Abstract
This paper discusses cyberinformation studies of the amino acid composition of insulin, in particular the identification of scientific terminology that could describe this phenomenon, ie, the study of genetic information, as well as the relationship between the genetic language of proteins and theoretical aspects of this system and cybernetics. The results of this research show that there is a matrix code for insulin. It also shows that the coding system within the amino acid language gives detailed information, not only on the amino acid “record”, but also on its structure, configuration, and various shapes. The issue of the existence of an insulin code and coding of the individual structural elements of this protein are discussed. Answers to the following questions are sought. Does the matrix mechanism for biosynthesis of this protein function within the law of the general theory of information systems, and what is the significance of this for understanding the genetic language of insulin? What is the essence of existence and functioning of this language? Is the genetic information characterized only by biochemical principles or it is also characterized by cyberinformation principles? The potential effects of physical and chemical, as well as cybernetic and information principles, on the biochemical basis of insulin are also investigated. This paper discusses new methods for developing genetic technologies, in particular more advanced digital technology based on programming, cybernetics, and informational laws and systems, and how this new technology could be useful in medicine, bioinformatics, genetics, biochemistry, and other natural sciences.
Introduction
The biologic role of any given protein in essential life processes, eg, insulin, depends on the positioning of its component amino acids, and is understood by the “positioning of letters forming words”. Each of these words has its own biochemical base. If this base is expressed by corresponding discrete numbers, it can be seen that any given base has its own program, along with its own unique cybernetic and information characteristics. Indeed, the sequencing of the molecule is determined not only by distint biochemical features, but also by cybernetic and information principles. For this reason, research in this field deals more with the quantitative rather than qualitative characteristics of genetic information and its biochemical basis. For the purposes of this paper, specific physical and chemical factors have been selected in order to express the genetic information for insulin. Numeric values are then assigned to these factors, enabling them to be measured. In this way it is possible to determine if a connection really exists between the quantitative ratios in the process of transfer of genetic information and the qualitative appearance of the insulin molecule. To select these factors, preference is given to classical physical and chemical parameters, including the numbers of atoms in the relevant amino acids, their analog values, the position of these amino acids in the peptide chain, and their frequencies. There is a large number of these parameters, and each of them gives important genetic information. Going through this process, it becomes clear that there is a mathematical relationship between quantitative ratios and the qualitative appearance of the biochemical “genetic processes”, and that there is a measurement method that can be used to describe the biochemistry of insulin.
Methods
Insulin can be represented by two different forms, ie, a discrete form and a sequential form. In the discrete form, a molecule of insulin is represented by a set of discrete codes or a multiple dimension vector. In the sequential form, an insulin molecule is represented by a series of amino acids according to the order of their position in the 1AI0 chains. Therefore, the sequential form can naturally reflect all the information about the sequence order and length of an insulin molecule. The key issue is whether we can develop a different discrete method of representing an insulin molecule that will allow accommodation of partial, if not all, sequence order information? Because a protein sequence is usually represented by a series of amino acid codes, what kind of numerical values should be assigned to these codes in order to optimally convert the sequence order information into a series of numbers for the discrete form representation? This section discusses how biochemical function of the molecule is determined by cybernetic information principles.
Expression of insulin code matrix 1AI0
The matrix mechanism of insulin, evolution of biomacromolecules and, especially, biochemical evolution of the insulin language have been analyzed by the application of cybernetic methods, information theory, and system theory, respectively. The primary structure of a molecule of insulin is an expression of the exact specification of its atomic composition and the chemical bonds between those atoms.
R6 insulin hexamer (d1ai02)
The structure 1AI0 has a total of 12 chains. Of these, two are sequence-unique identical chains of BFHJL ().
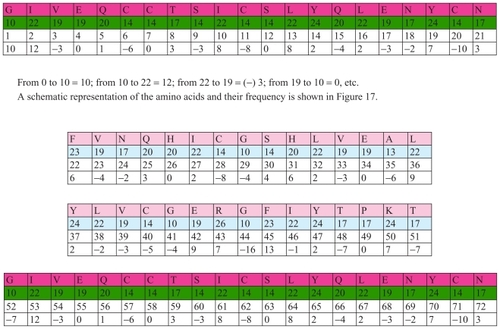
Figure 1 Number of atoms in insulin B chain.
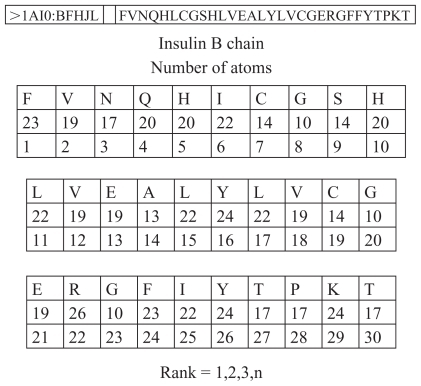
This positioning is of key importance for understanding the programmatic, cybernetic, and information elements of this protein. The scientific key for interpretation of biochemical processes is the same for insulin as for other proteins and sequences in biochemistry. The first amino acid in this example has 23 atoms, the second 19, the third 17, etc. They have exactly these numbers of atoms because there are many codes in the insulin molecules, analog codes, and other coded features. In fact, there is a cybernetic algorithm in which it is “recorded” that the first amino acid has to have 23 atoms, the second 19, the third 17, etc. The first amino acid has its own biochemistry, as does the second and the third, etc. The obvious conclusion is that there is a concrete relationship between quantitative ratios in the process of transfer of genetic information and qualitative appearance, ie, the characteristics of the organism.
Bio insulin code
The bio insulin code is an area of biomacromolecular processes in biochemistry (chemical engineering, bioprocess engineering, information technology, biorobotics) that treats signals as stochastic processes, dealing with their biosignal properties (eg, frequencies, mean, and covariance). In this context, biocodes are modeled as functions consisting of both deterministic and stochastic components. A simple example and also a common model of many bio systems is a signal Bio (t) that consists of a deterministic part x (t) as a white biocode ().
Figure 2 Fragment of insulin B chain.
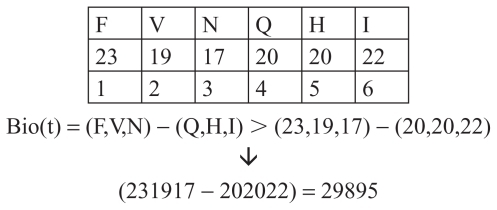
Bio (t) = [X1,2,3,n(t) – X1,2,3,n(t)] X (t) = Biocode 1,2,3,n
(A,B,C) – (B,C,D) = biocode 1
(B,C,D – C,D,E) = biocode 2
(D,E,F – E,F,G) = biocode 3 (1) etc.
where A, B, C, and n represent the connection of group amino acid positions, 1,2,3,n
Thus, the union of these amino acids generates the number 29895, which is a code representing one of the quantitative characteristics of the given genetic information. In a similar way we can calculate biocodes for other groups of amino acids, which are connected by various biocodes and analog codes, as well as other quantitative features. Connection is one of the numerical expressions that connect various corresponding features in biochemistry. It has a very prominent place in our mathematical understanding of all processes in biochemistry. This is a recently discovered phenomenon, the role and significance of which will hopefully be clarified in the future. Those bioprocesses are well correlated, and the autocorrelation function is a biocode:
Bio I = (X1,2,3,n(t) – X1,2,3,n(t)) = [(−)Y ↔ (+)Y]
[(−)Y = (+)Y] (2)
where Y represents the result as a functional biocode.
Insulin B chain
Mathematical evidence is provided here to prove that in the biochemistry of insulin there really is a programmatic and cybernetic algorithm in which it is “recorded”, in the language of mathematics, how the molecule will be built and what will be the quantitative characteristics of the given genetic information.
Bio(t) = X1,2,3,n(t) – X1,2,3,n(t)
Where:
X(t) = biocodes 1,2,3,n
Bio 1 = (A,B,C) – (B,C,D) = biocodes 1
Examples are presented in .
Figure 3 Examples of biocodes.
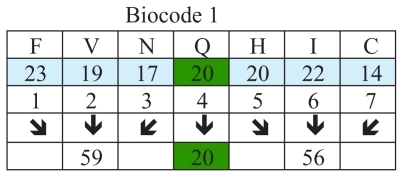
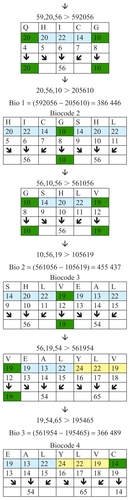
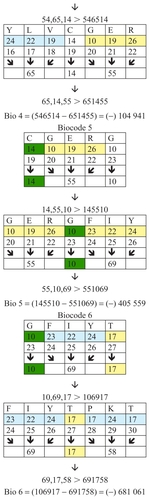
The biocodes presented in are calculated using the relationship between corresponding groups of amino acids. These are groups with different numbers of amino acids. There are different ways and methods of selecting these groups of amino acids, and it is hoped that science will soon determine which method is most efficient. Some biocodes have a positive numeric value and some have a negative one. gives these codes (see also ).
Figure 4 The formula for calculating of biocodes.
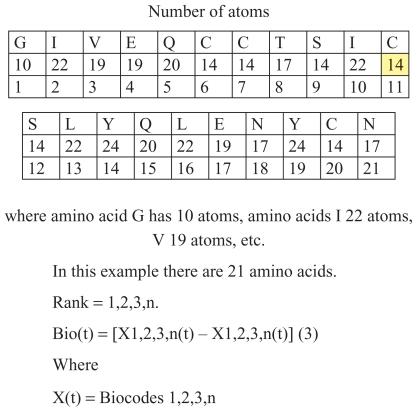

Figure 5 Examples of the connection of numbers of atoms in group of three amino acids of the insulin chain.
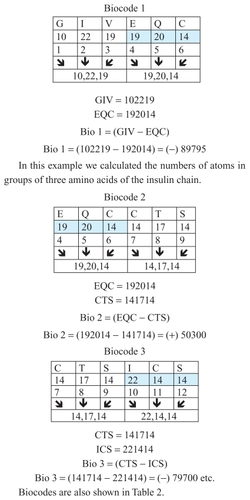

Table 1 Overview of positive and negative values of biocodes for insulin chain B showing some of the quantitative characteristics of the insulin molecule and the exact mathematical balance between its components. Schematic representation of the biocode processing (1)
Analog biocode
Insulin A chain
Each numerical value has its analog expression. For example, the analog expression for number 19 is 91 (91 || 19). In a similar way we can calculate the analog expression for any numerical value (). Our research has shown that analog codes are quantitative characteristics in biochemistry. An analog biocode is a discrete code that protects and guards genetic information coded in biochemical processes. This is a recently discovered code, and more detailed knowledge about it is necessary.
Figure 6 Examples of the analog codes of insulin chain A.
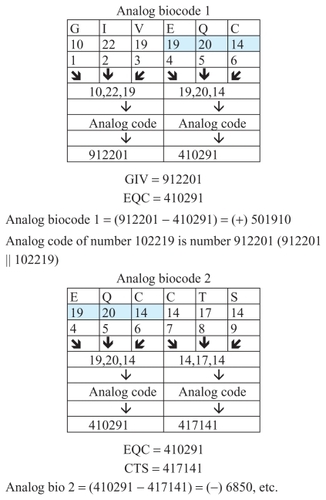
Figure 7 Group of chains E,F,G,H.
Positive and negative analog codes of insulin chain A are shown in .
Table 3 mathematical balance between positive and negative values of analog codes of insulin chain A. Schematic representation of biocode processing (3)
Correlation between code and analog code
Arithmetical expression for biocode = 190 001; there is a correlation between the code and analog code = 899 809; 100091 | | 190001 > 89 910; 908998 | | 899809 > 9189. This phenomenon will be investigated into more detail in the future.
Insulin model
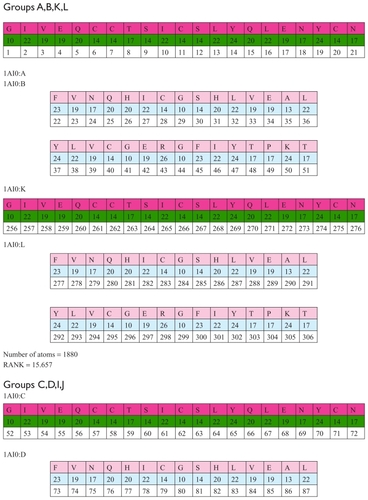
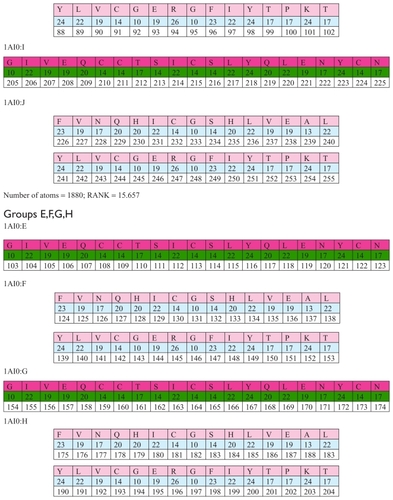
The structure 1AI0 has in total 12 chains, ie, A,B,C,D,E,F, G,H,I,J,K,L. In this group of chains there are three unions with four chains each. Each of these three groups of chains has an identical number of amino acids, identical number of atoms, and an identical sum of position numbers for these amino acids. shows the atomic structure 1A10.
Figure 9 Groups of chains (A,B,K,L), (C,D,I,J) and (E,F,G,H).
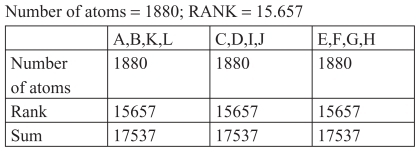
Table 2 Overview of the mathematical balance between the positive and negative values of the biocodes of the insulin A chain. Schematic representation of biocode processing (2)
Each of these three groups of chains has an identical number of amino acids, an identical number of atoms, and an identical sum of position numbers for these amino acids.
Code 6
The insulin matrix code can be developed into a periodic system in which each of 20 natural amino acids has an exactly established place. On the basis of this system, amino acids are distributed according to the criteria of mathematical similarity or difference ().
Figure 10 Distribution of amino acids according to the criteria of mathematical similarity or difference.
In this example, the quantitative characteristics of genetic information are given in the sign of the code 6. That code links the corresponding groups of amino acids.
Rank of the biocodes
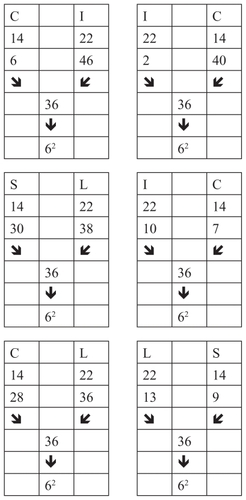
Some groups of amino acids that are discussed here are connected by the discrete code 11. Here are some examples ()
Figure 11 Groups of amino acids connected by the discrete code 11.
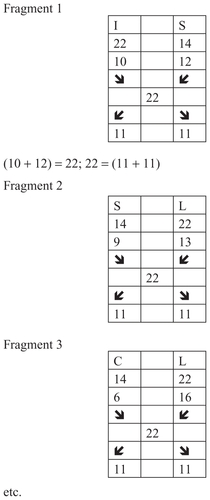
In the aforementioned examples, amino acids are connected by the code 6, and their position numbers are connected with code 11. As can be seen, the quantitative characteristics of the biochemistry of insulin apply to the position number.
Biocode 11
The aforementioned code 11 also connects groups of amino acids. We can see that in the following example ().
Figure 12 Discrete code 11 connects groups of amino acids.
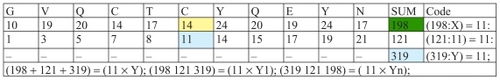
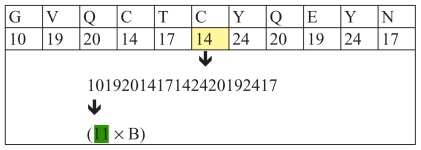
We will give a few more examples of coding of amino acids of insulin with code 11: (–).
Figure 13 Connection of the rank of amino acids G and N.
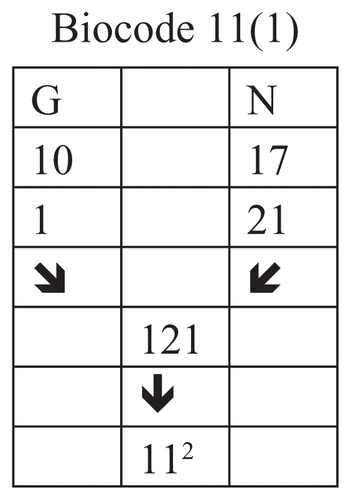
Figure 14 Connection of the rank of amino acids V and Y.
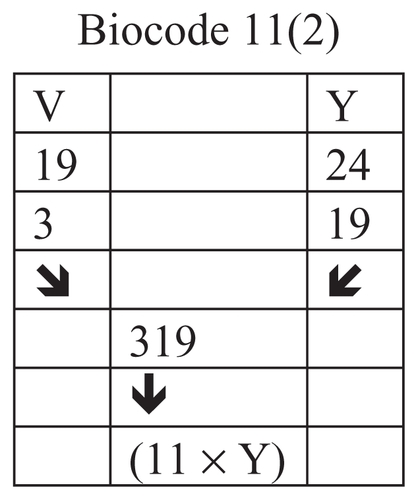
Figure 15 Connection of the rank of amino acids Q and E.
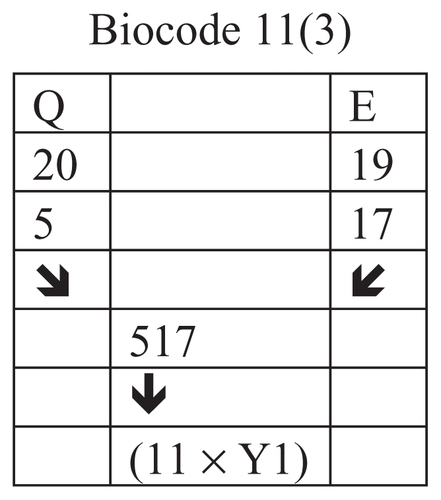
Figure 16 A schematic diagram to show the positive and negative output biocodes of the insulin A chain.
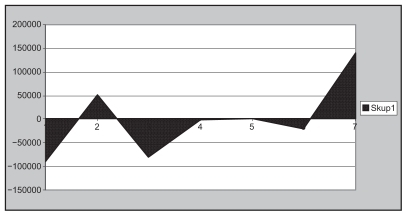
Figure 17 A schematic diagram to show the positive and negative output analog biocodes of the insulin C chain.
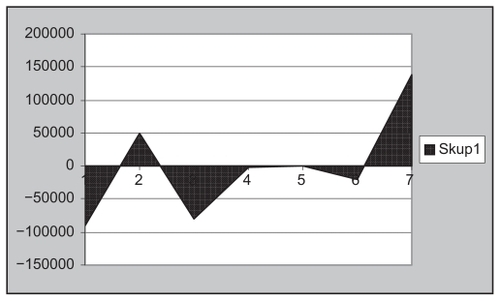
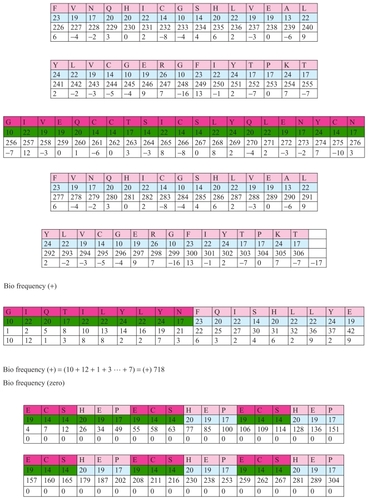
Figure 18 Frequency of insulin.
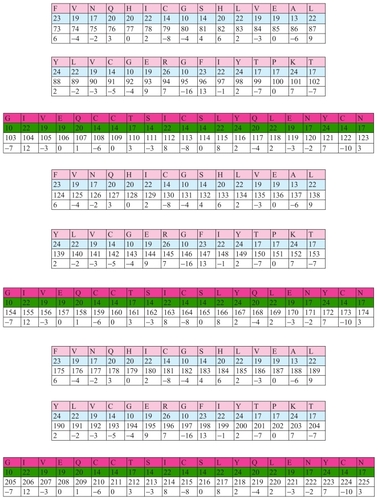
In this example we connected the position number of amino acids G and N, and got number 121 or 112 as result.
From the previous examples it can be seen that this protein has quantitative characteristics. It can be concluded that there is a connection between these quantitative characteristics in the process of transfer of genetic information and the qualitative appearance of the given genetic processes.
Biofrequency
r6 insulin hexamer (d1ai02)
Insulin is composed of amino acids with various numeric values. These numeric values are in an irregular order. For example, the first one has 10 atoms, the second one 22. Their frequency is X. The second amino acid has 22 atoms, and the third one 19. Their frequency is Y, etc. Frequency is the measurement for establishment of intervals of numeric values of amino acids in proteins. This value can be positive, negative, or zero. These frequencies demonstrate a completely new dimension of protein sequencing. Using these frequencies we can establish which of the amino acids are of primary and which are of secondary significance in the biochemical processes of insulin. A practical example: etc.
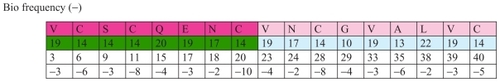
Bio frequency (−) = [(−3) + (−6) + (−3) … + (−17)] = (−)718
Results
Bio frequency = [(−) 718 ↔ (+) 718]
Therefore, there is a mathematical balance between the group of amino acids with a positive frequency and those with a negative frequency. Amino acids with a positive frequency have a primary role in the mathematical picture of that protein, and the negative frequencies have a secondary role in it. We assume that amino acids with a positive frequency have a primary role in the biochemical picture of that protein, and the negative frequencies have a secondary role in it. If this really is the case and research on an experimental level proves it, a radically new way of learning about biochemical processes will emerge.
Within insulin’s constituents, 133 amino acids have a positive frequency. These amino acids have 2848 atoms. There are 137 amino acids with a negative frequency. These amino acids have 2174 atoms. Number 133 from the union of acids with a positive frequency is the discrete number that we can use for decoding a matrix with positive and negative frequencies. Here are some examples: Example 1
[(137 + 2174) + (133 + 2848)] = [(133 + 133) + (718 × Y)]
Y = 7;
133 = (7 + 7 + 7 …, + 7)
Example 2
4712 || 2174
8482 || 2848
(4712 + 2174 + 8482 + 2848) = [(133 + 133) + (718 × Y)]
Y = 25
Levels of frequencies
In the group of amino acids with a positive frequency there are two subunions. One is the subunion with positive frequencies, and the other one is a subunion with negative frequencies. Also, in the union of amino acids with negative frequency, there are subunions with positive and negative frequencies. These groups also have their subunions with positive and negative frequencies. These groups demonstrate a new dimension of the biochemistry of insulin. We expect that this discovery will unlock many of the secrets of protein biochemistry in the future.
Discussion
The results of our research show that the process of sequencing molecules is conditioned and arranged not only according to chemical and biochemical laws, but also according to program, cybernetic, and informational laws. At the first stage of our research we replaced nucleotides from the amino acid code matrix with the numbers of atoms in those nucleotides. Translation of the biochemical language of these amino acids into a digital language may be very useful for developing new methods of predicting protein subcellular localization, membrane protein type, secondary protein structure prediction, or indeed any other protein attribute. Since the concept of Chou’s pseudo amino acid composition was proposed,Citation1,Citation2 there have been many attempts to use various digital numbers to represent the 20 native amino acids in order to reflect better the sequence-order effects involved. Some investigators have used complexity measure factor,Citation3 whereas others have used values derived from cellular automata,Citation4–Citation7 hydrophobic and/or hydrophilic values,Citation8–Citation16 Fourier transform techniques,Citation17,Citation18 or physicochemical distance.Citation19
It is going to be possible to use a completely new strategy of research in genetics in the future. However, close investigation of all these relationships, which are the outcomes of periodic laws (more specifically the law of binary coding) is necessary to be able to decode the conformational, stereo-chemical, and digital structure of proteins.
Disclosure
The author reports no conflict of interest in this research.
References
- ChouKCA new branch of proteomics: Prediction of protein cellular attributesWeinrerPWLuQGene Cloning and Expression TechnologiesWestborough, MAEaton Publishing2002
- ChouKCPrediction of protein cellular attributes using pseudo amino acid compositionProteins20014324625511288174
- XiaoXShaoSDingYHuangZHuangYChouKCUsing complexity measure factor to predict protein subcellular locationAmino Acids200528576115611847
- XiaoXShaoSDingYHuangZChenXChouCKUsing cellular automata to generate image representation for biological sequencesAmino Acids200528293515700108
- XiaoXShaoSDingYHuangZChenXChouKCAn application of gene comparative image for predicting the effect on replication ratio by HBV virus gene missense mutationJ Theor Biol200523555556515935173
- XiaoXShaoSHHuangZDChouKCUsing pseudo amino acid composition to predict protein structural classes: Approached with complexity measure factorJ Comput Chem20062747848216429410
- XiaoXShaoSHDingYSHuangZDChouKDUsing cellular automata images and pseudo amino acid composition to predict protein sub-cellular locationAmino Acids200630495416044193
- ChouKCUsing amphiphilic pseudo amino acid composition to predict enzyme subfamily classesBioinformatics20052110915308540
- ChouKCCaiYDPrediction of membrane protein types by incorporating amphipathic effectsJ Chem Inf Model20054540741315807506
- FengZPPrediction of the subcellular location of prokaryotic proteins based on a new representation of the amino acid compositionBiopolymers20015849149911241220
- FengZPAn overview on predicting the subcellular location of a proteinIn Silico Biol2002229130312542414
- WangMJYangJXuZJChouKCSLLE for predicting membrane protein typesJ Theor Biol200523271515498588
- WangSQYangJChouKCUsing stacked generalization to predict membrane protein types based on pseudo amino acid compositionJ Theor Biol200624294194616806277
- WangMYangJLiuGPXuZJChouKCWeighted-support vector machines for predicting membrane protein types based on pseudo amino acid compositionProtein Eng Des Sel20041750951615314209
- ZhangSWPanQZhangHCShaoZCShiJYPrediction protein homo-oligomer types by pseudo amino acid composition: Approached with an improved feature extraction and naive Bayes feature fusionAmino Acids20063046146816773245
- GaoYShaoSHXiaoHUsing pseudo amino acid composition to predict protein subcellular location: approached with Lyapunov index, Bessel function, and Chebyshev filterAmino Acids20052837337615889221
- GuoYZLiMLuMClassifying G protein-coupled receptors and nuclear receptors based on protein power spectrum from fast Fourier transformAmino Acids20063039740216773242
- LiuHWangMChouKCLow-frequency Fourier spectrum for predicting membrane protein typesBiochem Biophys Res Comm200533673773916140260
- ChouKCPrediction of protein subcellular locations by incorporating quasi-sequence-order effectBiochem Biophys Res Comm200027847748311097861
- ChouCKA novel approach to predicting protein structural classes in a (20-1)-D amino acid composition spaceProteins1995213193447567954
- ChouKCZhangCTPredicting protein folding types by distance functions that make allowances for amino acid interactionsJ Biol Chem199426922014220208071322
- ChouKCZhangCTReview: Prediction of protein structural classesCrit Rev Biochem Mol Biol1995302753497587280
- ChouKCElrodDWProtein subcellular location predictionProtein Eng19991210711810195282
- ChouKCReview: Prediction of protein structural classes and subcellular locationsCurr Protein Pept Sci2001117120812369916
- ChouKCElrodDWPrediction of membrane protein types and subcellular. LocationsProteins19993413715310336379
- ChouKCElrodDWPrediction of enzyme family classesJ Proteome Res2003218319012716132
- ChouKCCaiYDPredicting enzyme family class in a hybridization spaceProtein Sci2004132857286315498934
- ChouKCElrodDWBioinformatical analysis of G-protein-coupled receptorsJ Proteome Res2002142943312645914
- ChouKCPrediction of G-protein-coupled receptor classesJ Proteome Res200541413141816083294
- ChouKCCaiYDPrediction of protease types in a hybridization spaceBiochem Biophys Res Comm20063391015102016325146
- ChouKCCaiYDPredicting protein-protein interactions from sequences in a hybridization spaceJ Proteome Res2006531632216457597
- ChouKCCaiYDZhongWZPredicting networking couples for metabolic pathways of arabidopsisEXCLI Journal200655565
- ChouKCCaiYDPredicting protein quaternary structure by pseudo amino acid compositionProteins200353282914517979
- KuriLThe digital language of amino acidsAmino Acids200765366117252309
- KuriLThe atomic genetic codeJ Comput Biol2009210116
- KuriLMesure complexe des caracteristiques dynamiques de series temporellesJournal de la Societe de statistique de Paris. tome12721986