Figures & data
Table 1. Significant correlations with data corrected for cell-type (versus uncorrected data)
Figure 1. Exposure-attributable FEV1 decline pathway correlations. (a) Correlation between exposure-attributable changes (6H–0H) in FEV1 and log uLTE4 levels. r = −0.51, p = 0.04. (b) Correlation between exposure-attributable changes (6H–0H) in CysLTR1 cg26848126 and log uLTE4 levels. r = 0.59, p = 0.02. (c) Correlation between exposure-attributable changes (6H–0H) in CysLTR1 expression and log uLTE4 levels. r = −0.45, p = 0.08. (d) Correlation between exposure-attributable changes (6H–0H) in CysLTR1 expression and CysLTR1 cg26848126 methylation. r = −0.49, p = 0.06. (e) Correlation between exposure-attributable changes (6H-0H) in FEV1 and CysLTR1 expression r = 0.76, p = 0.0007
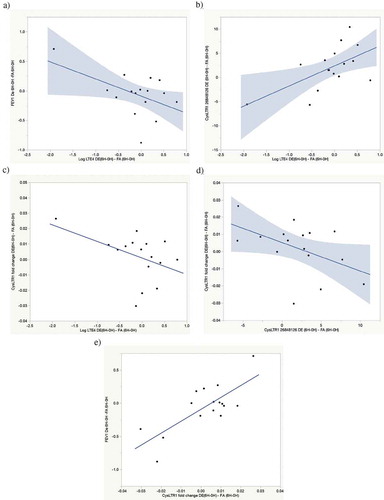
Figure 2. Exposure-attributable FEV1 increase pathway correlations. (a) Correlation between exposure-attributable changes (6H-0H) in CysLTR1 cg0081399 methylation and GPR17 cg05361273 methylation r = −0.73, p = 0.001. (b) Correlation between exposure-attributable changes (6H-0H) in CysLTR1 expression and GPR17 cg05361273 methylation r = 0.47, p = 0.07. 9 c) Correlation between exposure-attributable changes (6H-0H) in CysLTR1 expression and CysLTR1 cg0081399 methylation r = −0.49, p = 0.06
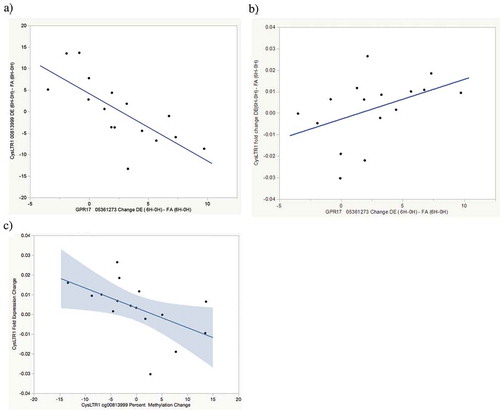
Figure 3. Biological pathways linking DE exposure to acute FEV1 changes through methylation and expression changes on CysLTR1-related genes
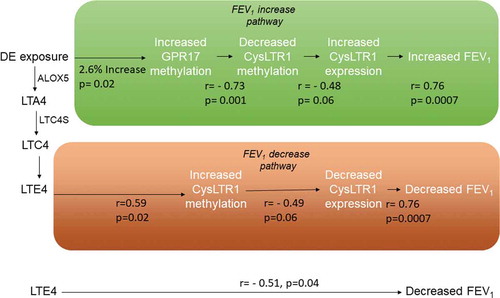
Table 2. CpG percent methylation by pyrosequencing at baseline and changes after DE, FA
Table 3. Participant characteristics
Table 4. Primers used for gene expression analysis
Table 5. Primers used for methylation analysis
Availability of data and materials
The data set analyzed during the current study are available from the corresponding authors on reasonable request.
