ABSTRACT
Acinetobacter nosocomialis is an important nosocomial pathogen that causes a variety of human infections. However, the specific virulence factors of this microorganism have not yet been determined. We investigated the role of outer membrane protein A (OmpA) in the pathogenesis of A. nosocomialis. A ΔompA mutant of the A. nosocomialis ATCC 17903T strain was constructed using markerless gene deletion. The ΔompA mutant displayed reduced biofilm formation in polystyrene tubes and reduced adherence to A549 cells in comparison to the wild-type strain. These virulence traits of the ΔompA mutant strain were restored when the ompA gene was complemented. Cytotoxicity was not significantly different between the wild-type strain and the ΔompA mutant when A549 cells were infected with bacteria or treated with outer membrane vesicles (OMVs). However, OMVs from the wild-type strain induced cytotoxicity in HEp-2 cells, whereas OMVs from the ΔompA mutant did not induce cytotoxicity. Proteomic analysis of OMVs revealed that OmpA influenced the distribution of envelope and periplasmic proteins. Overall, this study is the first report that links OmpA to A. nosocomialis pathogenesis, and highlights OmpA as a putative target to develop anti-virulence agents or vaccines against A. nosocomialis infection.
Introduction
Acinetobacter species are emerging nosocomial pathogens that cause a variety of human infections, especially in intensive care units.Citation1,2 Among the 39 genomic species described to date (http://www.bacterio.net/acinetobacter.html), Acinetobacter baumannii, Acinetobacter nosocomialis, and Acinetobacter pittii are the most clinically relevant species, notorious for the prevalence of nosocomial infections and multi-drug resistant phenotypes.Citation3-8 A. baumannii is the most prevalent species among the Acinetobacter species worldwide and the clinical importance of nosocomial infections caused by this microorganism has been well recognized by clinicians. A. nosocomialis is increasingly common as a causative agent of nosocomial infections and accounts for 21% of Acinetobacter species in the United States,Citation9 24.4% in Korea,Citation10 26.7% and 35.8% in Taiwan,Citation5,11 and 46.9% in Norway.Citation12
A. baumannii and A. nosocomialis are phenotypically similar and genetically closely related. They can be differentiated into different genomic species by molecular methods, including sequence analysis of the 16S rRNA gene, the rpoB gene,Citation13 and the 16S-23S rRNA gene spacer region,Citation14 as well as by amplified fragment length polymorphismCitation15 and amplified 16S rRNA gene restriction analysis.Citation16 However, previous studies have demonstrated that A. baumannii is associated with greater resistance to antimicrobial agents and higher mortality than A. nosocomialis,Citation5,17,18 which may result from differences in the biological and virulence traits between the two species. A. baumannii possesses several virulence traits, including biofilm formation,Citation19,20 cellular adherence and invasion of host cells,Citation21,22 serum resistance,Citation23,24 iron acquisition,Citation25,26 and cytotoxicity in host cells.Citation27-30 We recently reported that A. nosocomialis secrets cytotoxic factors via outer membrane vesicles (OMVs), which induce cytotoxicity and an innate immune response in epithelial cells in vitro.Citation31 However, the fundamental mechanisms of pathogenicity of A. nosocomialis remain to be elucidated.
Outer membrane protein A of A. baumannii (AbOmpA) is a major outer membrane porin that contributes significantly to the pathogenesis of A. baumannii. AbOmpA inhibits the alternative pathway of complement activation by binding factor H,23 and is also involved in biofilm formation on abiotic surfacesCitation32 and adherence to and invasion of host cells.Citation22 These strategies promote bacterial survival both inside and outside of the host. Furthermore, a large amount of AbOmpA is secreted via OMVs to stimulate innate immune responses and induce host cell death.Citation33,34 The secretion of AbOmpA via OMVs may be associated with host pathology upon A. baumannii infection in vivo. OmpA of A. nosocomialis ATCC 17903T shows 92% amino acid sequence homology with that of A. baumannii ATCC 17978T. Moreover, A. nosocomialis OmpA is secreted via OMVsCitation31 and carries putative nuclear localization signals that may direct OmpA to the nuclei of host cells.Citation28 Therefore, we postulated that the major outer membrane protein OmpA might directly or indirectly contribute to A. nosocomialis pathogenesis, similar to AbOmpA of A. baumannii. The aim of this study was to investigate the role of OmpA in the pathogenesis of A. nosocomialis by evaluating its effect on biofilm formation, adherence to host cells, and the induction of cytotoxicity in host cells. The virulence traits were compared for wild-type A. nosocomialis ATCC 17903T, an ompA deletion mutant, and ompA-complemented strains.
Results
Construction of A. nosocomialis ΔompA mutant and its ompA-complemented strain
To investigate whether OmpA played a role in the pathogenesis of A. nosocomialis, the ΔompA mutant of A. nosocomialis ATCC 17903T was constructed using a cloning method based on overlap extension polymerase chain reactions (PCR).Citation35 Two sets of PCRs were performed using the primer sets of OmpA01F/OmpA01R and OmpA02F/OmpA02R (), and two PCR products with a size of approximately 1.0 kb, respectively, were combined by overlap extension PCR using the primers OmpA01F/OmpA02R. The amplicon in which the ompA gene was deleted was cloned into the pOH1 plasmids, resulting in the pOH2 plasmid (). The ompA gene in the chromosome of A. nosocomialis ATCC 17903T was deleted by conjugal gene transfer of pOH2 plasmids and homologous recombination. To determine whether the ompA gene was deleted in the chromosome of A. nosocomialis ATCC 17903T, a PCR was performed using the primers Omp01F/Omp02R. The PCR products of the wild-type strain were 3.1 kb in size, whereas the ΔompA mutant OH1 strain showed a PCR product of approximately 2.1 kb (), which was in good agreement with the projected size of the DNA fragment after ompA deletion. For complementation of the ompA gene in the ΔompA mutant OH1 strain, the pOH3 plasmid was constructed by subcloning the ompA coding region under the control of the blaCTX-M14 gene promoter into the Acinetobacter-E. coli shuttle vector pWH1266 (). This recombinant plasmid was transformed into the ΔompA mutant OH1 strain, resulting in the ompA-complemented OH2 strain. PCR analysis using the primers OmpA04F/OmpA04R showed that the wild-type and the ompA-complemented OH2 strains carried the full-length 1,066 bp ompA gene, while it was not amplified in the ΔompA mutant OH1 strain (). To determine whether the wild-type, ΔompA mutant OH1, and ompA-complemented OH2 strains expressed OmpA in the outer membrane, outer membrane proteins were extracted from each of the three bacterial strains, and sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and subsequent western blotting with anti-AbOmpA antibodies were performed. The expected 37.4 kDa OmpA protein was expressed in the wild-type and the ompA-complemented OH2 strains, but was not detected in the outer membrane of the ΔompA mutant OH1 strain (). To determine whether the absence of OmpA in the outer membrane affected bacterial survival and growth, in vitro growth studies were carried out. The three bacterial strains were grown in Luria-Bertani (LB) medium at 37°C for 48 h, and colony forming units (CFUs) were determined at the indicated times. The growth of the ompA-complemented OH2 strains was slightly decreased at 24 h when compared to that of wild-type and ΔompA mutant OH1 strains, but no significant difference in growth rates was observed between the three bacterial strains ().
Figure 1. Generation of the ΔompA mutant OH1 strain and the ompA complemented OH2 strain. (A) PCR analysis of the genomic DNA of wild-type and ΔompA mutant OH1 strains using OmpA01F and OmpA01R primers. The genomic DNA of A. nosocomialis ATCC 17903T produced a 3.1 kb amplicon (lane 1), whereas the PCR using genomic DNA from the ΔompA mutant OH1 strain resulted in a 2.1 kb amplicon (lane 2). Lane M: 1-kb DNA molecular marker. (B) PCR analysis of the genomic DNA of wild-type, ΔompA mutant OH1, and ompA complemented OH2 strains using the OmpA04F and OmpA04R primers. The full-length ompA gene (expected size of 1,066 bp) was amplified in the wild-type (lane 1) and ompA complemented OH2 strains (lane 3). It was not amplified in the ΔompA mutant OH1 strain (lane 2). Lane M: 1-kb DNA molecular marker. (C) SDS-PAGE analysis and protein gel blot analysis of outer membrane proteins prepared from the wild-type (lane 1), ΔompA mutant OH1 (lane 2), and ompA complemented OH2 (lane 3) strains. Protein samples were separated by SDS-PAGE on a 12% gel, transferred to membranes, and immunoblotted with polyclonal anti-mouse AbOmpA immune serum. In lane M and 4, the molecular marker and recombinant OmpA protein with a molecular weight of 44 kDa were loaded, respectively. Arrows indicate the 37.4 kDa OmpA proteins. (D) Bacterial growth curve of A. nosocomialis strains cultured in LB broth under static conditions at 37°C for 48 h. CFUs were determined at the indicated time points. The data are the mean CFUs of duplicate in a representative of three independent experiments.
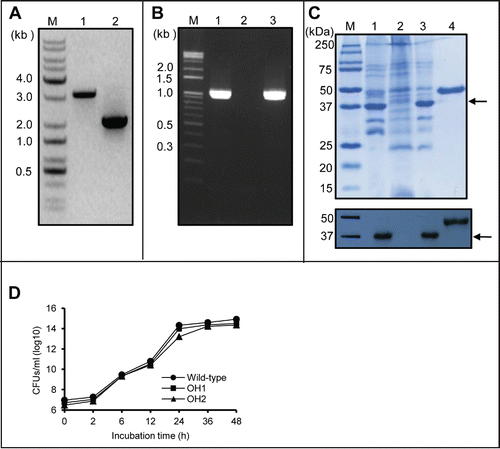
Table 1. Oligonucleotides used in this study.
Table 2. Bacterial strains and plasmids used in this study.
OmpA contributes to biofilm formation of A. nosocomialis
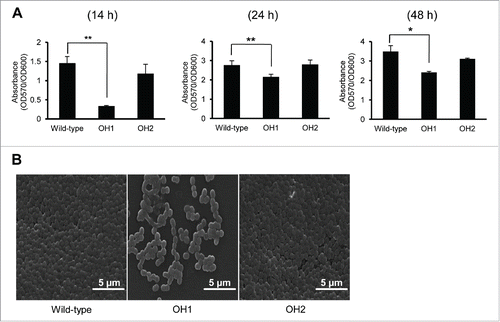
To evaluate the involvement of OmpA in biofilm formation, the ability of the wild-type and ΔompA mutant OH1 strains to produce biofilm on a polystyrene surface was assessed by crystal violet staining at different time points (). The ability of the wild-type strain to produce the biofilm was gradually increased when the bacteria was cultured for 14, 24, and 48 h. However, the ΔompA mutant OH1 strain showed a significant reduction in biofilm formation compared with the wild-type strain when bacteria were cultured during the periods of incubation. Rescue of the ompA gene in the ΔompA mutant restored the wild-type biofilm phenotype, as demonstrated by both the biofilm assay stained with crystal violet () and scanning electron microscopy (SEM) (). These results suggest that the decreased biofilm formation in the ΔompA mutant OH1 strain results from inactivation of the ompA gene rather than from any polar effect on genes downstream of the ompA gene.
Figure 2. Biofilm formation of A. nosocomialis strains. (A) Bacteria were cultured in polystyrene tubes at 30°C for 14 h, 24 h, and 48 h. Biofilm formation was quantified by calculating the ratio of A570/A600. (B) SEM analysis of bacterial biofilms formed on the plastic surfaces at 14 h. Means ± standard deviation were calculated based on the results of three independent experiments. Wild-type, A. nosocomialis ATCC 17903T; OH1, ΔompA mutant; OH2, ompA complemented strain. *,P < 0.05; **,P < 0.01.
OmpA contributes to adherence of A. nosocomialis to epithelial cells
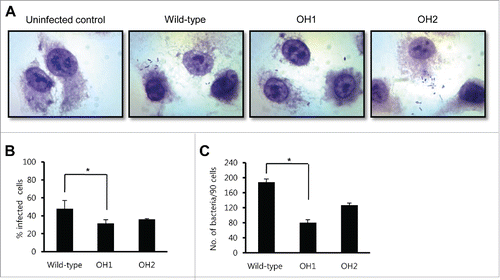
To investigate whether OmpA was involved in adherence of A. nosocomialis to epithelial cells, A549 cells were infected with the wild-type and the ΔompA mutant strains at a multiplicity of infection (MOI) of 100 for 1 h. The wild-type strain showed a dispersed adherence to A549 cells rather than an aggregated adherence at localized areas of the cells (). There was a significant difference in the percentage of infected cells between the wild-type strain (47.7% ± 9.4%) and the ΔompA mutant (31.7% ± 3.9%) (). Moreover, the ΔompA mutant showed a significant reduction in the number of adherent bacteria per 90 cells (80.5 ± 7.8) when compared with the wild-type strain (188.0 ± 8.5) (). The observed decrease in adherence for the ΔompA mutant OH1 strain was restored in the ompA-complemented OH2 strain (127.0 ± 5.7). These results suggest that OmpA plays a role in the adherence of A. nosocomialis in A549 epithelial cells.
Figure 3. Adherence of A. nosocomialis strains to A549 cells. (A) Cells were infected with A. nosocomialis at an MOI of 100 for 1 h and stained with Giemsa solution. Magnification: 100 ×. (B and C) A549 cells were infected with A. nosocomialis at an MOI of 100 for 1 h, and the percentage of infected cells (B) and cell-associated bacteria per 90 cells (C) were counted. The adherence data are expressed as mean ± standard deviation based on two independent experiments, each performed in triplicate. *,P < 0.05.
OmpA is not associated with epithelial cytotoxicity induced by A. nosocomialis infection
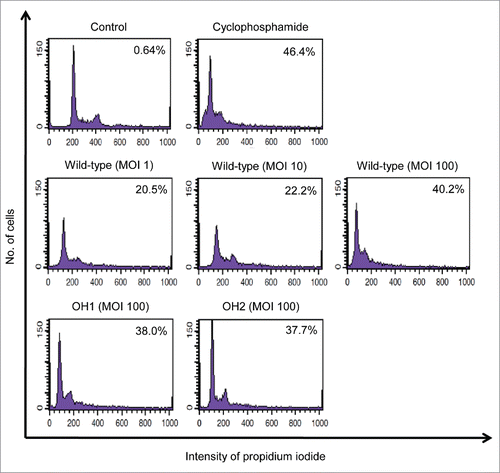
To determine whether A. nosocomialis induced cytotoxicity of epithelial cells, A549 cells were infected with the wild-type strain at an MOI of 1, 10, and 100 for 16 h. The cells infected with the bacteria were stained with propidium iodide (PI) and the hypo-diploid cells were counted using flow cytometry. The wild-type strain clearly induced cytotoxicity in A549 cells (, middle panel). To investigate whether OmpA contributed to cytotoxicity of epithelial cells, A549 cells were infected with the ΔompA mutant and the ompA-complemented strains at an MOI of 100 for 16 h. There was no significant difference in the cells with hypo-diploid DNA between the three A. nosocomialis strains (, middle and lower panels). These results suggest that OmpA does not directly contribute to the cytotoxicity of A549 cells induced by A. nosocomialis infection.
Figure 4. Cytotoxicity of A. nosocomialis infection for epithelial cells. A549 cells were infected with A. nosocomialis strains at an MOI of 100 for 16 h. Cells were stained with propidium iodide. The DNA content of each cell was analyzed using flow cytometry. A total of 9,000 cells were analyzed per sample. The percentage of cells in the hypo-diploid population is indicated in the diagrams. Representative data from three independent experiments are shown.
OmpA associated with OMVs is responsible for cytotoxicity of HEp-2 cells
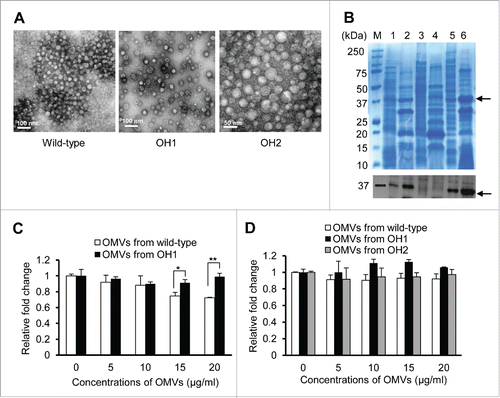
A. nosocomialis produces and secretes OMVs containing a large amount of OmpA, and A. nosocomialis OMVs induce cytotoxicity of HEp-2 cells.Citation31 To investigate whether OmpA in the A. nosocomialis OMVs was responsible for the cytotoxicity toward the epithelial cells, OMVs were purified from the wild-type, the ΔompA mutant, and the ompA-complemented strains (). SDS-PAGE and protein gel blotting analysis revealed that OmpA was detected in the OMVs purified from the wild-type and the ompA-complemented strains, but not in the OMVs from the ΔompA mutant (). We previously showed that cytotoxicity of HEp-2 cells was induced by treatment of 20 μg/ml of OMVs purified from A. nosocomialisCitation31 and, therefore, HEp-2 cells were treated with 5-20 μg/ml OMVs from the wild-type and the ΔompA mutant strains, and cell viability was analyzed using the WST-1 assay. The OMVs from the wild-type strain were cytotoxic to HEp-2 cells at a concentration of 15 μg/ml, whereas the OMVs from the ΔompA mutant did not induce cytotoxicity in HEp-2 cells (). The OMVs from the wild-type strain, the ΔompA mutant, and the ompA-complemented strain did not induce cytotoxicity in A549 cells treated with ≤20 μg/ml of OMVs (). These results suggest that OmpA in the A. nosocomialis OMVs is associated with epithelial cell death, but that the cytotoxic activity of A. nosocomialis OMVs is dependent on the epithelial cell type.
Figure 5. Purification of A. nosocomialis OMVs and their cytotoxicity in epithelial cells. (A) TEM analysis of OMVs from A. nosocomialis ATCC 17903T, ΔompA mutant OH1 strain, and ompA complemented OH2 strain. (B) SDS-PAGE and western blotting analysis of OMVs from A. nosocomialis strains. Lane 1, bacterial lysate of the wild-type strain; 2, OMVs from the wild-type strain; 3, bacterial lysate of the ΔompA mutant; 4, OMVs from the ΔompA mutant; 5, bacterial lysate of the ompA complemented strain; 6, OMVs from the ompA complemented strain. Arrows indicate the 37.4 kDa OmpA proteins. (C and D) HEp-2 (C) and A549 (D) cells were treated with OMVs from A. nosocomialis strains for 24 h. Cellular viability was determined by a WST-1 assay. The values are expressed as relative fold changes of optical density of cells treated with A. nosocomialis OMVs at A450 compared to those of untreated control cells. Data are presented as the mean ± standard deviation of three independent experiments, each performed in duplicate. *,P <0.05; **,P <0.01.
OmpA determines the composition of proteins in OMVs
Proteomic analysis of OMVs from the ΔompA mutant was performed to identify proteins associated with OMVs. We previously identified a total of 147 proteins in the OMVs of A. nosocomialis ATCC 17903T.Citation31 In this study, a total of 168 proteins were identified in the ΔompA mutant OMVs (Table S1). Among these, 87 (51.8%) overlapped with the proteins found in the OMVs from the wild-type strain (). Sixty proteins were identified only in the OMVs from the wild-type strain, whereas 81 were unique to the ΔompA mutant OMVs. Hypothetical protein W9I_03611 (gi|407441077), translocation protein TolB (gi|515963786), and hypothetical protein W9I_03176 (gi|407439503) were most abundant in the OMVs from the ΔompA mutant. In contrast, the most abundant proteins specific to the wild-type strain OMVs were hypothetical protein (gi|515187519), hypothetical protein (gi|446575849), and hypothetical protein W9I_00996 (gi|407440186).Citation31 These results suggest that OmpA may influence the distribution of membrane and periplasmic proteins, which may result in compositional difference in OMVs from the wild-type strain and the ΔompA mutant.
Figure 6. Venn diagram of proteins identified in OMVs from A. nosocomialis ATCC 17903T and ΔompA mutant OH1 strain. The number of proteins identified in the OMVs is presented. A total of 147 and 168 proteins were identified in OMVs from A. nosocomialis ATCC 17903T and the ΔompA mutant, respectively. Eighty-seven proteins were commonly identified in OMVs from both the wild-type strain and the ΔompA mutant.

Discussion
This study shows that OmpA plays a role in the pathogenesis of A. nosocomialis. OmpA expressed in the outer membrane of A. nosocomialis does not only contribute to biofilm formation on abiotic surfaces and adherence to human epithelial cells, but OmpA present in OMVs is also responsible for cytotoxicity in epithelial HEp-2 cells. These results indicate that OmpA is a specific virulence factor of A. nosocomialis that may contribute to pathogenesis upon infection in vivo.
The ability of nosocomial pathogens, including A. baumannii, to form biofilms on abiotic surfaces plays a pivotal role in the persistence and survival of these bacteria in hospital environments. Biofilm formation is considered to be an important virulence factor common to clinical isolates of A. baumannii.Citation20,36,37 The expression of pili-like structures on the surface of A. baumannii is essential for biofilm formation.Citation19 Although biofilm formation is a complex process involving a wide range of bacterial factors and multiple cellular signals, as well as a sequential expression of genes, several determinants associated with biofilm formation in A. baumannii have been characterized, including the CsuAB/ABCDE pilus chaperone-usher system,Citation19,38 the 854 kDa outer membrane protein with a high similarity to the staphylococcal biofilm-associated protein (Bap),Citation39 AbaI of the quorum sensing system,Citation40 and the pgaABCD operon responsible for the production of poly-β-1,6-N-acetyl glucosamine.Citation41 Assembly of pili and production of the Bap protein are responsible for the initiation and maturation of biofilms in abiotic surfaces, respectively. Gaddy et al.Citation32 reported that OmpA played a role in the biofilm formation of A. baumannii on abiotic surfaces. The csuAB/ABCDE genes essential for pili assembly was identified in the A. nosocomialis RUH2624 strain,Citation42 suggesting that the pili-like structures might also be associated with biofilm formation in A. nosocomialis. The present study clearly showed that OmpA contributed to the biofilm formation of A. nosocomialis in polystyrene tubes. The reduction of biofilm mass in the ΔompA mutant was most prominent after 14 h of culture and was significantly different for 48 h of culture (). These results suggest that OmpA may be associated with the initiation of biofilm development, although the present study did not specifically address whether OmpA plays a role in the initiation or maturation step of biofilm formation.
Adherence of pathogenic bacteria to epithelial cells is an essential initial step for colonization and infection. Lee et al.Citation21 demonstrated that A. baumannii showed two different types of adherence to human bronchial epithelial NCI-H292 cells: an aggregated adherence with connections of thread-like material between bacteria and a dispersed adherence with small pili-like structures connected to the surface of epithelial cells. The latter type of adherence was more frequently observed in clinical isolates of A. baumannii. Choi et al.Citation22 have demonstrated that OmpA plays a major role in the interactions of A. baumannii with epithelial cells in in vitro cell culture but also in an in vivo mouse pneumonia model. The present study demonstrated that A. nosocomialis ATCC 17903T showed a dispersed-like adherence to A549 cells. Although A. baumannii adherence to host cells was highly dependent on the epithelial cell type, the adherence efficiency of A. nosocomialis in A549 cells was similar to that of A. baumannii in NCI-H292 cells.Citation21,22 In the present study, A. nosocomialis infected 47% of A549 cells and the number of adhering bacteria per cell was 2.1 (). The ΔompA mutant showed a significant reduction in both the number of infected cells and adhering bacteria. These results suggest that the OmpA plays a role in the adherence of A. nosocomialis to epithelial cells. However, it should be determined whether OmpA is a non-fimbrial adhesin or influences the expression of other adhesins.
A. baumannii induces apoptosis of epithelial cells through cell surface death receptors and mitochondrial disintegration.Citation27 Furthermore, AbOmpA is considered to be an important cytotoxic factor through both mitochondrial and nuclear targeting.Citation27-29,43 The present study showed that A. nosocomialis ATCC 17903T induced cytotoxicity of A549 cells in an MOI-dependent manner (). In addition, both the ΔompA mutant and the ompA-complemented strains could induce cytotoxicity in A549 cells like the wild-type strain. Epithelial cells infected with A. nosocomialis strains showed morphological changes such as cellular shrinkage, round-up and detachment from the culture dish, and nuclear condensation (data not shown). These results suggest that A. nosocomialis can induce apoptosis of A549 cells, but that OmpA in the outer membrane is not directly responsible for cytotoxicity. Other bacterial factors are likely associated with cytotoxicity in A549 cells. A. baumannii secreted cytotoxic AbOmpA via OMVs.Citation33 Nho et al.Citation31 recently reported that A. nosocomialis secreted OMVs and these vesicles were also cytotoxic in HEp-2 cells. These OMVs delivered OmpA to the cytoplasm of host cells, but was unclear whether OmpA was associated with cytotoxicity in the case of A. nosocomialis OMVs. In the present study, OMVs from the wild-type strain could induce cytotoxicity observed upon A. nosocomial infection in HEp-2 cells, whereas OMVs from the ΔompA mutant did not induce cytotoxicity in HEp-2 cells (). HEp-2 cells were treated with 20 μg/ml OMVs from the ompA-complemented OH2 strain, but cytotoxicity was not induced (data not shown). One possible explanation for the defect of ompA complementation in the OMV-induced cytotoxicity is that the genetic elements other than the ompA gene in the pOH3 plasmids or antibiotics to maintain pOH3 plasmids in the ompA-complemented strain may influence on the composition or exact localization of proteins in OMVs. OMVs from the wild-type strain, the ΔompA mutant, and the ompA-complemented strain did not induce cytotoxicity in A549 cells at a concentration of ≤20 μg/ml. Our results suggest that OmpA plays a role in the cytotoxicity mediated by OMVs, and its cytotoxicity is dependent on the type of epithelial cells.
AbOmpA is a transmembrane protein in the outer membrane of A. baumannii and its C-terminal OmpA-like domain interacts with diaminopimelate (DAP) of peptidoglycan.Citation44 Moon et al.Citation45 demonstrated that OMV production in the ΔompA mutant constructed by random transposon mutagenesis was significantly higher than that of wild-type A. baumannii. Furthermore, proteomic analysis of OMVs showed that there were differences in protein content between the wild-type and the ΔompA mutant. The present study also showed a significant difference between the protein constituents of OMVs from the wild-type strain and from the ΔompA mutant. OmpA of A. nosocomialis ATCC 17903T and A. baumannii 19606T is composed of 347 and 356 amino acids, respectively. Differences in amino acid sequence homology between A. nosocomialis OmpA and A. baumannii AbOmpA were focused on the transmembrane domain, whereas the C-terminal OmpA-like domain (residues 226 to 328) of A. nosocomialis OmpA shows 100% homology with that of A. baumannii AbOmpA. Asp271 and Arg286 of AbOmpA are important among the residues in contact with DAP of peptidoglycan.Citation44 These two amino acids are also conserved in OmpA of A. nosocomialis. These results suggest that the C-terminal OmpA-like domain of A. nosocomialis OmpA may also interact with peptidoglycan. This may influence the protein composition of OMVs, especially for proteins located in the membrane and periplasmic space.
In conclusion, the present study demonstrates for the first time that OmpA is a specific virulence factor of A. nosocomialis. OmpA is considered to be a potential target to develop anti-virulence agents or vaccines against multi-drug resistant A. nosocomialis.
Materials and methods
Bacterial strains, plasmids, and culture conditions
The bacterial strains and plasmids used in this study are listed in . A. nosocomialis and E. coli were grown in LB media at 37°C. Chloramphenicol (20 μg/ml) or kanamycin (50 μg/ml) was added to the growth media to maintain the plasmids in E. coli. A. nosocomialis merodiploids were selected on the medium supplemented with kanamycin (30 μg/ml) and ampicillin (100 μg/ml). Bacterial growth was monitored by determining the optical density at 600 nm (OD at A600).
Cell culture
Type II pneumocyte cell line A549 originating from human lung carcinoma and HEp-2 originating from human laryngeal epithelial cells were obtained from the Korean Cell Line Bank (Seoul, Korea). A549 cells and HEp-2 cells were grown in RPMI 1640 medium (HyClone, Logan, UT) and Dulbecco's modified Eagle medium (HyClone) supplemented with 10% fetal bovine serum (FBS; HyClone), 2.0 mM L-glutamine, 100 U/ml penicillin, and 50 µg/ml streptomycin at 37°C in 5% CO2, respectively. Confluent cells were harvested and seeded into 12-, 24- or 96-well plates for infection with A. nosocomialis strains or incubation with A. nosocomialis OMVs.
DNA manipulations
Genomic and plasmid DNA was purified from bacteria using a SolGent™ Genomic DNA prep kit (SolGent, Daejeon, Korea) and AccuPrep® Plasmid Extraction Kit (Bioneer, Daejeon, Korea), respectively. DNA fragments were purified with an AccuPrep Gel Purification Kit (Bioneer). Restriction and DNA modifying enzymes were purchased from New England Biolabs (Ipswich, MA). Routine DNA manipulations were performed as previously describedCitation46 or according to the manufacturer's recommendations. Synthetic DNA fragments containing promoter of the blaCTX-M14 gene were purchased from Integrated DNA Technologies (IDT) (Coralville, IA).
Construction of the ΔompA mutant
To construct the ompA deletion mutant, a previously described markerless gene deletion method was modified.Citation35 A new suicide vector was constructed as follows: 1.2 kb nptI DNA conferring resistance to kanamycin was amplified using primers U1 and U2 () and the pUC4K vector was used as a template. The PCR product was cloned into pDM4 digested with SmaI. The new suicide vector was named pOH1 (). The ompA gene was inactivated in vitro by deletion of the ompA open reading frame (1,044 bp) using an overlap extension PCR method. To combine the upstream and downstream regions of the ompA gene by overlap extension PCR, a reverse primer, OmpA01R, for the amplification of the upstream region was designed to carry an additional 30 nucleotides at its 5′-end, which are homologous to the downstream region. The upstream and downstream regions of the ompA gene were amplified from the genomic DNA of A. nosocomialis ATCC 17903T using the primer pairs OmpA01F/OmpA01R and OmpA02F/OmpA02R, respectively (). The two PCR products obtained in the first step were mixed at equimolar concentrations and subjected to overlap extension PCR with the OmpA01F and OmpA02R primers. The resulting 2,075 bp DNA fragment, in which the upstream and downstream regions of the ompA gene were sequentially assembled, was ligated with XbaI-SpeI-digested pOH1 to generate pOH2 (). The correct assembly of the plasmids was confirmed by DNA sequencing. The plasmid pOH2 was integrated into the chromosome of A. nosocomialis by using conjugation-based gene transfer and homologous recombination. The E. coli S17-1 λ pir tra strain carrying pOH2 was used as a conjugal donor to A. nosocomialis. The conjugation and isolation of the transconjugants were conducted as follows: donor and recipient strains were grown in LB broth until late log phase (OD of 0.8 at A600). Bacterial cells were then mixed at an equal ratio and spotted onto an LB plate, and incubated for 12 h at 30°C. The bacteria were resuspended into LB broth and then plated on LB agar plates containing ampicillin and kanamycin to eliminate the donor strain and to select the merodiploid strain produced by first single cross-over homologous recombination. For plasmid excision, second single crossover homologous recombination of the bacteria was achieved by the sacB conferring sucrose sensitivity on LB agar plates with 10% sucrose and without NaCl. Bacteria susceptible to kanamycin were selected, and the deletion of the ompA gene was confirmed by PCR analysis.
Complementation of the ompA gene in the ΔompA mutant
For complementation of the ompA gene in the mutant strain, the ompA coding region under control of the blaCTX-M14 gene promoter with the kanamycin-resistance cassette was constructed by overlap extension PCR. The primers were designed as follow: A forward primer, OmpA03F, and reverse primer, OmpA03R, for amplification of the ompA coding region were designed to contain additional 25 nucleotides at their 5′ end that are homologous to the blaCTX-M14 gene promoter and kanamycin-resistance cassette, respectively (). The blaCTX-M14 gene promoter, ompA coding region, and nptI were amplified from the synthetic DNA fragment containing the blaCTX-M14 gene promoter, the genomic DNA of A. nosocomialis ATCC 17903T, and pUC4K using the primer pairs CtxF/CtxR, OmpA03F/OmpA03R, and U1/U2, respectively (). The three PCR products were mixed at equimolar concentrations and subjected to overlap extension PCR with the OmpA03F and U2 primers. The resulting DNA fragment, in which the blaCTX-M14 gene promoter, ompA coding region, and nptI were sequentially assembled, was ligated with PstI-SphI-digested pWH1266 to generate pOH03 (). The plasmid pOH3 was transformed into the ompA mutant by electroporation, and kanamycin-resistant colonies were selected.
Bacterial growth studies
Overnight cultures of A. nosocomialis strains were diluted 1:20 in LB broth and incubated for 48 h at 37°C under static conditions. Bacteria were sampled at the indicated times and CFUs as well as the OD at A600 were determined. Bacterial growth studies were performed in duplicate in a total of three independent experiments.
Preparation of outer membrane proteins and western blotting
A. nosocomialis strains were grown in LB broth to reach late logarithmic phase and were lysed by sonication. Outer membrane proteins were extracted with sodium lauryl sarcosine (Sigma-Aldrich, St. Louis, MO) and recovered by ultracentrifugation at 100,000 × g for 3 h at 4°C. The outer membrane protein profiles were determined by SDS-PAGE using 12% gels. Western blot analysis was performed following SDS-PAGE by incubating the Hybond-ECL membranes (Amersham Pharmacia Biotech, Piscataway, NJ) with a polyclonal anti-mouse AbOmpA immune serum.Citation31 The membranes were incubated with a secondary antibody coupled to horseradish peroxidase and developed using the ECL plus system (Amersham Pharmacia Biotech).
Purification of A. nosocomialis OMVs and western blotting
A. nosocomialis OMVs were prepared as previously described.Citation31 Briefly, A. nosocomialis strains were grown in 500 ml of LB broth to reach late log phase at 37°C with shaking. Bacteria were removed by centrifugation at 6,000 × g for 20 min at 4°C. The culture supernatants were filtered using a QuixStand Benchtop System (GE Healthcare, Buckinghamshire, UK) using a 0.2 μm-sized hollow fiber membrane (GE Healthcare) and then concentrated using a 100-kDa hollow fiber membrane (GE Healthcare). OMVs were collected by ultracentrifugation of the concentrated samples at 150,000 × g for 3 h at 4°C and resuspended in a small volume of phosphate-buffered saline (PBS). The protein concentration was determined using a modified BCA assay (Thermo Scientific, Waltham, MA). The purified OMVs were streaked onto blood agar plates to check sterility and stored at −70°C until use. Western blot analysis was performed to detect OmpA in the OMVs. The purified A. nosocomialis OMVs were separated via SDS-PAGE on 12% gels, followed by electrotransfer onto the Hybond-ECL membranes. The membranes were incubated with a polyclonal anti-mouse AbOmpA immune serum and then incubated with a secondary antibody coupled to horseradish peroxidase. The OmpA signals were detected using the ECL plus system.
Identification of proteins associated with OMVs
Proteins in the OMVs purified from the ΔompA mutant were identified using 1-dimensional gel electrophoresis and liquid chromatography-tandem mass spectrometry (1-DE-LC-MS/MS) as previously described.Citation31,33 All MS and MS/MS spectra were acquired in data-dependent mode. The MS/MS spectra were analyzed with MASCOT software (Matrix Science, Boston, MA) using all A. nosocomialis genome data from NCBInr (http://www.ncbi.nlm.nih.gov/) and the decoy sequence database. The locations of proteins were predicted using a subcellular location prediction program, Cello version 2.5 (http://cello.life.nctu.edu.tw/).
Biofilm assay
A biofilm formation assay was performed for all A. nosocomialis strains as previously described.Citation19,20 In brief, overnight cultures were adjusted to OD 2.0 at A600 and diluted 200-fold in LB medium containing 0.5% NaCl. Aliquots (1 or 2 ml) of the bacterial suspension were inoculated into 5 ml polystyrene tubes and incubated without shaking at 30°C for 48 h. Planktonic cells were removed and then the tubes were washed twice with 1 ml of PBS. The biofilm cells on the wall were stained with 0.1% (wt/vol) crystal violet solution for 15 min at room temperature. The biofilms were quantified at A570 using biofilm cell-associated dye, which was eluted with 100% ethanol, and normalized to total bacterial growth at A600. The biofilm assay was performed in triplicate in a total of three independent experiments.
Cell viability
The cytotoxicity of A. nosocomialis infection in A549 cells was measured using flow cytometric analysis as previously described.Citation27 Briefly, cells were infected with the wild-type, ΔompA mutant, and ompA complemented strains at an MOI of 1, 10, or 100 for 16 h. The adherent and detached cells were collected and resuspended in PBS containing 5% FBS. Cells were stained with PtdIns and the DNA content of 9,000 cells per sample was assessed in a FACSCalibur flow cytometry (BD Biosciences, San Jose, CA) by plotting the PI fluorescent intensity in a histogram plot. The cytotoxicity observed for A549 cells and HEp-2 cells after treatment with A. nosocomialis OMVs was measured using the Premix WST-1 cell proliferation assay system (Takara Bio, Otsu, Japan).Citation27 Cells were seeded at a concentration of 2.0 × 105 cells/ml in 96-well microplates. Cells were treated with different concentrations of OMVs from the wild-type, ΔompA mutant, and ompA complemented strains for 24 h and cellular cytotoxicity was measured at 450 nm, 3 h after treatment with WST1. Each experiment was performed in duplicate in minimum three independent experiments.
Adherence assay
The adherence evaluation of A. nosocomialis strains to A549 cells was performed as previously described.Citation21,22 Briefly, A549 cells were seeded at a density of 6 × 104 cells in 24-well culture dishes. Cells were infected with the A. nosocomialis strains at an MOI of 100 for 1 h. The cells were washed five times with PBS, fixed with methanol for 20 min, and stained with Giemsa solution. Bacterial adherence to the cells was determined by light microscopy. The percentage of infected cells (bacteria-associated cells/bacteria-non-associated cells × 100) and the number of bacteria associated with 90 cells were counted. Adherence assay were performed in triplicate in two independent experiments.
TEM and SEM analysis
The purified A. nosocomialis OMVs were applied to copper grids and stained with 2% uranyl acetate. The samples were visualized on a transmission electron microscope (TEM) (H-7500; Hitachi Ltd, Tokyo, Japan) operating at 120 kV. Biofilm formation of A. nosocomialis was examined by SEM (S-4800; Hitachi Ltd). Bacterial strains were statically cultured in 50-ml conical tubes with plastic coverslips semi-immersed in 5 ml of LB broth for 14 h at 30°C. Samples were pulled out of the LB media and processed as previously described.Citation19
Statistical analysis
The statistical significance of the data was calculated using Student's t-test. A P value of <0.05 was considered to be statistically significant.
Disclosure of potential conflicts of interest
No potential conflicts of interest were disclosed.
Author contributions
All authors contributed to the design of the experiments, analysis of the data, and writing of the manuscript. SWK and KHK performed the biofilm formation and bacterial adherence assays. MHO constructed the ompA deletion mutant and its complemented strain and performed biofilm assay and SEM analysis. SHJ and HJ performed cytotoxicity assay. SIK and YCL performed proteomic analysis.
KVIR_A_Supp_1140298.xlsx
Download MS Excel (22.9 KB)Funding
This research was supported by a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health &Welfare, Republic of Korea (grant number: HI14C0257).
References
- Dijkshoorn L, Nemec A, Seifert H. An increasing threat in hospitals: multidrug-resistant Acinetobacter baumannii. Nat Rev Microbiol 2007; 5:939-51; PMID:18007677; http://dx.doi.org/10.1038/nrmicro1789
- Peleg AY, Seifert H, Paterson DL. Acinetobacter baumannii: emergence of a successful pathogen. Clin Microbiol Rev 2008; 21:538-82; PMID:18625687; http://dx.doi.org/10.1128/CMR.00058-07
- Nemec A, Krizova L, Maixnerova M, van der Reijden TJ, Deschaght P, Passet V, Vaneechoutte M, Brisse S, Dijkshoorn L. Genotypic and phenotypic characterization of the Acinetobacter calcoaceticus-Acinetobacter baumannii complex with the proposal of Acinetobacter pittii sp. nov. (formerly Acinetobacter genomic species 3) and Acinetobacter nosocomialis sp. nov. (formerly Acinetobacter genomic species 13TU). Res Microbiol 2011; 162:393-404; PMID:21320596; http://dx.doi.org/10.1016/j.resmic.2011.02.006
- Wang J, Ruan Z, Feng Y, Fu Y, Jiang Y, Wang H, Yu Y. Species distribution of clinical Acinetobacter isolates revealed by different identification techniques. PLoS One 2014; 9:e104882; PMID:25120020; http://dx.doi.org/10.1371/journal.pone.0104882
- Lee YC, Huang YT, Tan CK, Kuo YW, Liao CH, Lee PtdIns, Hsueh PR. Acinetobacter baumannii and Acinetobacter genospecies 13TU and 3 bacteraemia: comparison of clinical features, prognostic factors and outcomes. J Antimicrob Chemother 2011; 66:1839-46; PMID:21653602; http://dx.doi.org/10.1093/jac/dkr200
- Schleicher X, Higgins PG, Wisplinghoff H, Körber-Irrgang B, Kresken M, Seifert H. Molecular epidemiology of Acinetobacter baumannii and Acinetobacter nosocomialis in Germany over a 5-year period (2005-2009). Clin Microbiol Infect 2013; 19:737-42; PMID:23034071; http://dx.doi.org/10.1111/1469-0691.12026
- Chusri S, Chongsuvivatwong V, Rivera JI, Silpapojakul K, Singkhamanan K, McNeil E, Doi Y. Clinical outcomes of hospital-acquired infection with Acinetobacter nosocomialis and Acinetobacter pittii. Antimicrob Agents Chemother 2014; 58:4172-9; PMID:24820079; http://dx.doi.org/10.1128/AAC.02992-14
- Lee YT, Huang LY, Chiang DH, Chen CP, Chen TL, Wang FD, Fung CP, Siu LK, Cho WL. Differences in phenotypic and genotypic characteristics among imipenem-non-susceptible Acinetobacter isolates belonging to different genomic species in Taiwan. Int J Antimicrob Agents 2009; 34:580-4; PMID:19733035; http://dx.doi.org/10.1016/j.ijantimicag.2009.06.027
- Wisplinghoff H, Paulus T, Lugenheim M, Stefanik D, Higgins PG, Edmond MB, Wenzel RP, Seifert H. Nosocomial bloodstream infections due to Acinetobacter baumannii, Acinetobacter pittii and Acinetobacter nosocomialis in the United States. J Infect 2012; 64:282-90; PMID:22209744; http://dx.doi.org/10.1016/j.jinf.2011.12.008
- Lee SY, Shin JH, Kim SH, Shin MG, Suh SP, Ryang DW. Evaluation of matrix-assisted laser desorption ionization-time of flight mass spectrometry-based VITEK MS system for the identification of Acinetobacter species from blood cultures: comparison with VITEK 2 and MicroScan systems. Ann Lab Med 2015; 35:62-8; PMID:25553282; http://dx.doi.org/10.3343/alm.2015.35.1.62
- Chuang YC, Sheng WH, Li SY, Lin YC, Wang JT, Chen YC, Chang SC. Influence of genospecies of Acinetobacter baumannii complex on clinical outcomes of patients with Acinetobacter bacteremia. Clin Infect Dis 2011; 52:352-60; PMID:21193494; http://dx.doi.org/10.1093/cid/ciq154
- Karah N, Haldorsen B, Hegstad K, Simonsen GS, Sundsfjord A, Samuelsen Ø. Species identification and molecular characterization of Acinetobacter spp. blood culture isolates from Norway. J Antimicrob Chemother 2011; 66:738-44; PMID:21393175; http://dx.doi.org/10.1093/jac/dkq521
- Gundi VA, Dijkshoorn L, Burignat S, Raoult D, La Scola B. Validation of partial rpoB gene sequence analysis for the identification of clinically important and emerging Acinetobacter species. Microbiol 2009; 155:2333-41; PMID:19389786; http://dx.doi.org/10.1099/mic.0.026054-0
- Chang HC, Wei YF, Dijkshoorn L, Vaneechoutte M, Tang CT, Chang TC. Species-level identification of isolates of the Acinetobacter calcoaceticus-Acinetobacter baumannii complex by sequence analysis of the 16S-23S rRNA gene spacer region. J Clin Microbiol 2005; 43:1632-9; PMID:15814977; http://dx.doi.org/10.1128/JCM.43.4.1632-1639.2005
- Janssen P, Maquelin K, Coopman R, Tjernberg I, Bouvet P, Kersters K, Dijkshoorn L. Discrimination of Acinetobacter genomic species by AFLP fingerprinting. Int J Syst Bacteriol 1997; 47:1179-87; PMID:9336926; http://dx.doi.org/10.1099/00207713-47-4-1179
- Dijkshoorn L, Van Harsselaar B, Tjernberg I, Bouvet PJ, Vaneechoutte M. Evaluation of amplified ribosomal DNA restriction analysis for identification of Acinetobacter genomic species. Syst Appl Microbiol 1998; 21:33-9; PMID:9786720; http://dx.doi.org/10.1016/S0723-2020(98)80006-4
- Park KH, Shin JH, Lee SY, Kim SH, Jang MO, Kang SJ, Jung SI, Chung EK, Ko KS, Jang HC. The clinical characteristics, carbapenem resistance, and outcome of Acinetobacter bacteremia according to genospecies. PLoS One 2013; 8:e65026; PMID:23755171; http://dx.doi.org/10.1371/journal.pone.0065026
- Chiang MC, Kuo SC, Chen SJ, Yang SP, Lee YT, Chen TL, Fung CP. Clinical characteristics and outcomes of bacteremia due to different genomic species of Acinetobacter baumannii complex in patients with solid tumors. Infection 2011; 40:19-26; PMID:21887526; http://dx.doi.org/10.1007/s15010-011-0187-4
- Tomaras AP, Dorsey CW, Edelmann RE, Actis LA. Attachment to and biofilm formation on abiotic surfaces by Acinetobacter baumannii: involvement of a novel chaperone-usher pili assembly system. Microbiology 2003; 149:3473-84; PMID:14663080; http://dx.doi.org/10.1099/mic.0.26541-0
- Lee HW, Koh YM, Kim J, Lee JC, Lee YC, Seol SY, Cho DT, Kim J. Capacity of multidrug-resistant clinical isolates of Acinetobacter baumannii to form biofilm and adhere to epithelial cell surfaces. Clin Microbiol Infect 2008; 14:49-54; PMID:18005176; http://dx.doi.org/10.1111/j.1469-0691.2007.01842.x
- Lee JC, Koerten H, van den Broek P, Beekhuizen H, Wolterbeek R, van den Barselaar M, van der Reijden T, van der Meer J, van de Gevel J, Dijkshoorn L. Adherence of Acinetobacter baumannii strains to human bronchial epithelial cells. Res Microbiol 2006; 157:360-6; PMID:16326077; http://dx.doi.org/10.1016/j.resmic.2005.09.011
- Choi CH, Lee JS, Lee YC, Park TI, Lee JC. Acinetobacter baumannii invades epithelial cells and outer membrane protein A mediates interactions with epithelial cells. BMC Microbiol 2008; 8:216; PMID:19068136; http://dx.doi.org/10.1186/1471-2180-8-216
- Kim SW, Choi CH, Moon DC, Jin JS, Lee JH, Shin JH, Kim JM, Lee YC, Seol SY, Cho DT, et al. Serum resistance of Acinetobacter baumannii through the binding of factor H to outer membrane proteins. FEMS Microbiol Lett 2009; 301:224-31; PMID:19878322; http://dx.doi.org/10.1111/j.1574-6968.2009.01820.x
- de Léséleuc L, Harris G, KuoLee R, Xu HH, Chen W. Serum resistance, gallium nitrate tolerance and extrapulmonary dissemination are linked to heme consumption in a bacteremic strain of Acinetobacter baumannii. Int J Med Microbiol 2014; 304:360-9; http://dx.doi.org/10.1016/j.ijmm.2013.12.002
- Gaddy JA, Arivett BA, McConnell MJ, López-Rojas R, Pachón J, Actis LA. Role of acinetobactin-mediated iron acquisition functions in the interaction of Acinetobacter baumannii strain ATCC 19606T with human lung epithelial cells, Galleria mellonella caterpillars, and mice. Infect Immun 2012; 80:1015-24; PMID:22232188; http://dx.doi.org/10.1128/IAI.06279-11
- Gentile V, Frangipani E, Bonchi C, Minandri F, Runci F, Visca P. Iron and Acinetobacter baumannii biofilm formation. Pathogens 2014; 3:704-19; PMID:25438019; http://dx.doi.org/10.3390/pathogens3030704
- Choi CH, Lee EY, Lee YC, Park TI, Kim HJ, Hyun SH, Kim SA, Lee SK, Lee JC. Outer membrane protein 38 of Acinetobacter baumannii localizes to the mitochondria and induces apoptosis of epithelial cells. Cell Microbiol 2005; 7:1127-38; PMID:16008580; http://dx.doi.org/10.1111/j.1462-5822.2005.00538.x
- Choi CH, Hyun SH, Lee JY, Lee JS, Lee YS, Kim SA, Chae JP, Yoo SM, Lee JC. Acinetobacter baumannii outer membrane protein A targets the nucleus and induces cytotoxicity. Cell Microbiol 2008; 10:309-19; PMID:17760880
- Lee JS, Choi CH, Kim JW, Lee JC. Acinetobacter baumannii outer membrane protein A induces dendritic cell death through mitochondrial targeting. J Microbiol 2010; 48:387-92; PMID:20571958; http://dx.doi.org/10.1007/s12275-010-0155-1
- Rumbo C, Tomás M, Fernández Moreira E, Soares NC, Carvajal M, Santillana E, Beceiro A, Romero A, Bou G. The Acinetobacter baumannii Omp33-36 porin is a virulence factor that induces apoptosis and modulates autophagy in human cells. Infect Immun 2014; 82:4666-80; PMID:25156738; http://dx.doi.org/10.1128/IAI.02034-14
- Nho JS, Jun SH, Oh MH, Park TI, Choi CW, Kim SI, Choi CH, Lee JC. Acinetobacter nosocomialis secretes outer membrane vesicles that induce epithelial cell death and host inflammatory responses. Microb Pathog 2015; 81:39-45; PMID:25778390; http://dx.doi.org/10.1016/j.micpath.2015.03.012
- Gaddy JA, Tomaras AP, Actis LA. The Acinetobacter baumannii 19606 OmpA protein plays a role in biofilm formation on abiotic surfaces and the interaction of this pathogen with eukaryotic cells. Infect. Immun 2009; 77:3150-60; PMID:19470746; http://dx.doi.org/10.1128/IAI.00096-09
- Jin JS, Kwon SO, Moon DC, Gurung M, Lee JH, Kim SI, Lee JC. Acinetobacter baumannii secretes cytotoxic outer membrane protein A via outer membrane vesicles. PLoS One 2011; 6:e17027; PMID:21386968; http://dx.doi.org/10.1371/journal.pone.0017027
- Jun SH, Lee JH, Kim BR, Kim SI, Park TI, Lee JC, Lee YC. Acinetobacter baumannii outer membrane vesicles elicit a potent innate immune response via membrane proteins. PLoS One 2013; 8:e71751; PMID:23977136; http://dx.doi.org/10.1371/journal.pone.0071751
- Oh MH, Lee JC, Kim J, Choi CH, Han K. Simple method for markerless gene deletion in multidrug-resistant Acinetobacter baumannii. Appl Environ Microbiol 2015; 81:3357-68; PMID:25746991; http://dx.doi.org/10.1128/AEM.03975-14
- Rodríguez-Baño J, Martí S, Soto S, Fernández-Cuenca F, Cisneros JM, Pachón J, Pascual A, Martínez-Martínez L, McQueary C, Actis LA, et al. Biofilm formation in Acinetobacter baumannii: associated features and clinical implications. Clin Microbiol Infect 2008; 14:276-8; http://dx.doi.org/10.1111/j.1469-0691.2007.01916.x
- King LB, Swiatlo E, Swiatlo A, McDaniel LS. Serum resistance and biofilm formation in clinical isolates of Acinetobacter baumannii. FEMS Immunol Med Microbiol 2009; 55:414-21; PMID:19220466; http://dx.doi.org/10.1111/j.1574-695X.2009.00538.x
- de Breij A, Gaddy J, van der Meer J, Koning R, Koster A, van den Broek P, Actis L, Nibbering P, Dijkshoorn L. CsuA/BABCDE-dependent pili are not involved in the adherence of Acinetobacter baumannii ATCC19606T to human airway epithelial cells and their inflammatory response. Res Microbiol 2009; 160:213-8; PMID:19530313; http://dx.doi.org/10.1016/j.resmic.2009.01.002
- Loehfelm TW, Luke NR, Campagnari AA. Identification and characterization of an Acinetobacter baumannii biofilm-associated protein. J Bacteriol 2008; 190:1036-44; PMID:18024522; http://dx.doi.org/10.1128/JB.01416-07
- Niu C, Clemmer KM, Bonomo RA, Rather PN. Isolation and characterization of an autoinducer synthase from Acinetobacter baumannii. J Bacteriol 2008; 190:3386-92; PMID:18281398; http://dx.doi.org/10.1128/JB.01929-07
- Choi AH, Slamti L, Avci FY, Pier GB, Maira-Litrán T. The pgaABCD locus of Acinetobacter baumannii encodes the production of poly-β-1-6-N-acetylglucosamine, which is critical for biofilm formation. J Bacteriol 2009; 191:5953-63; PMID:19633088; http://dx.doi.org/10.1128/JB.00647-09
- Peleg AY, de Breij A, Adams MD, Cerqueira GM, Mocali S, Galardini M, Nibbering PH, Earl AM, Ward DV, Paterson DL, et al. The success of acinetobacter species; genetic, metabolic and virulence attributes. PLoS One 2012; 7:e46984; PMID:23144699; http://dx.doi.org/10.1371/journal.pone.0046984
- Choi CH, Hyun SH, Kim J, Lee YC, Seol SY, Cho DT, Lee JC. Nuclear translocation and DNAse I-like enzymatic activity of Acinetobacter baumannii outer membrane protein A. FEMS Microbiol Lett 2008; 288:62-7; PMID:18783439; http://dx.doi.org/10.1111/j.1574-6968.2008.01323.x
- Park JS, Lee WC, Yeo KJ, Ryu KS, Kumarasiri M, Hesek D, Lee M, Mobashery S, Song JH, Kim SI, et al. Mechanism of anchoring of OmpA protein to the cell wall peptidoglycan of the gram-negative bacterial outer membrane. FASEB J 2012; 26:219-28; PMID:21965596; http://dx.doi.org/10.1096/fj.11-188425
- Moon DC, Choi CH, Lee JH, Choi CW, Kim HY, Park JS, Kim SI, Lee JC. Acinetobacter baumannii outer membrane protein A modulates the biogenesis of outer membrane vesicles. J Microbiol 2012; 50:155-60; PMID:22367951; http://dx.doi.org/10.1007/s12275-012-1589-4
- Sambrook J, Russell D. Molecular cloning: a laboratory manual, 3rd ed. 2001. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
- Simon R, Priefer U, Pühler A. A broad host range mobilization system for in vivo genetic engineering transposon mutagenesis in gram negative bacteria. Nat Biotechnol 1983; 1:784-91; http://dx.doi.org/10.1038/nbt1183-784
- Milton DL, O'Toole R, Horstedt P, Wolf-Watz H. Flagellin A is essential for the virulence of Vibrio anguillarum. J Bacteriol 1996; 178:1310-9; PMID:8631707
- Hunger M, Schmucker R, Kishan V, Hillen W. Analysis and nucleotide sequence of an origin of DNA replication in Acinetobacter calcoaceticus and its use for Escherichia coli shuttle plasmids. Gene 1990; 87:45-51; PMID:2185139; http://dx.doi.org/10.1016/0378-1119(90)90494-C
