citation1 However, P53 and PTEN germ-line mutations are very rare, and it is unlikely that these mutations would account for a proportion of breast cancers in the absence of their respective syndromes.Citation10,Citation11
Figure 1 Genetic susceptibility in hereditary breast cancer. Up to 10% of all breast cancers are caused by inherited germ-line mutations in breast cancer susceptibility genes. High-penetrance genes (BRCA1 and BRCA2) contribute to 25% of hereditary breast cancer, moderate-penetrance genes (CHEK2, ATM, PALB2, BRIP1, RAD51C) contribute less than 5% to the risk of breast cancer. The great majority of hereditary breast cancer may be due to common low-penetrance alleles or other still unknown genetic factors.
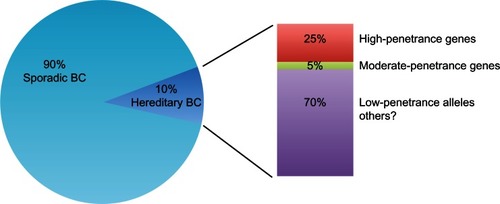
Overall, high-risk genes account for about 25% of inherited breast cancers ().Citation12 Variants in genes functionally related to BRCA1/2 in DNA repair pathways confer an intermediate risk of breast cancer. These variants are rare, occurring in less than 1% of the population, and their contribution to the risk of breast cancer is less than 5% ().Citation13 Recently, a third class of low-penetrance susceptibility alleles has been identified. These alleles, which may occur in genic or nongenic regions, confer a lower risk but are very common in the population.Citation13 Due to their low penetrance, the real contribution of these common variants to breast cancer risk is not entirely clear (). Overall, this scenario suggests that the majority of genetic factors involved in breast cancer susceptibility are still unknown.
High-penetrance breast cancer genes
BRCA1 and BRCA2 are the most important breast cancer susceptibility genes in high-risk families (). BRCA1/2 mutations are considered to be responsible for approximately 30% of breast cancer cases with a family history of breast/ovarian cancer, and it has been estimated that inherited BRCA1 and BRCA2 mutations account for 5%–10% of the total percentage of breast cancer.Citation14–Citation16
Table 1 Classes of genetic susceptibility and comparison of their different features
In women, germ-line BRCA1 and BRCA2 mutations confer a high risk for developing breast cancer by age 70 years. Initial studies based on multiple-case families, reported a female breast cancer risk at age 70 years in BRCA1 and BRCA2 mutation carriers of 85% and 84%, respectively.Citation17 Later meta-analyses showed that the average cumulative female breast cancer risk in BRCA1 mutation carriers by 70 years of age, unselected for family history, was 46%–65% and the corresponding estimates for BRCA2 were 43%–45%.Citation18,Citation19
In male breast cancer cases, BRCA2 mutations are much more common than BRCA1. Mutations in the BRCA2 gene are estimated to be responsible for 60%–76% of male breast cancers occurring in high-risk breast cancer families, whereas the BRCA1 mutation frequency ranges from 10% to 16%.Citation17,Citation20 In a large population-based male breast cancer series, we reported a mutation frequency of about 7% and 2% for BRCA2 and BRCA1, respectively.Citation21 Interestingly, a founder effect was observed in BRCA1-associated male breast cancer cases.Citation22
All known BRCA1/2 mutations are recorded in the Breast Information Core database (http://www.nhgri.nih.gov/Intramural_research/Lab_transfer/Bic/). To date, 1643 distinct germ-line BRCA1 mutations and 2015 BRCA2 mutations have been reported in the database. The great majority of BRCA1/2 mutations in breast cancer are predicted to truncate the protein product. The most common type of mutations are small frameshift insertions or deletions, nonsense mutations, or mutations affecting splice sites resulting in a deletion of complete or partial exons or insertion of intronic sequences. The Breast Cancer Linkage Consortium has reported that approximately 70% of BRCA1 mutations and 90% of BRCA2 mutations in linked families are truncating mutations.Citation4 In addition to truncating mutations, an elevated number of missense variants has been identified. The most frequent are the BRCA1 G61C in the RING-finger codon and the BRCA2 I2490T in exon 15.
Some studies also indicate that BRCA1/2 polymorphic variants could be associated with an increased risk of breast and ovarian cancer.Citation23,Citation24 Association between the BRCA2 N372H variant and increased breast cancer risk in particular has been reported from population-based studies.Citation25 Interestingly, we observed an association between the BRCA2 N372H variant and risk of male breast cancer in young patients.Citation26 Breast cancer risk in women is influenced by the position of the mutation within the gene sequence. Women with a mutation in the central region of BRCA1 were shown to have a lower breast cancer risk than women with mutations outside this region. BRCA2 mutations located in the central region, referred to as the ovarian cancer cluster region, also appear to be associated with a lower breast cancer risk and a higher ovarian cancer risk than other BRCA2 mutations.Citation27,Citation28
Specific BRCA1 and BRCA2 mutations show a high frequency in specific countries or ethnic groups, particularly in genetically isolated populations. These mutations descend from a single founder. For example, two founder mutations in BRCA1 (185delAG and 5382insC) and one in BRCA2 (6174delT) account for the majority of all BRCA1/2 mutations (>90%) in the Ashkenazi Jewish population.Citation29BRCA1 185delAG is present in about 1% of Ashkenazi Jews and in 20% of Ashkenazi women affected by breast cancer before the age of 50 years.Citation18,Citation30 A single BRCA2 mutation (999del5) has been found in the majority of multiple-case breast cancer families in the Icelandic population.Citation31,Citation32 The BRCA2 999del5 accounts for about 8% of female breast cancer, rising to 24% of female breast cancers before the age of 40 years, and for about 38% of male breast cancers.Citation32,Citation33 In Italy, a historically and genetically heterogeneous country, BRCA1/2 founder mutations are found in small isolated geographic areas. The BRCA1 5083del19 was found in a geographically homogeneous population from Calabria, a region of Southern Italy, where it accounts for 33% of overall gene mutations.Citation34 A high frequency of BRCA1 5083del19 mutation has also been identified in the population of Sicily, a region near to Calabria.Citation35 Other regional founder mutations have been reported in Tuscany (BRCA1 1499insA, BRCA1 3347delAG), a region in Central Italy of ancient settlement.Citation21,Citation22,Citation36
In addition to point mutations, small deletions and insertions, large-scale BRCA1/2 rearrangements, including insertions, deletions, or duplications of more than 500 kb of DNA, have been identified in both male and female breast cancer.Citation37–Citation40 Large genomic rearrangements may account for 3%–15% of all BRCA1 and BRCA2 mutations.Citation38 The higher density of Alu repeat sequences in BRCA1 and both Alu and non-Alu repetitive DNA in BRCA2 are thought to contribute to the large number of deletions and duplications observed in these genes.Citation37,Citation40–Citation43 The frequency of large BRCA1 genomic rearrangements in families with a strong family history of breast and/or ovarian cancer, varies greatly (0%–36%) in different populations.Citation37–Citation39 The frequency of large BRCA2 genomic rearrangements seems to be lower (1%–2%) in comparison with BRCA1.Citation41,Citation44 Interestingly, large genomic rearrangements in BRCA2 are more frequent in families with male breast cancer.Citation38,Citation43
Moderate-penetrance breast cancer genes
Overall, fewer than 10% of breast cancers are attributable to known mutations in the breast cancer susceptibility genes, BRCA1 and BRCA2.Citation13 Recently, direct interrogation of candidate genes involved in BRCA1/2-associated DNA damage repair pathway has led to the identification of other breast cancer susceptibility genes, classified as moderate-penetrance genes (). Variants found in this class of genes confer a smaller risk of breast cancer than BRCA1/2 and, because of their rarity, are very difficult to detect in the population. Overall, mutations in moderate-penetrance genes account for less than 3% of the familial risk of breast cancer.Citation13
CHEK2 1100delC was the first moderate breast cancer risk allele identified and was associated with a twofold risk among breast cancer cases unselected for family history and fivefold among familial breast cancer cases.Citation45,Citation46 The CHEK2 1100delC mutation has also been shown to confer approximately a tenfold increase in breast cancer risk in men lacking BRCA1/2 mutations, and was estimated to account for 9% of familial high-risk male breast cancer cases.Citation45 However, this association is not so evident in male breast cancer series unselected for family history, in which it was reported that the CHEK2 1100delC is unlikely to account for a significant proportion of male breast cancer cases.Citation47–Citation50 The contribution of the CHEK2 1100delC mutation to breast cancer predisposition in both genders varies by ethnic group and from country to country. A decreased frequency of the 1100delC allele in North to South orientation has been observed in Europe both for male and female breast cancer.Citation50–Citation53 Identification of the CHEK2 1100delC mutation as a breast cancer-associated allele induced mutational screening of the whole CHEK2 gene sequence. However, at present, only a small number of rare truncating mutations and missense variants have been reported in breast cancer cases.Citation54,Citation55
ATM was first proposed as a breast cancer predisposition gene by epidemiological studies that reported an increased breast cancer risk in relatives of patients with ataxia telangiectasia, a recessive syndrome caused by mutation in the ATM gene.Citation56,Citation57 However, molecular data corroborating this observation were provided after 20 years.Citation58 To date, many truncating splice site mutations and missense variants for ATM have been found and associated with a relative risk of breast cancer of about 2.4.Citation58 Currently there are no data about the role of ATM in men predisposed to breast cancer.
The involvement of BRCA2 in the Fanconi anemia pathway promoted mutation screening of other Fanconi anemia genes functionally linked to BRCA2, such as PALB2, BRIP1, and, more recently, RAD51C.Citation59PALB2 truncating mutations were estimated to be associated with a 2.3-fold increased risk.Citation60PALB2 mutations have now been identified in many countries, with frequencies varying from 0.6% to 2.7% in familial breast cancer cases.Citation61–Citation69 Two founder PALB2 mutations, 1592delT and 2323C > T, were respectively identified in 1% of Finnish and 0.5% of French-Canadian breast cancers unselected for family history.Citation70,Citation71 Interestingly, PALB2 mutations were found in families with both female and male breast cancer cases, suggesting that PALB2 may be involved in male breast cancer risk.Citation60,Citation67 To date, five studies have reported on the frequency of PALB2 mutations in male breast cancer.Citation70,Citation72–Citation75 Overall, PALB2 seems to have a role as a moderate-penetrance gene in male breast cancer to a comparable extent as for female breast cancer. Recently, it has been reported that PALB2 heterozygote mutation carriers were four times more likely to have a male relative with breast cancer.Citation76
Deleterious BRIP1 mutations were initially estimated to confer a twofold increased breast cancer risk and to account for about 1% of BRCA1/2 negative familial/early-onset breast cancer cases, but further studies suggested that the BRIP1 contribution to breast cancer susceptibility might be more limited than initially reported.Citation77–Citation80 Indeed, a total of only eight BRIP1 truncating mutations in 11 BRCA1/2 mutation-negative breast cancer cases from three independent studies have been identified worldwide.Citation77,Citation81,Citation82 Several studies in diverse populations failed to detect truncating mutations.Citation78–Citation80 To date, only one study has investigated the role of BRIP1 in male breast cancer susceptibility, and no evidence was found that germ-line variants in BRIP1 might contribute to male breast cancer predisposition.Citation83 Taken together, these data suggest that the contribution of BRIP1 to breast cancer predisposition in both females and males is less consistent compared with other moderate breast cancer susceptibility genes, such as CHEK2 and PALB2.
Recently, mutations in RAD51C, another gene associated with Fanconi anemia, were identified as breast cancer susceptibility alleles, accounting for 1.3% of female patients from families with at least one case each of breast and ovarian cancer.Citation84 However, further studies did not confirm this frequency.Citation85,Citation86 At present, there is no evidence that RAD51C mutations contribute to male breast cancer susceptibility.Citation87
Low-penetrance breast cancer genes
A polygenic model, in which many genes that confer low risk individually act in combination to confer much larger risk in the population, has been suggested for susceptibility to breast cancer and other common cancers.Citation88 Breast cancers unaccounted for by currently known high-penetrance and moderate-penetrance breast cancer susceptibility genes can be explained by this model. This hypothesis, speculated for years, has only recently been confirmed by multigroup collaborations working in genome-wide association studies performed in a very large series of cases and controls from different countries, in order to increase the power to detect small effects on risk.Citation89,Citation90 Common low-penetrance breast cancer susceptibility single nucleotide polymorphisms have thus far been reported in regions that cover known protein-coding genes, including CASP8, FGFR2, TNRC9, MAP3K1, LSP1, RAD51L1, and ESR1 and in regions such as 8q24, 2q35, 5p12, and 1p11 with no known protein-coding genes ().Citation90–Citation95 The relative risk conferred by these alleles ranges from 1.07 to 1.26. Overall, these single nucleotide polymorphisms are estimated to account for less than 4% of the familial risk of breast cancer in women.Citation90 Interestingly, many of the alleles are in intronic portions of genes, and often are noncoding regions that may confer susceptibility. This might be explained by the observation that some of these loci are located in regions of linkage disequilibrium that cover different genes, but it is very difficult to establish which of a set of variants in linkage disequilibrium is the most functionally relevant.Citation96 Furthermore, some of these single nucleotide polymorphisms, including CASP8, FGFR2, TOX3, and MAP3K1, could act as modulators of the risk conferred by mutations in the high-penetrance breast cancer susceptibility genes, BRCA1 and BRCA2.Citation97 Recently, a subtle regulatory effect of one allele in the prostate/breast cancer-associated 8q24 block was also demonstrated, which acts as a cis enhancer of the MYC promoter.Citation98 Interestingly, different haplotype blocks within 8q24 were specifically associated with risk of different cancers.Citation99 In particular, four blocks were site-specific (one for breast cancer and three for prostate cancer), and a fifth was a multicancer susceptibility marker because it was associated with a risk of various cancers, including prostate, colon, ovarian, kidney, thyroid, and laryngeal cancer, but not breast cancer.Citation99–Citation101 None of the presently identified loci is directly linked to the DNA repair pathway. Instead, many of the coding loci are in genes somatically mutated in diverse cancers, including breast cancer. Recently, it was observed that genetic germ-line variations in genes encoding for “driver kinases” may influence breast cancer risk, thus suggesting that low-penetrance alleles might be a link between germ-line and somatic alterations in breast cancer.Citation102
Specific single nucleotide polymorphisms seem to be associated with specific clinicopathological features. In particular, loci at FGFR2, MAP3K1, and 2q35 were found to associate specifically with estrogen receptor-positive breast cancer.Citation92,Citation103,Citation104 Data are still limited for less common tumor subtypes, such as estrogen receptor-negative or basal-like breast tumors. Whether these loci are associated with the risk of breast cancer in males has not yet been investigated, but an involvement of low-penetrance alleles in male breast cancer susceptibility cannot be excluded and warrants ad hoc studies.
Acquired alterations in breast cancer
Breast cancer development and progression is a multistep process resulting from the accumulation of genetic alterations, such as mutations and copy number variations, and also epigenetic alterations, such as promoter methylation, resulting in aberrant gene expression. The increasing number of deregulated genes subsequently affects important cellular networks, such as cell cycle control, DNA repair, cell adhesion or migration, and differentiation, driving normal breast cells into highly malignant derivatives with metastatic potential. Such alterations can result either in inactivation of tumor suppressor genes (eg, TP53, BRCA1) or activation of proton-cogenes (eg, PIK3CA, MYC), both of which contribute to the malignant state of a transformed cell. Recent landmark studies have shed new light on the genomic landscape of breast cancer. Within a breast tumor there are many infrequently mutated genes and a few frequently mutated genes, resulting in incredible genetic heterogeneity.Citation105,Citation106 The great majority of somatic mutations frequently lie in hotspot regions that might represent targets in cancer therapy. Both genetic and epigenetic alterations are also frequently associated to specific biological and clinicopathological tumor characteristics, allowing the identification of personalized therapies targeting the associated molecular pathways.Citation107–Citation116
Genetic alterations in breast cancer
A number of gene and chromosome alterations have been identified in sporadic breast carcinomas. Indeed, the great majority of breast cancer cases are due to solely somatic genetic alterations without germ-line ones.Citation117,Citation118 A heterogeneous set of somatic alterations, including gene amplification, deletion, mutations, and rearrangements, were reported to be involved in the etiology of breast cancer.Citation6 The amount of information on these alterations has been dramatically increased by the introduction of high-throughput molecular cytogenetic approaches. Using large-scale approaches, the sequence of about 18,000 genes has been analyzed in breast cancer cases, and it has been reported that about 10% of these had at least one nonsilent mutation.Citation105 The great majority of alterations are single base substitutions (about 90%), with a prevalence of missense changes (60%, ). The remainder are somatic mutations resulting in stop codon or splice site alterations, and only a few of these are insertions, deletions, or duplicationsCitation105 (). Somatic mutations found in cancers can be subdivided mainly into two biological classes, ie, “driver” and “passenger” mutations. Driver mutations confer proliferative advantage to tumor cells and are positively selected during cancer development. Passenger mutations are present in the tumor progenitor cells, are biologically neutral, and do not confer a growth advantage.Citation105
Figure 2 Somatic mutations in breast cancer. The great majority of somatic mutations are single base substitutions, mainly missense mutations; missense changes account for about 60% of somatic alterations; the remaining somatic mutations result in stop codon (5.1%) or in alterations of splice site (2.8%) and only a few percentages of these are insertions, deletions, or duplications (5.6%).
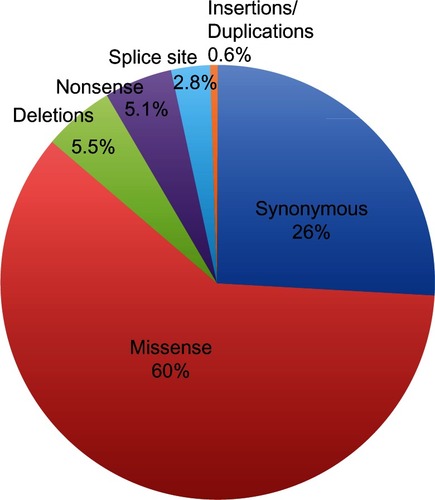
Several studies have shown a bimodal distribution of mutations in breast cancer. It has been proposed that the genomic landscape of breast cancer consists of “mountains” and “hills,” where the mountains correspond to the most frequently mutated genes, specifically PIK3CA and TP53, and the hills consist of a much larger number of less frequently mutated cancer-associated genes (<5%).Citation105,Citation106 Additional to mutations in PIK3CA and P53, alterations in several genes implicated in pathways involved in breast tumorigenesis, including the phosphatidylinositol 3-kinase/Akt and NF-kB pathways, have been also identified (eg, IKBKB, IRS4, NFKBIA, NFKBIE, NFKB1, PIK3R1, PIK3R4, RPS6KA3, MAP3K1, AKT1, and GATA3 genes).Citation105,Citation119,Citation120 These genes could be considered the hills of the mutational landscape of breast cancer, because their mutation frequencies are lower than for genes considered to be the mountains.
Mutations of the PIK3CA gene are observed in 16%–40% of female breast cancers and in about 18% of male breast cancers.Citation109,Citation121–Citation124PIK3CA mutations are associated with a positive estrogen receptor and progesterone receptor status, nodal involvement, and high histological grade, suggesting that they could be strong prognostic factors in breast cancer.Citation107–Citation110 The great majority (85%) of PIK3CA mutations are in exons 9 and 20, encoding the helical and kinase domains, respectively. The majority of mutations are located in two hot spot regions, including the central helical domain and the COOH terminal kinase domain. The three most common hot spot mutations lead to amino acid changes in the helical domain (E542K and E545K) and in the kinase domain (H1047R).Citation125 In particular E542K, E545K, and H1047R represent 3.6%, 6.2%, and 14.8% of the total PIK3CA mutations in breast cancer, respectively.Citation125
Mutations of the TP53 gene in breast cancer range from 15% to 71% among different populations.Citation126 More than 90% of TP53 mutations reported in breast cancer are located in conservative regions within exons 5 to 8.Citation126,Citation127 More than 2% of all TP53 mutations are represented by three TP53 hot spot mutations, ie, 273 (CGT > CAT), 158 (CGC > CTC), and 248 (CGG > CAG).Citation128 However, an overrepresentation of codon 163 (TAC > TGC) mutation has been observed in breast cancer. Indeed, this codon, rarely mutated in most cancers, accounts for over 2% of all breast cancer mutations. Interestingly, codon 163 is a hot spot for TP53 mutation in breast cancer among BRCA1/2 carriers.Citation127 Overall, TP53 mutations are strongly associated with high histological grade, negative estrogen receptor status, increased global genomic instability, and germ-line BRCA1 mutations.Citation111–Citation113 At least 14 different common polymorphisms have been described in the TP53 gene (IARC p53 data base www.p53.iarc.fr/p53main.html). The most common TP53 polymorphic variants are the16 bp duplication in intron 3 (TP53PIN3) and the TP53 G215C (Arg72Pro). There is some evidence of an association between TP53PIN3 and Arg72Pro variants and elevated breast cancer risk, although some studies suggest a neutral or protective effect for these polymorphisms.Citation129–Citation131 Genotype and haplotype analyses of these two TP53 polymorphisms also revealed that the presence of a specific haplotype carrying the consensus sequence for TP53PIN3 (allele without the 16 bp insertion), and the variant allele for Arg72Pro (72Pro) is associated with an earlier age at onset of breast cancer in BRCA2 mutation carriers.Citation132,Citation133
In addition to nonsynonymous mutations arising from single nucleotide substitutions, several splice variants specific to breast cancer have been reported.Citation134,Citation135 Interestingly, breast cancer-specific alternative splicing is not restricted to splicing defects resulting in loss of protein functions, and may also include modifications that generate proteins with new functions.Citation134
Alternative splicing events can involve breast cancer-specific genes, such as BRCA1, ESR2, and HER2, or genes involved in cell cycle progression, DNA damage response, and spliceosome assembly.Citation134,Citation136–Citation138 The number of known BRCA1 mRNA variants representing aberrant splicing products is relatively high.Citation136 The four predominant mRNA variants with a molecular weight lower than full length BRCA1 are ∆(9,10), ∆(9,10,11q), ∆(11q), and ∆(11). The variants that would be expected to differ the greatest at the functional level from the full length species are those lacking the largest exon 11, containing many functional domains involved in protein-protein interaction.Citation136
Different ESR2 (estrogen receptor β) splice variants have been identified and studies on the function of some of these suggested that they might act as a dominant negative receptor in the estrogen receptor α and β pathways.Citation135,Citation139
A specific splicing variant of HER2 (∆HER2), which causes lack of exon 16 encoding the extracellular domain, has been identified in 9% of breast cancers overexpressing HER2 protein, suggesting that HER2 proteins carrying splicing variants may represent the oncogenic receptor population.Citation137,Citation140
Different alternative splicing events have been identified in which changes in splicing correlate with estrogen receptor status and histological tumor grade.Citation134 Thus, analysis of alternative splicing might provide information about the biology of the tumor.
Genomic instability, such as gene copy number alterations and DNA amplifications, has also been observed frequently in breast cancer. The most commonly amplified regions in breast cancer include 8q24, 11q13, 12q14, 17q11, 17q24, and 20q13, with amplification of genes such as HER2 (15%–20%), EGFR (14%), MYC (15%–20%), CCND1 (15%–20%), ESR1 (20%), and EMSY (7%–13%).Citation114,Citation141,Citation142 Amplification of these regions increases genetic instability in breast cancer and is generally associated with poor prognosis.Citation114
HER2 and EGFR are members of the epidermal growth factor receptor family, and both genes are targets for copy number amplification in breast cancer.Citation143 Amplification of the HER2 gene causes HER2 protein levels that are 10–100 times greater than normal. HER2-positive breast cancers are associated with a worse prognosis and resistance to hormonal therapyCitation144,Citation145EGFR upregulation in breast cancer is not only due to gene amplification but often results from either high polysomy of chromosome 7 or transcriptional induction by the transcription factor YBX1 (Y box binding protein 1).Citation146EGFR amplification is frequently associated with poor prognosis parameters in breast cancer patients, such as large tumor size, high histological grade, high proliferative index, and negative estrogen receptor status.Citation147,Citation148 Moreover, increased EGFR gene copy numbers are observed in triple-negative (estrogen receptor, progesterone receptor, and HER2 negative) breast cancers together with decreased BRCA1 mRNA expression.Citation149
MYC functions as a transcription factor, regulating up to 15% of all human genes. Although the relationship between amplification and overexpression is not clearly delineated, MYC amplification is significantly correlated with aggressive tumor phenotypes and poor clinical outcomes. MYC amplification is emerging as an important predictor of response to HER2-targeted therapies, and its role in BRCA1-associated breast cancers makes it an important target in basal-like/triple-negative breast cancers.Citation150
Other two genes frequently amplified in breast cancer are ESR1 and CCND1. Amplification of ESR1 is associated with the expression of the estrogen receptor in breast cancer.Citation151 Overall, higher ESR1 gene amplification is found in tumors with CCND1 gene amplification in comparison with tumors without CCND1 gene amplification.Citation151CCND1 amplification occurred preferentially in estrogen receptor-positive breast cancer and is associated with reduced overall survival.Citation152–Citation155 Tumors which are sufficiently genetically unstable to develop one gene amplification have increased probability of developing multiple gene amplifications. The coamplification of one or several oncogenes, such as EGFR, ErbB2, CCND1, or ESR1, occurs commonly in breast cancer, and is reported in up to 30% of CCND1-amplified and up to 40% of ErbB2-amplified tumors.Citation114
EMSY amplification in sporadic breast tumors has been shown to be associated with a poor prognosis.Citation141 Amplification of EMSY has been reported in sporadic breast cancer but not in BRCA2-associated breast cancer, suggesting that BRCA2 mutations and EMSY gene amplification may be mutually exclusive.Citation156 Indeed, EMSY protein interacts with the transactivation domain of BRCA2, reducing its activity, and it has been suggested that amplification of EMSY can explain somatic BRCA2 inactivation.Citation156
Recently it has been reported that male breast cancers have a lower frequency of gene copy number alterations than female breast cancers.Citation157 Moreover, different chromosomal regions were found to be altered in male and female breast cancer. Male breast cancer alterations targeted Xp11.23 and 14q13.1 regions in more than 50% of cases. Shared amplified regions between male and female breast cancers are 8q24 (53%), 11q13 (50%), and 17q24 (30%), mapping for MYC, EMSY, and HER2, respectively.Citation117,Citation157 Furthermore, HER2 and CCND1 gene amplification is observed in about 8% and 12% of male breast cancer cases, respectively.Citation158,Citation159
Epigenetic alterations
Epigenetic changes, in particular DNA methylation, are emerging as one of the most important events involved in breast cancer initiation and progression, and there is evidence that DNA methylation may serve as a link between genome and environment. Interestingly, factors that can be modulated by the environment, such as estrogens, elicit epigenetic changes (such as DNA methylation) and this could contribute to breast cancer risk. Furthermore, tumor-specific CpG island hypermethylation profiles are now emerging in breast cancer.Citation7 Tumor-related genes that become hypermethylated may play a significant role in breast cancer, including BRCA1 and hormone response genes, such as estrogen, progesterone, androgen, and prolactin receptors.Citation160 Epigenetic silencing is one of the mechanisms by which mammary epithelial cells repress estrogen receptor expression, leading to the estrogen receptor-negative molecular subtypes of breast cancer.Citation161ESR1 methylation is more common in estrogen receptor-negative than in estrogen receptor-positive tumors, but there is no clear link between ESR1 methylation and estrogen receptor status. It has been suggested that a heterogeneous ESR1 gene methylation pattern may evolve during breast cancer progression and play a role in estrogen receptor-negative recurrences or metastases in patients with estrogen receptor-positive tumors.Citation162 Interestingly, breast tumors with BRCA1 methylation show a high frequency of ESR1 promoter methylation. BRCA1 somatic mutations are extremely rare in sporadic breast cancer, but 9%–13% of these tumors reveal aberrant BRCA1 methylation, especially when loss of heterozygosity occurs at the BRCA1 locus.Citation160BRCA1-associated breast cancers are generally basal-like tumors, and promoter methylation is one mechanism of BRCA1 gene silencing in sporadic basal-like breast cancers. However, there is no significant difference in BRCA1 methylation between sporadic basal-like breast cancers (14%) and matched sporadic nonbasal-like breast cancers (11%). BRCA1 methylation also appears to be similar across distinct breast cancer molecular subtypes (14%–17%) including ductal, mucinous, and lobular breast cancers.Citation163,Citation164
Tumor-specific CpG island hypermethylation profiles are emerging, and the growing list of genes inactivated by promoter hypermethylation in breast cancer include genes involved in evasion of apoptosis (RASSF1A, HOXA5, TWIST1), in cell cycle control (CCND2, p16, RARβ), and tissue invasion and metastasis (CDH1). Tumor suppressor genes, such as GSTP1, RIL, HIN-1, CDH13, APC, and RUNX3, are frequently methylated in breast cancer tissues.Citation115,Citation161,Citation165 These genes are not only hypermethylated in tumor cells, but show increased epigenetic silencing in normal epithelium surrounding the tumor site. Thus, methylation frequently represents an early event in breast cancer tumorigenesis. For example, CCND2, an important regulator of the cell cycle, has been frequently found to be methylated in breast cancer and is also methylated in ductal carcinoma in situ, suggesting that it may represent an early event in tumorigenesis.Citation161 Another gene frequently hypermethylated in breast cancer is RASSF1A. RASSF1A methylation is also an early epigenetic event in breast cancer and is found in ductal carcinoma in situ and in lobular carcinoma in situ.Citation160 Its diverse functions include regulation of apoptosis, growth regulation, and microtubule dynamics during mitotic progression.Citation161
There is some evidence that DNA hypermethylation patterns can identify breast cancer subgroups having distinctive biological properties that could be used for prognostication and for prediction of response to therapy.Citation115,Citation166,Citation167 An association between methylation in five genes, including RARb, CDH1, CCND2, p16, and ESR1, and poor histological differentiation of breast cancer is frequently reported.Citation115,Citation161 Furthermore, distinct epigenetic profiles can be identified when dividing breast tumors into groups based on hormone receptor status.Citation166,Citation168 Differences in methylation status of the promoter region CpG islands of major breast cancer tumor-related genes, such as RASSF1, CCND2, GSTP1, TWIST, RARb, and CDH1, have been found relating to estrogen receptor and HER2 status.Citation115 In particular, methylation of these tumor-related genes resulted in significantly higher estrogen receptor-positive and HER2-positive breast tumors.Citation115,Citation161 On the other hand, double-negative (estrogen receptor-negative, HER2-negative) breast cancers have significantly lower frequencies of RASSF1, GSTP, and APC methylation. Interestingly, epigenetic differences between estrogen receptor-positive and estrogen receptor-negative breast cancer arise early in cancer development and persist during cancer progression.Citation115
Micro RNA
miRNAs are small noncoding, double-stranded RNA molecules involved in post-transcriptional regulation of target genes. Aberrant expressions of miRNA are associated with cancer progression, by acting either as tumor suppressor genes or oncogenes.Citation8,Citation169 Much attention has been paid to deregulation of gene expression through the action of specific miRNA in breast cancer. Microarray studies demonstrated that overall miRNA expression could clearly separate normal versus cancerous breast tissue, with the most significantly deregulated miRNAs being mir-125b, mir-145, mir-21, mir-155, and mir–335.Citation170–Citation172 Interestingly, a large number of miRNAs, overexpressed or underexpressed in breast cancer, are predicted to regulate expression of key breast cancer proteins, such as BRCA1/2, ATM, PTEN, CHEK2, MLH1, P53, and ER.Citation169 Moreover, specific miRNAs, including miRNAs that regulate genes involved in cell proliferation, such as MAPK, RAS, HER2, HER3, and ESR1, have been shown to play a direct role in male breast cancer development.Citation173,Citation174 Indeed, cluster analysis of miRNA expression profiles reveals cancer-specific alterations of miRNA expression in male breast cancer distinct from female breast cancer, such as downregulation of miRNAs that suppress HOXD10, a protein involved in cell proliferation and migration, and vascular endothelial growth factor.Citation173
Significant differences in miRNA expression profiles associated with molecular subtypes of breast cancer and correlated with specific clinicopathological factors, such as estrogen receptor, progesterone receptor, and HER2 status, emerged in breast cancer.Citation170 Thus, evaluation of the associations between miRNA expression profiles and clinicopathological characteristics may be important to identify distinct breast cancer subgroups and may lead to improvements in the clinical management of breast cancer patients. Indeed, some miRNAs, including mir-21 and mir-145, have been shown to have potential clinical applications as novel biomarkers in the diagnosis and prognosis of breast cancer.Citation175,Citation176 Moreover, miRNAs may act as strong inhibitors of cellular pathways via regulation of entire sets of genes, thus suggesting a possibly great potential for miRNAs in breast cancer prevention and therapeutics.Citation116
Future research directions
Recent advances in technology have shed more light on the complexity of breast cancer biology and have provided data that allow risk estimation for patients with inherited mutations, prognostic and predictive determinations for patients with sporadic breast cancer, and targets for therapies. Recently, loci identified by genome-wide association studies have greatly expanded the list of genes associated with breast cancer risk.
However, evaluation of the functional consequences of low-penetrance alleles in breast cancer risk and their association with breast cancer molecular subtypes and clinicopathological characteristics are still challenging, but are needed for clinical application. Moreover, exploration of the polygenic model proposed for low-penetrance alleles requires further research in diverse and large populations. Overall, despite the remarkable efforts made in recent years, much of the complex landscape of familial breast cancer risk remains unknown, suggesting the need for ongoing efforts in this field.Citation96
The introduction of high-throughput molecular approaches has greatly increased the amount of information on the genomic landscape of breast tumors. Serial analysis of the cancer genome in different phases of its evolution might lead to improved management of the individual breast cancer patient.Citation118
Moreover, studying the global methylation status as well as miRNA expression profiles of different types of tumors will allow the development of profiles unique for breast cancer and its subtypes, staging, and prognostic categories, leading to diagnostic applications and identification of new therapeutic targets.Citation116,Citation171
Conclusion
The identification of breast cancer susceptibility genes, in particular BRCA1 and BRCA2, has changed the management of breast cancer patients with a family history of breast cancer. Several models have been developed, and are currently used to assess the pretest probability of identifying BRCA1/2 germ-line mutations in individuals at risk for hereditary breast and ovarian cancer. Moreover, novel therapeutic strategies specific for BRCA1 and BRCA2 cancers are emerging, including crosslinking agents and poly ADP ribose polymerase inhibitors.Citation156
Both genetic and epigenetic acquired alterations are frequently associated with specific biological and clinicopathological tumor characteristics, allowing identification of personalized therapies targeting specific molecular pathways. In particular, a number of compounds, including trastuzumab, lapatinib, and pertuzumab, are currently under clinical evaluation for HER2-targeted therapy.Citation177 However, the majority of HER2-overexpressing breast cancers do not respond to HER2-targeted therapy alone.Citation178 There is evidence showing that combination therapy involving the use of HER2 and endothelial growth factor receptor inhibitors, such as trastuzumab and lapatinib, may have promising results in breast cancer treatment.Citation178 Crosstalk between the endothelial growth factor receptor/HER2 and phosphatidylinositol 3-kinase/Akt pathways provides a rationale for combining anti-endothelial growth factor receptor/HER2 agents and inhibitors of phosphatidylinositol 3-kinase/Akt/mTOR in breast cancer. In addition, DNA methylation as well as miRNAs are currently emerging as interesting candidates for the development of therapeutic strategies against breast cancer. In conclusion, elucidation of the inherited and acquired genetic and epigenetic alterations involved in breast cancer has not only clarified the molecular pathways involved in development and progression of breast cancer itself, but may also have important clinical and therapeutic implications in the management of patients with breast cancer.
Disclosure
The authors report no conflicts of interest in this work.
References
- WillemsPGSusceptibility genes in breast cancer: more is less?Clin Genet200772493496
- AndersonWFJatoiITseJRosenbergPSMale breast cancer: a population-based comparison with female breast cancerJ Clin Oncol20102823223919996029
- MartinAMWeberBLGenetic and hormonal risk factors in breast cancerJ Natl Cancer Inst2000921126113510904085
- ThompsonDEastonDThe genetic epidemiology of breast cancer genesJ Mammary Gland Biol Neoplasia2004922123615557796
- LeeEYMullerWJOncogenes and tumor suppressor genesCold Spring Harb Perspect Biol20102a00323620719876
- StephensPJMcBrideDJLinMLComplex landscapes of somatic rearrangement in human breast cancer genomesNature20094621005101020033038
- EstellerMCancer epigenomics: DNA methylomes and histone-modification mapsNat Rev Genet2007828629817339880
- CroceCMCauses and consequences of microRNA dysregulation in cancerNat Rev Genet20091070471419763153
- HirshfieldKMRebbeckTRLevineAJGermline mutations and polymorphisms in the origins of cancers in womenJ Oncol2010 Epub 2010 Jan 10
- BorresenALAndersenTIGarberJScreening for germ line TP53 mutations in breast cancer patientsCancer Res199252323432361591732
- ChenJLindblomPLindblomAA study of the PTEN/MMAC1 gene in 136 breast cancer familiesHum Genet19981021241259490290
- WoosterRWeberBLBreast and ovarian cancerN Engl J Med20033482339234712788999
- StrattonMRRahmanNThe emerging landscape of breast cancer susceptibilityNat Genet200840172218163131
- NathansonKLWoosterRWeberBLBreast cancer genetics: what we know and what we needNat Med2001755255611329055
- MartinAMBlackwoodMAAntin-OzerkisDGermline mutations in BRCA1 and BRCA2 in breast-ovarian families from a breast cancer risk evaluation clinicJ Clin Oncol2001192247225311304778
- LiebensFPCarlyBPastijnARozenbergSManagement of BRCA1/2 associated breast cancer: a systematic qualitative review of the state of knowledge in 2006Eur J Cancer20074323825717095205
- FordDEastonDFStrattonMGenetic heterogeneity and penetrance analysis of the BRCA1 and BRCA2 genes in breast cancer families. The Breast Cancer Linkage ConsortiumAm J Hum Genet1998626766899497246
- AntoniouAPharoahPDNarodSAverage risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: a combined analysis of 22 studiesAm J Hum Genet2003721117113012677558
- ChenSIversenESFriebelTCharacterization of BRCA1 and BRCA2 mutations in a large United States sampleJ Clin Oncol20062486387116484695
- FrankTSDeffenbaughAMReidJEClinical characteristics of individuals with germline mutations in BRCA1 and BRCA2: analysis of 10,000 individualsJ Clin Oncol2002201480149011896095
- OttiniLRizzoloPZannaIBRCA1/BRCA2 mutation status and clinical-pathologic features of 108 male breast cancer cases from Tuscany: a population-based study in central ItalyBreast Cancer Res Treat200911657758618819001
- PapiLPutignanoALCongregatiCFounder mutations account for the majority of BRCA1-attributable hereditary breast/ovarian cancer cases in a population from Tuscany, Central ItalyBreast Cancer Res Treat200911749750418821011
- DurocherFShattuck-EidensDMcClureMComparison of BRCA1 polymorphisms, rare sequence variants and/or missense mutations in unaffected and breast/ovarian cancer populationsHum Mol Genet199658358428776600
- DunningAMChianoMSmithNRCommon BRCA1 variants and susceptibility to breast and ovarian cancer in the general populationHum Mol Genet199762852899063749
- QiuLXYaoLXueKBRCA2 N372H polymorphism and breast cancer susceptibility: a meta-analysis involving 44,903 subjectsBreast Cancer Res Treat201012348749020135345
- PalliDFalchettiMMasalaGAssociation between the BRCA2 N372H variant and male breast cancer risk: a population-based case-control study in Tuscany, Central ItalyBMC Cancer2007717017767707
- ThompsonDEastonDBreast Cancer Linkage Consortium. Variation in cancer risks, by mutation position, in BRCA2 mutation carriersAm J Hum Genet20016841041911170890
- ThompsonDEastonDBreast Cancer Linkage Consortium. Variation in BRCA1 cancer risks by mutation positionCancer Epidemiol Biomarkers Prev20021132933611927492
- PhelanCMKwanEJackEA low frequency of non-founder BRCA1 mutations in Ashkenazi Jewish breast-ovarian cancer familiesHum Mutat20022035235712402332
- StruewingJPAbeliovichDPeretzTThe carrier frequency of the BRCA1 185delAG mutation is approximately 1 percent in Ashkenazi Jewish individualsNat Genet1995111982007550349
- GudmundssonJJohannesdottirGArasonAFrequent occurrence of BRCA2 linkage in Icelandic breast cancer families and segregation of a common BRCA2 haplotypeAm J Hum Genet1996587497568644738
- ThorlaciusSOlafsdottirGTryggvadottirLA single BRCA2 mutation in male and female breast cancer families from Iceland with varied cancer phenotypesNat Genet1996131171198673089
- JohannesdottirGGudmundssonJBergthorssonJTHigh prevalence of the 999del5 mutation in Icelandic breast and ovarian cancer patientsCancer Res199656366336658706004
- BaudiFQuaresimaBGrandinettiCEvidence of a founder mutation of BRCA1 in a highly homogeneous population from southern Italy with breast/ovarian cancerHum Mutat20011816316411462242
- RussoACalòVBrunoLIs BRCA1-5083del19, identified in breast cancer patients of Sicilian origin, a Calabrian founder mutation? Breast Cancer Res Treat2009113677018228134
- CaligoMAGhimentiCCipolliniGBRCA1 germline mutational spectrum in Italian families from Tuscany: a high frequency of novel mutationsOncogene199613148314888875986
- SluiterMDvan RensburgEJLarge genomic rearrangements of the BRCA1 and BRCA2 genes: review of the literature and report of a novel BRCA1 mutationBreast Cancer Res Treat201112532534920232141
- HansenTOJønsonLAlbrechtsenAAndersenMKEjlertsenBNielsenFCLarge BRCA1 and BRCA2 genomic rearrangements in Danish high risk breast-ovarian cancer familiesBreast Cancer Res Treat200911531532318546071
- KarhuRLaurilaEKallioniemiASyrjäkoskiKLarge genomic BRCA2 rearrangements and male breast cancerCancer Detect Prev20063053053417113724
- WoodwardAMDavisTASilvaAGKirkJALearyJAkConFab InvestigatorsLarge genomic rearrangements of both BRCA2 and BRCA1 are a feature of the inherited breast/ovarian cancer phenotype in selected familiesJ Med Genet200542e3115863663
- WalshTCasadeiSCoatsKHSpectrum of mutations in BRCA1, BRCA2, CHEK2, and TP53 in families at high risk of breast cancerJAMA20062951379138816551709
- WangTLererIGuetaZA deletion/insertion mutation in the BRCA2 gene in a breast cancer family: a possible role of the Alu-polyA tail in the evolution of the deletionGenes Chromosomes Cancer200131919511284040
- TournierIPailleretsBBSobolHSignificant contribution of germline BRCA2 rearrangements in male breast cancer familiesCancer Res2004648143814715548676
- Gutiérrez-EnríquezSde la HoyaMMartínez-BouzasCScreening for large rearrangements of the BRCA2 gene in Spanish families with breast/ovarian cancerBreast Cancer Res Treat200710310310717063271
- Meijers-HeijboerHvan den OuwelandAKlijnJLow-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutationsNat Genet20023555911967536
- WeischerMBojesenSEEllervikCTybjaerg-HansenANordestgaardBGCHEK2*1100delC genotyping for clinical assessment of breast cancer risk: meta-analyses of 26,000 patient cases and 27,000 controlsJ Clin Oncol20082654254818172190
- NeuhausenSDunningASteeleLRole of CHEK2*1100delC in unselected series of non-BRCA1/2 male breast cancersInt J Cancer200410847747814648718
- OhayonTGalIBaruchRGSzaboCFriedmanECHEK2*1100delC and male breast cancer risk in IsraelInt J Cancer200410847948014648719
- SyrjäkoskiKKuukasjärviTAuvinenAKallioniemiOPCHEK2 1100delC is not a risk factor for male breast cancer populationInt J Cancer200410847547614648717
- FalchettiMLupiRRizzoloPBRCA1/BRCA2 rearrangements and CHEK2 common mutations are infrequent in Italian male breast cancer casesBreast Cancer Res Treat200811016116717661168
- Martinez-BouzasCBeristainEGuerraICHEK2 1100delC is present in familial breast cancer cases of the Basque CountryBreast Cancer Res Treat200710311111317063278
- NarodSALynchHTCHEK2 mutation and hereditary breast cancerJ Clin Oncol2007256717132696
- CaligoMAAgataSAcetoGThe CHEK2 c.1100delC mutation plays an irrelevant role in breast cancer predisposition in ItalyHum Mutat20042410010115221794
- SchutteMSealSBarfootRBreast Cancer Linkage Consortium. Variants in CHEK2 other than 1100delC do not make a major contribution to breast cancer susceptibilityAm J Hum Genet2003721023102812610780
- BogdanovaNEnssen-DubrowinskajaNFeshchenkoSAssociation of two mutations in the CHEK2 gene with breast cancerInt J Cancer200511626326615810020
- SwiftMReitnauerPJMorrellDChaseCLBreast and other cancers in families with ataxia-telangiectasiaN Engl J Med1987316128912943574400
- ThompsonDDuedalSKirnerJCancer risks and mortality in heterozygous ATM mutation carriersJ Natl Cancer Inst20059781382215928302
- RenwickAThompsonDSealSATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility allelesNat Genet20063887387516832357
- Levy-LahadEFanconi anemia and breast cancer susceptibility meet againNat Genet20104236836920428093
- RahmanNSealSThompsonDPALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility geneNat Genet20073916516717200668
- TischkowitzMXiaBSabbaghianNAnalysis of PALB2/FANCN-associated breast cancer familiesProc Natl Acad Sci USA20071046788679317420451
- CaoAYHuangJHuZThe prevalence of PALB2 germline mutations in BRCA1/BRCA2 negative Chinese women with early onset breast cancer or affected relativesBreast Cancer Res Treat200911445746218446436
- HeikkinenTKärkkäinenHAaltonenKThe breast cancer susceptibility mutation PALB2 1592delT is associated with an aggressive tumor phenotypeClin Cancer Res2009153214322219383810
- PapiLPutignanoALCongregatiCA PALB2 germline mutation associated with hereditary breast cancer in ItalyFam Cancer2010918118519763884
- Dansonka-MieszkowskaAKluskaAMoesJA novel germline PALB2 deletion in Polish breast and ovarian cancer patientsBMC Med Genet2010112020122277
- SluiterMMewSvan RensburgEJPALB2 sequence variants in young South African breast cancer patientsFam Cancer2009834735319333784
- GarcíaMJFernándezVOsorioAAnalysis of FANCB and FANCN/PALB2 Fanconi anemia genes in BRCA1/2-negative Spanish breast cancer familiesBreast Cancer Res Treat200911354555118302019
- BaliaCSensiELombardiGRoncellaMBevilacquaGCaligoMAPALB2: a novel inactivating mutation in a Italian breast cancer familyFam Cancer2010953153620852946
- DingYCSteeleLChuLHGermline mutations in PALB2 in African-American breast cancer casesBreast Cancer Res Treat201112622723021113654
- ErkkoHXiaBNikkiläJA recurrent mutation in PALB2 in Finnish cancer familiesNature200744631631917287723
- FoulkesWDGhadirianPAkbariMRIdentification of a novel truncating PALB2 mutation and analysis of its contribution to early-onset breast cancer in French-Canadian womenBreast Cancer Res20079R8318053174
- Sauty de ChalonATeoZParkDJAre PALB2 mutations associated with increased risk of male breast cancer?Breast Cancer Res Treat201012125325520091115
- SilvestriVRizzoloPZannaIPALB2 mutations in male breast cancer: a population-based study in Central ItalyBreast Cancer Res Treat201012229930120180015
- AdankMAvan MilSEGilleJJWaisfiszQMeijers-HeijboerHPALB2 analysis in BRCA2-like familiesBreast Cancer Res Treat201112735736220582465
- DingYCSteeleLKuanCJGreilacSNeuhausenSLMutations in BRCA2 and PALB2 in male breast cancer cases from the United StatesBreast Cancer Res Treat201112677177820927582
- CasadeiSNorquistBMWalshTContribution to familial breast cancer of inherited mutations in the BRCA2-interacting protein PALB2Cancer Res2011712222222921285249
- SealSThompsonDRenwickATruncating mutations in the Fanconi anemia J gene BRIP1 are low-penetrance breast cancer susceptibility allelesNat Genet2006381239124117033622
- KarppinenSMVuoskuJHeikkinenKAllinenMWinqvistRNo evidence of involvement of germline BACH1 mutations in Finnish breast and ovarian cancer familiesEur J Cancer20033936637112565990
- GuénardFLabrieYOuelletteGMutational analysis of the breast cancer susceptibility gene BRIP1/BACH1/FANCJ in high-risk non-BRCA1/BRCA2 breast cancer familiesJ Hum Genet20085357959118414782
- CaoAYHuangJHuZMutation analysis of BRIP1/BACH1 in BRCA1/BRCA2 negative Chinese women with early onset breast cancer or affected relativesBreast Cancer Res Treat2009115515518483852
- LewisAGFlanaganJMarshAMutation analysis of FANCD2, BRIP1/BACH1, LMO4 and SFN in familial breast cancerBreast Cancer Res20057R1005R101616280053
- De NicoloATancrediMLombardiGA novel breast cancer-associated BRIP1 (FANCJ/BACH1) germ-line mutation impairs protein stability and functionClin Cancer Res2008144672468018628483
- SilvestriVRizzoloPFalchettiMMutation analysis of BRIP1 in male breast cancer cases: a population-based study in Central ItalyBreast Cancer Res Treat201112653954321165771
- MeindlAHellebrandHWiekCGermline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility geneNat Genet20104241041420400964
- AkbariMRToninPFoulkesWDGhadirianPTischkowitzMNarodSARAD51C germline mutations in breast and ovarian cancer patientsBreast Cancer Res20101240420723205
- ZhengYZhangJHopeKNiuQHuoDOlopadeOIScreening RAD51C nucleotide alterations in patients with a family history of breast and ovarian cancerBreast Cancer Res Treat201012485786120697805
- SilvestriVRizzoloPFalchettiMMutation screening of RAD51C in male breast cancer patientsBreast Cancer Res20111340421392410
- PharoahPDAntoniouABobrowMZimmernRLEastonDFPonderBAPolygenic susceptibility to breast cancer and implications for preventionNat Genet200231333611984562
- Breast Cancer Association ConsortiumCommonly studied single-nucleotide polymorphisms and breast cancer: results from the Breast Cancer Association ConsortiumJ Natl Cancer Inst2006981382139617018785
- EastonDFPooleyKADunningAMGenome-wide association study identifies novel breast cancer susceptibility lociNature20074471087109317529967
- CoxADunningAMGarcia-ClosasMA common coding variant in CASP8 is associated with breast cancer riskNat Genet20073935235817293864
- StaceySNManolescuASulemPCommon variants on chromosome 5p12 confer susceptibility to estrogen receptor-positive breast cancerNat Genet20084070370618438407
- ThomasGJacobsKBKraftPA multistage genome-wide association study in breast cancer identifies two new risk alleles at 1p11.2 and 14q24.1 (RAD51L1)Nat Genet20094157958419330030
- DunningAMHealeyCSBaynesCAssociation of ESR1 gene tagging SNPs with breast cancer riskHum Mol Genet2009181131113919126777
- TurnbullCAhmedSMorrisonJGenome-wide association study identifies five new breast cancer susceptibility lociNat Genet20104250450720453838
- FletcherOHoulstonRSArchitecture of inherited susceptibility to common cancerNat Rev Cancer20101035336120414203
- AntoniouACSpurdleABSinilnikovaOMCommon breast cancer-predisposition alleles are associated with breast cancer risk in BRCA1 and BRCA2 mutation carriersAm J Hum Genet20088293794818355772
- WassermanNFAneasINobregaMAAn 8q24 gene desert variant associated with prostate cancer risk confers differential in vivo activity to a MYC enhancerGenome Res2010201191119720627891
- GhoussainiMSongHKoesslerTMultiple loci with different cancer specificities within the 8q24 gene desertJ Natl Cancer Inst200810096296618577746
- WokołorczykDLubińskiJNarodSACybulskiCGenetic heterogeneity of 8q24 region in susceptibility to cancerJ Natl Cancer Inst200910127827919211453
- WokolorczykDGliniewiczBSikorskiAA range of cancers is associated with the rs6983267 marker on chromosome 8Cancer Res2008689982998619047180
- BonifaciNGórskiBMasojćBExploring the link between germline and somatic genetic alterations in breast carcinogenesisPLoS One20105e1407821124932
- StaceySNManolescuASulemPCommon variants on chromosomes 2q35 and 16q12 confer susceptibility to estrogen receptor-positive breast cancerNat Genet20073986586917529974
- Garcia-ClosasMChanockSGenetic susceptibility loci for breast cancer by estrogen receptor statusClin Cancer Res2008148000800919088016
- WoodLDParsonsDWJonesSThe genomic landscapes of human breast and colorectal cancersScience20073181108111317932254
- GreenmanCStephensPSmithRPatterns of somatic mutation in human cancer genomesNature200744615315817344846
- BachmanKEArganiPSamuelsYThe PIK3CA gene is mutated with high frequency in human breast cancersCancer Biol Ther2004377277515254419
- SaalLHHolmKMaurerMPIK3CA mutations correlate with hormone receptors, node metastasis, and ERBB2, and are mutually exclusive with PTEN loss in human breast carcinomaCancer Res2005652554255915805248
- LiSYRongMGrieuFIacopettaBPIK3CA mutations in breast cancer are associated with poor outcomeBreast Cancer Res Treat200696919516317585
- MaruyamaNMiyoshiYTaguchiTTamakiYMondenMNoguchiSClinicopathologic analysis of breast cancers with PIK3CA mutations in Japanese womenClin Cancer Res2007132 Pt 140841417202311
- JongYJLiLHTsouMHChromosomal comparative genomic hybridization abnormalities in early- and late-onset human breast cancers: correlation with disease progression and TP53 mutationsCancer Genet Cytogenet2004148556514697642
- LangerødAZhaoHBorganØTP53 mutation status and gene expression profiles are powerful prognostic markers of breast cancerBreast Cancer Res20079R3017504517
- HolstegeHJoosseSAvan OostromCTNederlofPMde VriesAJonkersJHigh incidence of protein-truncating TP53 mutations in BRCA1-related breast cancerCancer Res2009693625363319336573
- Al-KurayaKSchramlPTorhorstJPrognostic relevance of gene amplifications and coamplifications in breast cancerCancer Res2004648534854015574759
- SunamiEShinozakiMSimMSEstrogen receptor and HER2/neu status affect epigenetic differences of tumor-related genes in primary breast tumorsBreast Cancer Res200810R4618485221
- Nana-SinkamSPCroceCMMicroRNAs as therapeutic targets in cancerTransl Res201115721622521420032
- LereboursFLidereauRMolecular alterations in sporadic breast cancerCrit Rev Oncol Hematol20024412114112413631
- BellDWOur changing view of the genomic landscape of cancerJ Pathol201022023124319918804
- KanZJaiswalBSStinsonJDiverse somatic mutation patterns and pathway alterations in human cancersNature201046686987320668451
- ArnoldJMChoongDYThompsonERFrequent somatic mutations of GATA3 in non-BRCA1/BRCA2 familial breast tumors, but not in BRCA1-, BRCA2- or sporadic breast tumorsBreast Cancer Res Treat201011949149619189213
- CampbellIGRussellSEChoongDYMutation of the PIK3CA gene in ovarian and breast cancerCancer Res2004647678768115520168
- ButtittaFFelicioniLBarassiFPIK3CA mutation and histological type in breast carcinoma: high frequency of mutations in lobular carcinomaJ Pathol200620835035516353168
- DunlapJLeCShuklaAPhosphatidylinositol-3-kinase and AKT1 mutations occur early in breast carcinomaBreast Cancer Res Treat201012040941819418217
- BenvenutiSFrattiniMArenaSPIK3CA cancer mutations display gender and tissue specificity patternsHum Mutat20082928428818022911
- BaderAGKangSZhaoLVogtPKOncogenic PI3K deregulates transcription and translationNat Rev Cancer2005592192916341083
- Børresen-DaleALTP53 and breast cancerHum Mutat20032129230012619115
- OlivierMHainautPTP53 mutation patterns in breast cancers: searching for clues of environmental carcinogenesisSemin Cancer Biol20011135336011562177
- McKinziePBDelongchampRRHeflichRHParsonsBLProspects for applying genotypic selection of somatic oncomutation to chemical risk assessmentMutat Res2001489477811673089
- PietschECHumbeyOMurphyMEPolymorphisms in the p53 pathwayOncogene2006251602161116550160
- DaminAPFrazzonAPDaminDCEvidence for an association of TP53 codon 72 polymorphism with breast cancer riskCancer Detect Prev20063052352917113725
- KhadangBFattahiMJTaleiADehaghaniASGhaderiAPolymorphism of TP53 codon 72 showed no association with breast cancer in Iranian womenCancer Genet Cytogenet2007173384217284368
- OsorioAMartínez-DelgadoBPollánMA haplotype containing the p53 polymorphisms Ins16bp and Arg72Pro modifies cancer risk in BRCA2 mutation carriersHum Mutat20062724224816419081
- MartinAMKanetskyPAAmirimaniBGermline TP53 mutations in breast cancer families with multiple primary cancers: is TP53 a modifier of BRCA1? J Med Genet200340e3412676907
- VenablesJPKlinckRBramardAIdentification of alternative splicing markers for breast cancerCancer Res2008689525953119010929
- WatsonPMWatsonDKAlternative splicing in prostate and breast cancerThe Open Cancer Journal201036276
- OrbanTIOlahEEmerging roles of BRCA1 alternative splicingMol Pathol20035619119712890739
- CastiglioniFTagliabueECampiglioMPupaSMBalsariAMénardSRole of exon-16-deleted HER2 in breast carcinomasEndocr Relat Cancer20061322123216601290
- AndréFMichielsSDessenPExonic expression profiling of breast cancer and benign lesions: a retrospective analysisLancet Oncol20091038139019249242
- LaVoieHADeSimoneDCGillio-MeinaCHuiYYCloning and characterization of porcine ovarian estrogen receptor β isoformsBiol Reprod20026661662311870066
- KwongKYHungMA novel splice variant of HER2 with increased transformation activityMol Carcinog19982362689808159
- RodriguezCHughes-DaviesLVallèsHAmplification of the BRCA2 pathway gene EMSY in sporadic breast cancer is related to negative outcomeClin Cancer Res2004105785579115355907
- CiampaAXuBAyataGHER-2 status in breast cancer: correlation of gene amplification by FISH with immunohistochemistry expression using advanced cellular imaging systemAppl Immunohistochem Mol Morphol20061413213716785779
- Vanden BemptIDrijkoningenMDe Wolf-PeetersCThe complexity of genotypic alterations underlying HER2-positive breast cancer: an explanation for its clinical heterogeneityCurr Opin Oncol20071955255717906451
- RossJSFletcherJAThe HER-2/neu oncogene in breast cancer: prognostic factor, predictive factor, and target for therapyStem Cells1998164134289831867
- FrancisGBeadleGThomasSMengersenKSteinSEvaluation of oestrogen and progesterone receptor status in HER-2 positive breast carcinomas and correlation with outcomePathology20063839139817008275
- StratfordALFryCJDesiletsCY-box binding protein-1 serine 102 is a downstream target of p90 ribosomal S6 kinase in basal-like breast cancer cellsBreast Cancer Res200810R9919036157
- TsutsuiSOhnoSMurakamiSHachitandaYOdaSPrognostic value of epidermal growth factor receptor (EGFR) and its relationship to the estrogen receptor status in 1029 patients with breast cancerBreast Cancer Res Treat200271677511859875
- UberallIKolárZTrojanecRBerkovcováJHajdúchMThe status and role of ErbB receptors in human cancerExp Mol Pathol200884798918279851
- ToyamaTYamashitaHKondoNFrequently increased epidermal growth factor receptor (EGFR) copy numbers and decreased BRCA1 mRNA expression in Japanese triple-negative breast cancersBMC Cancer2008830918950515
- ChenYMYC in breast tumor progressionExpert Rev Anticancer Ther200881689169818925859
- HolstFStahlPRRuizCEstrogen receptor alpha (ESR1) gene amplification is frequent in breast cancerNat Genet20073965566017417639
- RoyPGThompsonAMCyclin D1 and breast cancerBreast20061571872716675218
- CourjalFLouasonGSpeiserPKatsarosDZeillingerRTheilletCCyclin gene amplification and overexpression in breast and ovarian cancers: evidence for the selection of cyclin D1 in breast and cyclin E in ovarian tumorsInt J Cancer1996692472538797862
- BärlundMMonniOKononenJMultiple genes at 17q23 undergo amplification and overexpression in breast cancerCancer Res2000605340534411034067
- KirkegaardTNielsenKVJensenLBGenetic alterations of CCND1 and EMSY in breast cancersHistopathology20085269870518393977
- TanDSMarchiòCReis-FilhoJSHereditary breast cancer: from molecular pathology to tailored therapiesJ Clin Pathol2008611073108218682420
- TommasiSMangiaAIannelliGGene copy number variation in male breast cancer by aCGHAnal Cell Pathol (Amst)20103311311921045282
- FonsecaRRTomásARAndréSSoaresJEvaluation of ERBB2 gene status and chromosome 17 anomalies in male breast cancerAm J Surg Pathol2006301292129817001161
- BärlundMKuukasjärviTSyrjäkoskiKAuvinenAKallioniemiAFrequent amplification and overexpression of CCND1 in male breast cancerInt J Cancer200411196897115300811
- DworkinAMHuangTHMEwart TolandAEpigenetic alterations in the breast: implications for breast cancer detection, prognosis and treatmentSemin Cancer Biol20091916517119429480
- JovanovicaJAnders RønnebergbJTostJKristensenaVThe epigenetics of breast cancerMol Oncol2010424225420627830
- LapidusRFergusonATOttavianoYLMethylation of estrogen and progesterone receptor gene 50 CpG islands correlates with lack of estrogen and progesterone receptor gene expression in breast tumorsClin Cancer Res199628058109816234
- BaeYKBrownAGarrettEHypermethylation in histologically distinct classes of breast cancerClin Cancer Res2004105998600515447983
- FacklerMJMcVeighMEvronEDNA methylation of RASSF1A, HIN-1, RAR-beta, Cyclin D2 and Twist in in situ and invasive lobular breast carcinomaCancer2003107970975
- LiSRongMIacopettaBDNA hypermethylation in breast cancer and its association with clinicopathological featuresCancer Lett200623727228016029926
- FengWShenLWenSCorrelation between CpG methylation profiles and hormone receptor status in breast cancersBreast Cancer Res20079R5717764565
- ParrellaPPoetaMLGalloAPNonrandom distribution of aberrant promoter methylation of cancer-related genes in sporadic breast tumorsClin Cancer Res2004105349535415328171
- WidschwendterMSiegmundKDMullerHMAssociation of breast cancer DNA methylation profiles with hormone receptor status and response to tamoxifenCancer Res2004643807381315172987
- ShenoudaSKAlahariSKMicroRNA function in cancer: oncogene or a tumor suppressor? Cancer Metastasis Rev20092836937820012925
- IorioMVFerracinMLiuCGMicroRNA gene expression deregulation in human breast cancerCancer Res2005657065707016103053
- ShiMGuoNMicroRNA expression and its implications for the diagnosis and therapeutic strategies of breast cancerCancer Treat Rev20093532833419171434
- PngKJYoshidaMZhangXHMicroRNA-335 inhibits tumor reinitiation and is silenced through genetic and epigenetic mechanisms in human breast cancerGenes Dev20112522623121289068
- FassanMBaffaRPalazzoJPMicroRNA expression profiling of male breast cancerBreast Cancer Res200911R5819664288
- LehmannUStreichertTOttoBIdentification of differentially expressed microRNAs in human male breast cancerBMC Cancer20101010920331864
- SempereLFChristensenMSilahtarogluAAltered microRNA expression confined to specific epithelial cell subpopulations in breast cancerCancer Res200767116121162018089790
- YanLXHuangXFShaoQMicroRNA miR-21 overexpression in human breast cancer is associated with advanced clinical stage, lymph node metastasis and patient poor prognosisRNA2008142348236018812439
- OttiniLCapalboCRizzoloPHER2-positive male breast cancer: an updateBreast Cancer: Targets and Therapy201024558
- SaxenaRDwivediAErbB family receptor inhibitors as therapeutic agents in breast cancer: current status and future clinical perspectiveMed Res RevOctober 252010 [Epub ahead of print]