Figures & data
Table 1. List of Leea species collected in different Philippine provinces showing USTH voucher numbers.
Figure 1. Field photographs of Leea species. (A) Habit and (B) infructescence of Leea aculeata. (C) Habit and (D) close-up of inflorescence of Leea guineensis.

Table 2. Universal primers of the four candidate barcodes used in the study.
Table 3. Sequence characteristics from multiple sequence alignment of the four candidate barcodes used in this study.
Figure 2. Efficiency of polymerase chain reaction amplification and sequencing success rates of the four candidate barcodes for the selected Leea sp.
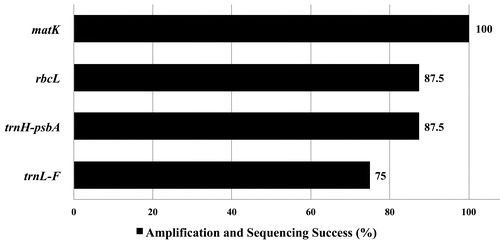
Table 4. Calculated mean interspecific and intraspecific divergences of the four candidate barcodes using Kimura two-parameter.
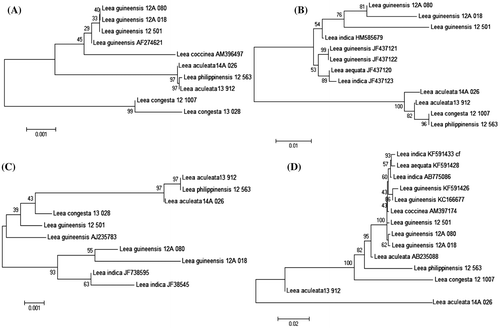
Figure 3. Neighbour-joining tree of (A) matK, (B) rbcL, (C) trnH-psbA and (D) trnL-F sequences inferred using Kimura two-parameter distances.