Figures & data
Table 1. Primers for PCR amplification of formaldehyde dismutase genes of FD1.
Figure 1. Amino acid alignment of the three Fdms of FD1, the Fdm of P. putida F61, and the glutathione-independent Fdh of P. putida C-83.
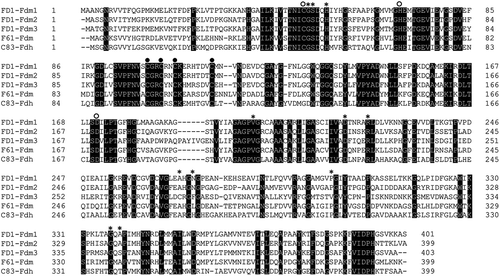
Figure 2. SDS-PAGE of Fdm1 (a), Fdm2 (b), and Fdm3 (c) of FD1 purified from each recombinant E. coli by IMAC.
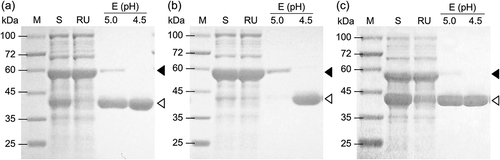
Table 2. Purification of Fdm1, Fdm2, and Fdm3 of FD1.
Figure 3. Effect of pH (a) and temperature (b) on the formaldehyde dismutase activity of the Fdms of FD1.
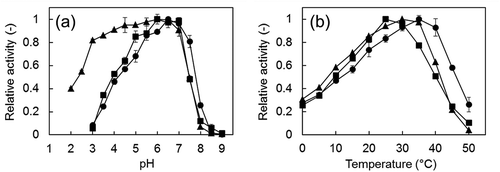
Figure 4. Effect of formaldehyde concentration on the formaldehyde dismutase activity of the Fdms of FD1 and the Lineweaver-Burk plots (inset).
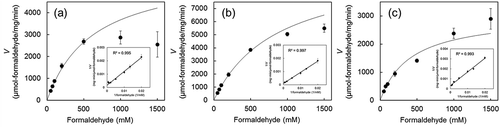
Table 3. Enzymatic kinetic parameters of Fdm1, Fdm2, and Fdm3 of FD1.
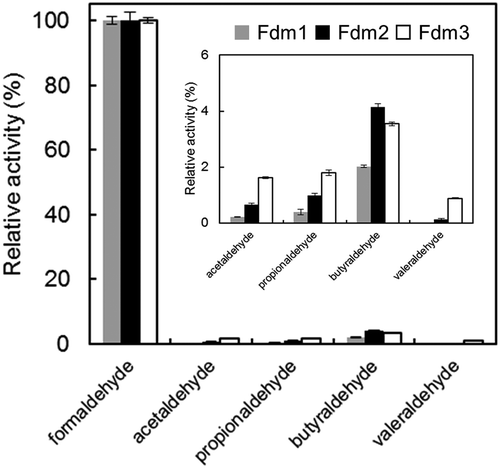
Table 4. Alcohol-NDMA oxidoreductase activities of Fdm1, Fdm2, and Fdm3 of FD1.