Figures & data
Figure 1. Tehama variety chromatogram which identifies:(1) gallic acid, (2) gallocatechin, (3) catechin, (4) procyanidin B2, (5) gallate epigallocatechin, (6) epicatechin and (7) gallate epicatechin at 280 nm. (8) syringic acid, (9) chlorogenic acid and (10) p-coumaric acid at 320 nm and (11) ellagic acid identified at 360 nm.
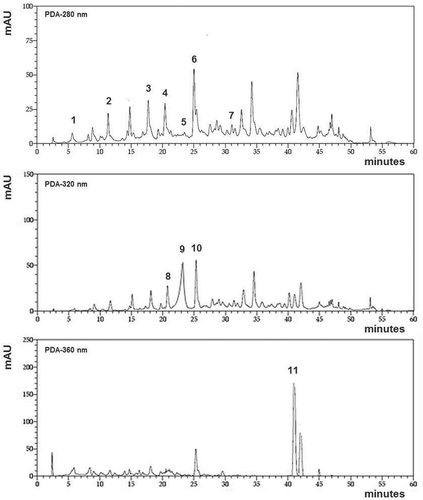
Table 1. Flavonoid phenolic compounds quantified in the different genotypes of walnuts.
Figure 2. Availability of flavonoid compounds identified after “in vitro” digestion. *Indicates statistical differences (p < 0.05).
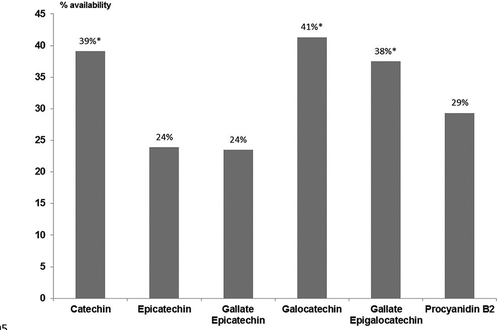
Table 2. Phenolic acids quantified in the different genotypes of walnuts.
Table 3. Flavonoid phenolic compounds quantified in the dialyzed fraction of the different genotypes of walnuts.
Figure 3. Availability of non-flavonoid compounds identified after “in vitro” digestion. *Indicates statistical differences (p < 0.05) with ellagic acid and syringic acid.
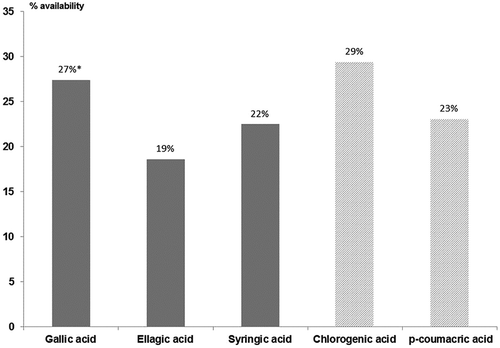
Table 4. Phenolic acids quantified in the dialyzed fraction of the different genotypes of walnuts.
